

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147404.5 + phase: 0
(498 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
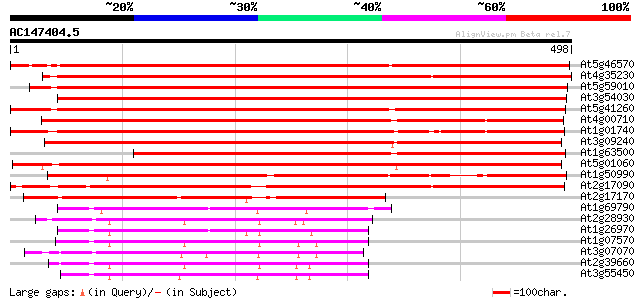
Score E
Sequences producing significant alignments: (bits) Value
At5g46570 protein kinase-like protein 810 0.0
At4g35230 unknown protein 607 e-174
At5g59010 protein kinase - like 603 e-173
At3g54030 protein kinase -like protein 591 e-169
At5g41260 protein kinase-like 586 e-167
At4g00710 unknown protein 581 e-166
At1g01740 protein kinase, putative 526 e-149
At3g09240 putative protein kinase 505 e-143
At1g63500 protein kinase, putative 499 e-141
At5g01060 putative protein - kinase 495 e-140
At1g50990 hypothetical protein 437 e-123
At2g17090 putative protein kinase 400 e-112
At2g17170 hypothetical protein 233 2e-61
At1g69790 protein kinase like protein 164 1e-40
At2g28930 protein kinase like protein 161 9e-40
At1g26970 protein kinase like protein 158 6e-39
At1g07570 protein kinase APK1A 155 4e-38
At3g07070 putative protein kinase 155 6e-38
At2g39660 putative protein kinase 155 6e-38
At3g55450 serine/threonine-specific protein kinase -like 154 1e-37
>At5g46570 protein kinase-like protein
Length = 489
Score = 810 bits (2091), Expect = 0.0
Identities = 402/497 (80%), Positives = 449/497 (89%), Gaps = 9/497 (1%)
Query: 1 MGCFHSKTAHLHSPEDPPTALPDSKKTDPGDDGGDGDQESQVPVFKEYGLNELRRATHEF 60
MGC HSKTA+L S +DP + P+ ++ GD DQE Q FKE+ LNELR+AT+ F
Sbjct: 1 MGCLHSKTANLPSSDDP--SAPNKPESVNGDQV---DQEIQN--FKEFELNELRKATNGF 53
Query: 61 STEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMV 120
S IVSE GEKAPNVVY+GKLE N LVA+KRFS+QSWPDAQQF+ EA GVGK+R+KR+V
Sbjct: 54 SPSCIVSEGGEKAPNVVYRGKLEGNHLVAIKRFSRQSWPDAQQFVVEATGVGKLRNKRIV 113
Query: 121 NLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDKQPLPWEMRVRVAYHVAQALDHCSMENR 180
+LIGCCAEGDERLLVAEYMPNDTLSKHLFHW+KQPLPW+MRVR+A ++A+ALD+C++ENR
Sbjct: 114 SLIGCCAEGDERLLVAEYMPNDTLSKHLFHWEKQPLPWDMRVRIADYIAEALDYCNIENR 173
Query: 181 KIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGKSYSTNLAYTPPEFLRTGRIIAESVIY 240
KIYHDLNAYRILFDE+GDPRLS+FGLMKNSRDGKSYSTNLAYTPPEFLRTGR+I ESVI+
Sbjct: 174 KIYHDLNAYRILFDEEGDPRLSTFGLMKNSRDGKSYSTNLAYTPPEFLRTGRVIPESVIF 233
Query: 241 SYGTVLLDLLSGKHIPPSHALDLIRGKNALLLMDSSLEGQYANDDATKLVELASKCLQFE 300
SYGT+LLDLLSGKHIPPSHALD+IRGKNALLLMDSSLEGQYANDDATKLV+LASKCLQ E
Sbjct: 234 SYGTILLDLLSGKHIPPSHALDIIRGKNALLLMDSSLEGQYANDDATKLVDLASKCLQSE 293
Query: 301 ARERPDIKFLLTAVTPLQKQKEVASHVLMGLTKTPAVLPLPTMLSPLGKACARMDLTAIH 360
A++RPD KFLL+AV PLQKQ+EVASHVLMGL K + LPTMLSPLGKACA+MDL H
Sbjct: 294 AKDRPDTKFLLSAVAPLQKQEEVASHVLMGLPKNTVI--LPTMLSPLGKACAKMDLATFH 351
Query: 361 DILLKTGYKDEEGAENELSFQEWTQQVQDILNTKKFGDIAFRDKDFKNAIEYYSKLVVMM 420
DILLKTGY+DEEGAENELSFQEWTQQVQ++LNTKKFGDIAFRDKDFKN+IEYYSKLV MM
Sbjct: 352 DILLKTGYRDEEGAENELSFQEWTQQVQEMLNTKKFGDIAFRDKDFKNSIEYYSKLVGMM 411
Query: 421 SVPSATVFARRAFAYLMNDQAELALRDAMQAQVCIPDWPTAFYLQALALSKLGMETDAQD 480
VPSATVFARRAF+YLM DQ ELALRDAMQAQVCIP+WPTAFYLQALALSKLGMETDAQD
Sbjct: 412 PVPSATVFARRAFSYLMTDQQELALRDAMQAQVCIPEWPTAFYLQALALSKLGMETDAQD 471
Query: 481 MLNDGAAFEAKRSNSWR 497
MLNDGAA++AKR NSWR
Sbjct: 472 MLNDGAAYDAKRQNSWR 488
>At4g35230 unknown protein
Length = 512
Score = 607 bits (1564), Expect = e-174
Identities = 293/470 (62%), Positives = 371/470 (78%), Gaps = 7/470 (1%)
Query: 30 GDDGGDGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVA 89
G GG G +P F E+ +L+ AT+ FS++ IVSESGEKAPN+VYKG+L+N R +A
Sbjct: 48 GGGGGGG-----IPSFSEFSFADLKAATNNFSSDNIVSESGEKAPNLVYKGRLQNRRWIA 102
Query: 90 VKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLF 149
VK+F+K +WP+ +QF EA GVGK+RH R+ NLIG C +GDERLLVAE+MPNDTL+KHLF
Sbjct: 103 VKKFTKMAWPEPKQFAEEAWGVGKLRHNRLANLIGYCCDGDERLLVAEFMPNDTLAKHLF 162
Query: 150 HWDKQPLPWEMRVRVAYHVAQALDHCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKN 209
HW+ Q + W MR+RV Y++A+ALD+CS E R +YHDLNAYR+LFDEDGDPRLS FGLMKN
Sbjct: 163 HWENQTIEWAMRLRVGYYIAEALDYCSTEGRPLYHDLNAYRVLFDEDGDPRLSCFGLMKN 222
Query: 210 SRDGKSYSTNLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIRGKNA 269
SRDGKSYSTNLAYTPPE+LR GR+ ESV YS+GTVLLDLLSGKHIPPSHALD+IRGKN
Sbjct: 223 SRDGKSYSTNLAYTPPEYLRNGRVTPESVTYSFGTVLLDLLSGKHIPPSHALDMIRGKNI 282
Query: 270 LLLMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAVTPLQKQKEVASHVLM 329
+LLMDS LEG+++ ++AT +VELAS+CLQ+E RERP+ K L+ + PLQ + +V S+V++
Sbjct: 283 ILLMDSHLEGKFSTEEATVVVELASQCLQYEPRERPNTKDLVATLAPLQTKSDVPSYVML 342
Query: 330 GLTKTPAVLPLPTM-LSPLGKACARMDLTAIHDILLKTGYKDEEGAENELSFQEWTQQVQ 388
G+ K P LSPLG+AC+RMDLTAIH IL+ T Y+D+EG NELSFQEWTQQ++
Sbjct: 343 GIKKQEEAPSTPQRPLSPLGEACSRMDLTAIHQILVMTHYRDDEGT-NELSFQEWTQQMK 401
Query: 389 DILNTKKFGDIAFRDKDFKNAIEYYSKLVVMMSVPSATVFARRAFAYLMNDQAELALRDA 448
D+L+ +K GD +FR+KDFK AI+ YS+ + + ++ S TVF RR+ YL+ DQ + ALRDA
Sbjct: 402 DMLDARKRGDQSFREKDFKTAIDCYSQFIDVGTMVSPTVFGRRSLCYLLCDQPDAALRDA 461
Query: 449 MQAQVCIPDWPTAFYLQALALSKLGMETDAQDMLNDGAAFEAKRSNSWRG 498
MQAQ PDWPTAFY+Q++AL+KL M TDA DMLN+ A E KR RG
Sbjct: 462 MQAQCVYPDWPTAFYMQSVALAKLNMNTDAADMLNEAAQLEEKRQRGGRG 511
>At5g59010 protein kinase - like
Length = 489
Score = 603 bits (1556), Expect = e-173
Identities = 296/478 (61%), Positives = 372/478 (76%), Gaps = 4/478 (0%)
Query: 18 PTALPDSKKTDPGDDGGDGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVV 77
PT L + D G D +P F E+ ++LR AT FST+ IVSE G KAPNVV
Sbjct: 14 PTHLKSTHNEASDLDNGTDD----LPSFTEFSFDQLRAATCGFSTDSIVSEHGVKAPNVV 69
Query: 78 YKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAE 137
YKG+LE++R +AVKRF++ +WPD +QFL EA VG++R++R+ NLIG C EGDERLLVAE
Sbjct: 70 YKGRLEDDRWIAVKRFNRSAWPDTRQFLEEAKAVGQLRNERLANLIGFCCEGDERLLVAE 129
Query: 138 YMPNDTLSKHLFHWDKQPLPWEMRVRVAYHVAQALDHCSMENRKIYHDLNAYRILFDEDG 197
+MP +TLSKHLFHWD QP+ W MR+RVA ++AQAL++CS + R +YHDLNAYRILFD+DG
Sbjct: 130 FMPFETLSKHLFHWDSQPMKWSMRLRVALYLAQALEYCSSKGRALYHDLNAYRILFDQDG 189
Query: 198 DPRLSSFGLMKNSRDGKSYSTNLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPP 257
+PRLS FGLMKNSRDGKSYSTNLA+TPPE+LRTGR+I ESV+YS+GT+LLDLLSGKHIPP
Sbjct: 190 NPRLSCFGLMKNSRDGKSYSTNLAFTPPEYLRTGRVIPESVVYSFGTLLLDLLSGKHIPP 249
Query: 258 SHALDLIRGKNALLLMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAVTPL 317
SHALDLIRGKN L+LMDS L+G ++NDD T LV LAS+CLQ+EARERP++K L++++ PL
Sbjct: 250 SHALDLIRGKNFLMLMDSCLDGHFSNDDGTDLVRLASRCLQYEARERPNVKSLVSSLAPL 309
Query: 318 QKQKEVASHVLMGLTKTPAVLPLPTMLSPLGKACARMDLTAIHDILLKTGYKDEEGAENE 377
QK+ ++ SHVLMG+ A T L+PLG AC+R DLTAIH+IL K GYKD+EG NE
Sbjct: 310 QKETDIPSHVLMGIPHGAASPKETTSLTPLGDACSRHDLTAIHEILEKVGYKDDEGVANE 369
Query: 378 LSFQEWTQQVQDILNTKKFGDIAFRDKDFKNAIEYYSKLVVMMSVPSATVFARRAFAYLM 437
LSFQ WT Q+Q+ LN+KK GD AF+ KDF A+E Y++ + ++ S TVFARR YLM
Sbjct: 370 LSFQVWTDQIQETLNSKKQGDAAFKGKDFVTAVECYTQFIEDGTMVSPTVFARRCLCYLM 429
Query: 438 NDQAELALRDAMQAQVCIPDWPTAFYLQALALSKLGMETDAQDMLNDGAAFEAKRSNS 495
++ + AL DAMQAQV P+WPTAFYLQA AL LGM+ DA + L DG + EAK+ N+
Sbjct: 430 SNMPQEALGDAMQAQVVSPEWPTAFYLQAAALFSLGMDKDACETLKDGTSLEAKKHNN 487
>At3g54030 protein kinase -like protein
Length = 490
Score = 591 bits (1523), Expect = e-169
Identities = 285/453 (62%), Positives = 362/453 (78%), Gaps = 1/453 (0%)
Query: 43 PVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQ 102
P FKE+ L +L+ AT FS++ IVSE GEKAPNVVY+G+L++ RL+AVKRF++ +W D +
Sbjct: 36 PTFKEFKLEQLKSATGGFSSDNIVSEHGEKAPNVVYRGRLDDGRLIAVKRFNRLAWADHR 95
Query: 103 QFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDKQPLPWEMRV 162
QFL EA VG +R R+ NLIGCC EG+ERLLVAE+MP++TL+KHLFHW+ P+ W MR+
Sbjct: 96 QFLDEAKAVGSLRSDRLANLIGCCFEGEERLLVAEFMPHETLAKHLFHWENNPMKWAMRL 155
Query: 163 RVAYHVAQALDHCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGKSYSTNLAY 222
RVA +AQAL++CS + R +YHDLNAYR+LFD+DG+PRLS FGLMKNSRDGKSYSTNLA+
Sbjct: 156 RVALCLAQALEYCSNKGRALYHDLNAYRVLFDKDGNPRLSCFGLMKNSRDGKSYSTNLAF 215
Query: 223 TPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIRGKNALLLMDSSLEGQYA 282
TPPE+LRTGR+ ESV++S+GTVLLDL+SGKHIPPSHALDLIRGKN +LMDS+LEG ++
Sbjct: 216 TPPEYLRTGRVTPESVVFSFGTVLLDLMSGKHIPPSHALDLIRGKNCAMLMDSALEGHFS 275
Query: 283 NDDATKLVELASKCLQFEARERPDIKFLLTAVTPLQKQKEVASHVLMGLT-KTPAVLPLP 341
N+D T+LV LA++CLQ+EARERP++K L+T++ LQK+ +VAS+VLMG+ +T A P
Sbjct: 276 NEDGTELVRLATRCLQYEARERPNVKSLVTSLVTLQKESDVASYVLMGIPHETEAEEESP 335
Query: 342 TMLSPLGKACARMDLTAIHDILLKTGYKDEEGAENELSFQEWTQQVQDILNTKKFGDIAF 401
L+P G AC R+DLTAI +IL K GYKD+EG NELSFQ WT Q+Q+ LN+KK GD+AF
Sbjct: 336 LSLTPFGDACLRVDLTAIQEILSKIGYKDDEGIANELSFQMWTNQMQESLNSKKQGDLAF 395
Query: 402 RDKDFKNAIEYYSKLVVMMSVPSATVFARRAFAYLMNDQAELALRDAMQAQVCIPDWPTA 461
R KDF A++ Y++ + ++ S TV ARR +YLMND A+ AL DA+QAQV PDWPTA
Sbjct: 396 RSKDFTTAVDCYTQFIDGGTMVSPTVHARRCLSYLMNDNAQEALTDALQAQVVSPDWPTA 455
Query: 462 FYLQALALSKLGMETDAQDMLNDGAAFEAKRSN 494
YLQA L KLGME DAQ L DG EAK+SN
Sbjct: 456 LYLQAACLFKLGMEADAQQALKDGTTLEAKKSN 488
>At5g41260 protein kinase-like
Length = 487
Score = 586 bits (1510), Expect = e-167
Identities = 292/493 (59%), Positives = 368/493 (74%), Gaps = 8/493 (1%)
Query: 1 MGCFHSKTAHLHSPEDPPTALPDSKKTDPGDDGGDGDQESQVPVFKEYGLNELRRATHEF 60
MGC SK + L + + PD D G D +P F+E+ + +R AT F
Sbjct: 1 MGCEVSKLSALCCVSESGRSNPDVTGLDEEGRGESND----LPQFREFSIETIRNATSGF 56
Query: 61 STEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMV 120
+ E IVSE GE+APNVVYKGKLEN R +AVKRF+++SWPD++QFL EA VG++R+ RM
Sbjct: 57 AAENIVSEHGERAPNVVYKGKLENQRRIAVKRFNRKSWPDSRQFLEEAKAVGQLRNHRMA 116
Query: 121 NLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDKQPLPWEMRVRVAYHVAQALDHCSMENR 180
NL+GCC E +ERLL+AE+MPN+TL+KHLFHW+ QP+ W MR+RVA H+AQAL++C+ + R
Sbjct: 117 NLLGCCYEDEERLLIAEFMPNETLAKHLFHWESQPMKWAMRLRVALHIAQALEYCTSKGR 176
Query: 181 KIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGKSYSTNLAYTPPEFLRTGRIIAESVIY 240
+YHDLNAYR+LFD+D +PRLS FGLMKNSRDGKSYSTNLA+TPPE+LRTGR+ ESVIY
Sbjct: 177 ALYHDLNAYRVLFDDDANPRLSCFGLMKNSRDGKSYSTNLAFTPPEYLRTGRVTPESVIY 236
Query: 241 SYGTVLLDLLSGKHIPPSHALDLIRGKNALLLMDSSLEGQYANDDATKLVELASKCLQFE 300
S+GT+LLDLLSGKHIPPSHALDLIR +N +LMDS LEGQ+++DD T+L+ LAS+CLQ+E
Sbjct: 237 SFGTLLLDLLSGKHIPPSHALDLIRDRNIQMLMDSGLEGQFSSDDGTELIRLASRCLQYE 296
Query: 301 ARERPDIKFLLTAVTPLQKQKEVASHVLMGLTKTPAVLPLPTMLSPLGKACARMDLTAIH 360
RERP+ K L++A+ PLQK E+ASH L+G+ + T LSPLG+AC R DLTAIH
Sbjct: 297 PRERPNPKSLVSAMIPLQKDLEIASHQLLGVPNSATT----TALSPLGEACLRSDLTAIH 352
Query: 361 DILLKTGYKDEEGAENELSFQEWTQQVQDILNTKKFGDIAFRDKDFKNAIEYYSKLVVMM 420
+I+ K GYKD+EGA ELSFQ WT Q+QD L KK GD AFR KDF AIE YS+ + +
Sbjct: 353 EIIEKLGYKDDEGATTELSFQMWTDQMQDTLVFKKKGDSAFRHKDFAKAIECYSQFIEVG 412
Query: 421 SVPSATVFARRAFAYLMNDQAELALRDAMQAQVCIPDWPTAFYLQALALSKLGMETDAQD 480
++ S TV AR++ YLMND AL +AMQAQV P W A YLQA+ALS LG E +A
Sbjct: 413 TMGSPTVHARQSLCYLMNDMPREALNNAMQAQVISPAWHIASYLQAVALSALGQENEAHT 472
Query: 481 MLNDGAAFEAKRS 493
L DGA E+KR+
Sbjct: 473 ALKDGAMLESKRN 485
>At4g00710 unknown protein
Length = 489
Score = 581 bits (1498), Expect = e-166
Identities = 287/463 (61%), Positives = 364/463 (77%), Gaps = 5/463 (1%)
Query: 29 PGDDGGDGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLV 88
P D G+ + + VP F+EY L +L+ AT F+ EYIVSE GEKAPNVVYKGKLEN + +
Sbjct: 24 PDVDNGESSEITDVPNFREYTLEQLKAATSGFAVEYIVSEHGEKAPNVVYKGKLENQKKI 83
Query: 89 AVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHL 148
AVKRF++ +WPD++QFL EA VG++R +RM NL+GCC EGDERLLVAE+MPN+TL+KHL
Sbjct: 84 AVKRFTRMAWPDSRQFLEEARSVGQLRSERMANLLGCCCEGDERLLVAEFMPNETLAKHL 143
Query: 149 FHWDKQPLPWEMRVRVAYHVAQALDHCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMK 208
FHW+ QP+ W MR+RV ++AQAL++C+ + R +YHDLNAYR+LFDE+ +PRLS+FGLMK
Sbjct: 144 FHWETQPMKWTMRLRVVLYLAQALEYCTSKGRTLYHDLNAYRVLFDEECNPRLSTFGLMK 203
Query: 209 NSRDGKSYSTNLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIRGKN 268
NSRDGKSYSTNLA+TPPE+LRTGRI ESVIYS+GT+LLDLLSGKHIPPSHALDLIR +N
Sbjct: 204 NSRDGKSYSTNLAFTPPEYLRTGRITPESVIYSFGTLLLDLLSGKHIPPSHALDLIRDRN 263
Query: 269 ALLLMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAVTPLQKQKEVASHVL 328
L DS L+GQ+++ D T+LV LAS+CLQ+EARERP+ K L+TA+TPLQK+ EV SHVL
Sbjct: 264 LQTLTDSCLDGQFSDSDGTELVRLASRCLQYEARERPNTKSLVTALTPLQKETEVLSHVL 323
Query: 329 MGLTKTPAVLPLPTMLSPLGKACARMDLTAIHDILLKTGYKDEEGAENELSFQEWTQQVQ 388
MGL + +V P LSPLG+AC+R DLTA+ +IL K GYKD+EG NELSF WT Q+Q
Sbjct: 324 MGLPHSGSVSP----LSPLGEACSRRDLTAMLEILEKLGYKDDEGVTNELSFHMWTDQMQ 379
Query: 389 DILNTKKFGDIAFRDKDFKNAIEYYSKLVVMMSVPSATVFARRAFAYLMNDQAELALRDA 448
+ LN+KK GD+AFR KDF+ AIE Y++ + + S TV ARR+ YLM+D + AL DA
Sbjct: 380 ESLNSKKKGDVAFRQKDFREAIECYTQFIDGGMI-SPTVCARRSLCYLMSDMPKEALDDA 438
Query: 449 MQAQVCIPDWPTAFYLQALALSKLGMETDAQDMLNDGAAFEAK 491
+QAQV P W A YLQ+ +L LGME ++Q L +G+ EAK
Sbjct: 439 IQAQVISPVWHVASYLQSASLGILGMEKESQIALKEGSNLEAK 481
>At1g01740 protein kinase, putative
Length = 483
Score = 526 bits (1354), Expect = e-149
Identities = 276/492 (56%), Positives = 358/492 (72%), Gaps = 16/492 (3%)
Query: 1 MGCFHSKTAHLHSPEDPPTALPDSKKTDPGDDGGDGDQESQVPVFKEYGLNELRRATHEF 60
MG SK S + PD + + G+ G V F+EY L +L+ AT F
Sbjct: 1 MGGQSSKIGTCCSHKTTALEAPDVENKENGEVNG-------VHSFREYSLEQLKIATSCF 53
Query: 61 STEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMV 120
+ E +VSE GE APNVVY+GKLEN+ +A+KRFS +WPD +QFL EA VG++R KRM
Sbjct: 54 ALENVVSEHGETAPNVVYQGKLENHMKIAIKRFSGTAWPDPRQFLEEARLVGQLRSKRMA 113
Query: 121 NLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDKQPLPWEMRVRVAYHVAQALDHCSMENR 180
NL+G C EG ERLLVAE+MPN+TL+KHLFHWD +P+ W MR+RVA ++++AL++CS
Sbjct: 114 NLLGYCCEGGERLLVAEFMPNETLAKHLFHWDTEPMKWAMRLRVALYISEALEYCSNNGH 173
Query: 181 KIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGKSYSTNLAYTPPEFLRTGRIIAESVIY 240
+YHDLNAYR+LFDE+ +PRLS+FGLMKNSRDGKSYSTNLA+TPPE+LRTGRI AESVIY
Sbjct: 174 TLYHDLNAYRVLFDEECNPRLSTFGLMKNSRDGKSYSTNLAFTPPEYLRTGRITAESVIY 233
Query: 241 SYGTVLLDLLSGKHIPPSHALDLIRGKNALLLMDSSLEGQYANDDATKLVELASKCLQFE 300
S+GT+LLDLL+GKHIPPSHALDLIR +N L DS LEGQ+++ D T+LV L S CLQ+E
Sbjct: 234 SFGTLLLDLLTGKHIPPSHALDLIRDRNLQTLTDSCLEGQFSDSDGTELVRLTSCCLQYE 293
Query: 301 ARERPDIKFLLTAVTPLQKQKEVASHVLMGLTKTPAVLPLPTMLSPLGKACARMDLTAIH 360
ARERP+IK L+TA+ LQK EV SHVLMGL ++ P SP +AC+ DLT++
Sbjct: 294 ARERPNIKSLVTALISLQKDTEVLSHVLMGLPQSGTFASPP---SPFAEACSGKDLTSMV 350
Query: 361 DILLKTGYKDEEGAENELSFQEWTQQVQDILNTKKFGDIAFRDKDFKNAIEYYSKLVVMM 420
+IL K GYKD+E +LSF WT+Q+Q+ +N+KK GDIAFR KDF AIE+Y++ + +
Sbjct: 351 EILEKIGYKDDE----DLSFM-WTEQMQEAINSKKKGDIAFRRKDFSEAIEFYTQFLDLG 405
Query: 421 SVPSATVFARRAFAYLMNDQAELALRDAMQAQVCIPDWPTAFYLQALALSKLGMETDAQD 480
+ SATV RR+ +YLM++ A+ AL DAM+AQ P W A YLQ+ ALS LGME ++Q
Sbjct: 406 MI-SATVLVRRSQSYLMSNMAKEALDDAMKAQGISPVWYVALYLQSAALSVLGMEKESQI 464
Query: 481 MLNDGAAFEAKR 492
L +G+ EA++
Sbjct: 465 ALTEGSILEARK 476
>At3g09240 putative protein kinase
Length = 477
Score = 505 bits (1301), Expect = e-143
Identities = 254/466 (54%), Positives = 339/466 (72%), Gaps = 9/466 (1%)
Query: 32 DGGDGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVK 91
D + D+ S P F E+ L +LR AT FS + IVSE E+ PN+VYKG+L + R +AVK
Sbjct: 13 DNKEEDEGSTCPNFLEFSLEQLRVATDGFSADNIVSEHNERVPNIVYKGQLNDGRKIAVK 72
Query: 92 RFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHW 151
RF + SWPD+ +F+ EA VG+ R + M NLIGCC+EG ERLLVAEYMPN+TL+KHLFHW
Sbjct: 73 RFQRLSWPDSLEFIEEAQAVGRCRSEHMANLIGCCSEGHERLLVAEYMPNETLAKHLFHW 132
Query: 152 DKQPLPWEMRVRVAYHVAQALDHCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNSR 211
+K+P+ WEMR+RVA H A AL++C+ +YHDLN YRILFD+ G+PRLS FGLMK SR
Sbjct: 133 EKRPMKWEMRLRVALHTATALEYCNDWGIDLYHDLNTYRILFDKVGNPRLSCFGLMKCSR 192
Query: 212 DGKSYSTNLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIRGKNALL 271
+GKSYSTNLA+ PPE+LR G +I ESV +S+GT+LLDL+SG+HIPP+HALDL RGKN L+
Sbjct: 193 EGKSYSTNLAFAPPEYLRLGTVIPESVTFSFGTLLLDLMSGRHIPPNHALDLFRGKNYLV 252
Query: 272 LMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAVTPLQKQKEVASHVLMGL 331
LMDS+L+GQ++++D T+L+ LAS+CL+ E ERP IKFL++A++ L+K+ E+ +V
Sbjct: 253 LMDSALDGQFSDEDRTELIHLASRCLRPEPDERPSIKFLMSALSRLEKRAELWPNVKEEN 312
Query: 332 TKTPAVL------PLPTMLSPLGKACARMDLTAIHDILLKTGY-KDEEGAENELSFQEWT 384
TP+ PLP L+P G+AC R+DL+ +H++L K GY +D+ NE SFQ WT
Sbjct: 313 IPTPSYTEPATKEPLP--LTPFGEACWRVDLSGMHELLEKLGYGEDDVVVTNEFSFQMWT 370
Query: 385 QQVQDILNTKKFGDIAFRDKDFKNAIEYYSKLVVMMSVPSATVFARRAFAYLMNDQAELA 444
Q+Q+ ++ KK GD AFR KDF+ AIE+Y++ + V S TV ARR YLM+D A
Sbjct: 371 GQMQENMDYKKHGDAAFRAKDFETAIEFYTEFMSGAPVVSPTVLARRCLCYLMSDMFREA 430
Query: 445 LRDAMQAQVCIPDWPTAFYLQALALSKLGMETDAQDMLNDGAAFEA 490
L DAMQ QV P++ A YLQA L KLGME +A++ L G++ EA
Sbjct: 431 LSDAMQTQVASPEFSIALYLQAACLLKLGMEAEAKEALRHGSSLEA 476
>At1g63500 protein kinase, putative
Length = 422
Score = 499 bits (1284), Expect = e-141
Identities = 242/383 (63%), Positives = 303/383 (78%), Gaps = 4/383 (1%)
Query: 111 VGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDKQPLPWEMRVRVAYHVAQ 170
VG++R+ RM NL+GCC EG+ERLLVAE+MPN+TL+KHLFHW+ QP+ W MR+RVA H+AQ
Sbjct: 42 VGQLRNYRMANLLGCCYEGEERLLVAEFMPNETLAKHLFHWESQPMKWAMRLRVALHIAQ 101
Query: 171 ALDHCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGKSYSTNLAYTPPEFLRT 230
AL++C+ + R +YHDLNAYR+LFD+D +PRLS FGLMKNSRDGKSYSTNLA+TPPE+LRT
Sbjct: 102 ALEYCTGKGRALYHDLNAYRVLFDDDSNPRLSCFGLMKNSRDGKSYSTNLAFTPPEYLRT 161
Query: 231 GRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIRGKNALLLMDSSLEGQYANDDATKLV 290
GR+ ESV+YSYGT+LLDLLSGKHIPPSHALDLIR +N +L+DS LEGQ+++DD T+L+
Sbjct: 162 GRVTPESVMYSYGTLLLDLLSGKHIPPSHALDLIRDRNIQMLIDSCLEGQFSSDDGTELI 221
Query: 291 ELASKCLQFEARERPDIKFLLTAVTPLQKQKEVASHVLMGLTKTPAVLPLPTMLSPLGKA 350
LAS+CLQ+E RERP+ K L+TA+ PLQK E SH LMG+ + + P LSPLG+A
Sbjct: 222 RLASRCLQYEPRERPNPKSLVTAMIPLQKDLETPSHQLMGIPSSASTTP----LSPLGEA 277
Query: 351 CARMDLTAIHDILLKTGYKDEEGAENELSFQEWTQQVQDILNTKKFGDIAFRDKDFKNAI 410
C R DLTAIH+IL K YKD+EGA ELSFQ WT Q+QD LN KK GD+AFR K+F NAI
Sbjct: 278 CLRTDLTAIHEILEKLSYKDDEGAATELSFQMWTNQMQDSLNFKKKGDVAFRHKEFANAI 337
Query: 411 EYYSKLVVMMSVPSATVFARRAFAYLMNDQAELALRDAMQAQVCIPDWPTAFYLQALALS 470
+ YS+ + ++ S TV+ARR+ YLMN+ + AL DAMQAQV P W A YLQA+ALS
Sbjct: 338 DCYSQFIEGGTMVSPTVYARRSLCYLMNEMPQEALNDAMQAQVISPAWHIASYLQAVALS 397
Query: 471 KLGMETDAQDMLNDGAAFEAKRS 493
LG E +A L DG+ E+KR+
Sbjct: 398 ALGQENEAHAALKDGSMLESKRN 420
>At5g01060 putative protein - kinase
Length = 499
Score = 495 bits (1275), Expect = e-140
Identities = 251/499 (50%), Positives = 338/499 (67%), Gaps = 16/499 (3%)
Query: 3 CFHSKTAHLHSPEDPPTALPDSKKT---DPGDDGGDGDQESQVPVFKEYGLNELRRATHE 59
CF S SP T + D + D DDGG +F+E+ L +LR AT
Sbjct: 5 CFKSWRRSSSSPSITSTVIDDLENVREYDADDDGGHYPL-----IFREFSLEQLRIATDG 59
Query: 60 FSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRM 119
FS IVSE + PN+VYKGKL + R +AVKRF + SWPD +F+ EA VG++R + M
Sbjct: 60 FSAGNIVSEHNDSVPNIVYKGKLGDGRRIAVKRFQRLSWPDPFEFINEAQAVGRLRSEHM 119
Query: 120 VNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDKQPLPWEMRVRVAYHVAQALDHCSMEN 179
NLIGCC + +ERLLVAEYMPN TL+KHLFHW+K+P+ WEMR++VA H A+AL++C+ +
Sbjct: 120 ANLIGCCCDDNERLLVAEYMPNGTLAKHLFHWEKRPMKWEMRLKVALHTARALEYCNDKG 179
Query: 180 RKIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGKSYSTNLAYTPPEFLRTGRIIAESVI 239
+YHDLN YRI+FD+ G P+LS FGLMKNS +GK YSTNLA+ PPE+LR G +IAESV
Sbjct: 180 IDLYHDLNPYRIMFDKTGIPKLSCFGLMKNSHEGKIYSTNLAFAPPEYLRLGTVIAESVT 239
Query: 240 YSYGTVLLDLLSGKHIPPSHALDLIRGKNALLLMDSSLEGQYANDDATKLVELASKCLQF 299
+S+GT+LLDL+SG+HIPP+HALDL RGKN L+LMDS+L+GQ++++D T+L+ +AS+C +
Sbjct: 240 FSFGTLLLDLMSGRHIPPNHALDLFRGKNYLVLMDSALDGQFSDEDRTELIHVASRCFKT 299
Query: 300 EARERPDIKFLLTAVTPLQKQKEVAS-HVLMGLTKTPAVLPLPT-------MLSPLGKAC 351
E ERP IKFL ++ LQK+ ++ +V ++ LP T L+P G AC
Sbjct: 300 EPEERPSIKFLKATLSRLQKRAKLCPINVKRPMSPPSKNLPEKTKPATESLKLTPFGDAC 359
Query: 352 ARMDLTAIHDILLKTGYKDEEGAENELSFQEWTQQVQDILNTKKFGDIAFRDKDFKNAIE 411
+R DL++IH++L K GY+++ G NE SFQ WT ++Q+ ++ KK GD AF KDF AIE
Sbjct: 360 SRADLSSIHELLEKLGYEEDNGVGNEFSFQMWTGEMQENMDYKKHGDAAFLAKDFDTAIE 419
Query: 412 YYSKLVVMMSVPSATVFARRAFAYLMNDQAELALRDAMQAQVCIPDWPTAFYLQALALSK 471
+Y++ + S TV ARR YLM + AL DAMQAQV P+WP YLQA L K
Sbjct: 420 FYTEFMTGAPTVSPTVLARRCLCYLMTEMFSEALSDAMQAQVASPEWPIPLYLQAACLFK 479
Query: 472 LGMETDAQDMLNDGAAFEA 490
L ME +A++ L G+A EA
Sbjct: 480 LEMEAEAKEALRHGSALEA 498
>At1g50990 hypothetical protein
Length = 476
Score = 437 bits (1124), Expect = e-123
Identities = 229/464 (49%), Positives = 311/464 (66%), Gaps = 36/464 (7%)
Query: 34 GDGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENN---RLVAV 90
G G S VP F E+ + LR AT+ F+ +VS ++ PN+VY+G + ++ RL+AV
Sbjct: 46 GRGWHFSDVPDFSEFSASVLRDATNNFNKNAVVSVCSDQEPNLVYQGCIRSDKDKRLIAV 105
Query: 91 KRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFH 150
K+FSK +WPD +QF EA +G +RH R+VNLIG C EGDERLLV+EYMPN++L+KHLFH
Sbjct: 106 KKFSKTTWPDPKQFATEARAIGSLRHVRLVNLIGYCCEGDERLLVSEYMPNESLTKHLFH 165
Query: 151 WDKQPLPWEMRVRVAYHVAQALDHCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNS 210
W+KQ + W MR+RVA +VA+AL++C K+YHDLN R+LFDE+G PRLS FG MKNS
Sbjct: 166 WEKQTMEWAMRLRVALYVAEALEYCRQSGLKLYHDLNTCRVLFDENGSPRLSCFGWMKNS 225
Query: 211 RDGKSYSTNLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIRGKNAL 270
+DGK++STNLAYTPPE+LR +ESV++S+GT LLDLLSGKHIPPSHA+ I+ +N
Sbjct: 226 KDGKNFSTNLAYTPPEYLR-----SESVVFSFGTFLLDLLSGKHIPPSHAVGTIQKQNLN 280
Query: 271 LLMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAVTPLQKQKEVASHVLMG 330
+LMDS LEG Y +DA + +LASKCL ERP+I +++ +T LQ++ +V S+ ++G
Sbjct: 281 VLMDSHLEGNYPEEDAAMVFDLASKCLHNNPNERPEIGDIISVITTLQQKLDVPSYTMLG 340
Query: 331 LTKTPAVLPLPTMLSPLGKACARMDLTAIHDILLKTGYKDEEGAENELSFQEWTQQVQDI 390
++K L + S + AC +MDL A+H IL YK++E ELSFQ+W QQ++D
Sbjct: 341 ISKLEK-LEMEHPKSLIYDACHQMDLAALHQILEAMEYKEDE-VTCELSFQQWAQQIKDF 398
Query: 391 LNTKKFGDIAFRDKDFKNAIEYYSKLVVMMSVPSATVFARRAFAYLMNDQAELALRDAMQ 450
+ ++ +M+ S TV+ARR+ YL DQ + ALRDAMQ
Sbjct: 399 I-----------------------EIGIMI---SPTVYARRSMCYLFCDQPDAALRDAMQ 432
Query: 451 AQVCIPDWPTAFYLQALALSKLGMETDAQDMLNDGAAFEAKRSN 494
AQ DWPTAFYLQA+ALSKL M D+ ML + E KR +
Sbjct: 433 AQCVYSDWPTAFYLQAVALSKLNMVEDSATMLKEALILEDKRGS 476
>At2g17090 putative protein kinase
Length = 465
Score = 400 bits (1028), Expect = e-112
Identities = 216/492 (43%), Positives = 313/492 (62%), Gaps = 27/492 (5%)
Query: 1 MGCFHSKTAHLHSPEDPPTALPDSKKTDPGDDGGDGDQESQVPVFKEYGLNELRRATHEF 60
MGC +S L S DP ++P + GG+ P ++ + L+ AT+ F
Sbjct: 1 MGCCYS----LSSTVDPVQDHTTDASSEPRNGGGED------PPLTKFSFSALKTATNHF 50
Query: 61 STEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMV 120
S E IVS+ + +VV+KG+L+N VA+KRF+ +W D + FL EA VGK+RHKR+V
Sbjct: 51 SPENIVSD---QTSDVVFKGRLQNGGFVAIKRFNNMAWSDPKLFLEEAQRVGKLRHKRLV 107
Query: 121 NLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDKQPLPWEMRVRVAYHVAQALDHCSMENR 180
NLIG C +GD+R LVA++M NDTL+K LF Q + W +R+RVAY VA+ALD+C+
Sbjct: 108 NLIGYCCDGDKRFLVADFMANDTLAKRLFQRKYQTMDWSIRLRVAYFVAEALDYCNTAGF 167
Query: 181 KIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGKSYSTNLAYTPPEFLRTGRIIAESVIY 240
Y++L+AY++LFDEDGD LS FGLMK + + + TG + E+VIY
Sbjct: 168 ASYNNLSAYKVLFDEDGDACLSCFGLMKEINNDQ-------------ITTGSVNPENVIY 214
Query: 241 SYGTVLLDLLSGKHIPPSHALDLIRGKNALLLMDSSLEGQYANDDATKLVELASKCLQFE 300
+GTVL++LLSGK IPPSHA ++I KN LMD L+G+++ D+A + +LAS+CL++E
Sbjct: 215 RFGTVLVNLLSGKQIPPSHAPEMIHRKNVFKLMDPYLKGKFSIDEANVVYKLASQCLKYE 274
Query: 301 ARERPDIKFLLTAVTPLQKQKEVASHVLMGLTKTPAVLPLPTMLSPLGKACARMDLTAIH 360
+E P+ K ++ + LQ + E S+ ++ +T + LSPLG+AC RMDL +IH
Sbjct: 275 GQESPNTKEIVATLETLQTRTEAPSYEVVEMTNQEKDASSSSNLSPLGEACLRMDLASIH 334
Query: 361 DILLKTGYKDEEGAENELSFQEWTQQVQDILNTKKFGDIAFRDKDFKNAIEYYSKLVVMM 420
IL+ GY D++ ELSF+EW Q+V+++ + ++ GD AF ++DFK AI YS+ V
Sbjct: 335 SILVLAGYDDDKDI-IELSFEEWIQEVKELQDVRRNGDRAFVEQDFKTAIACYSQFVEER 393
Query: 421 SVPSATVFARRAFAYLMNDQAELALRDAMQAQVCIPDWPTAFYLQALALSKLGMETDAQD 480
S+ +V+ARR+ +YL D+ E AL D M AQ PDWPTAFYLQ++AL+KL M TD+ D
Sbjct: 394 SLVYPSVYARRSLSYLFCDEPEKALLDGMHAQGVFPDWPTAFYLQSVALAKLDMNTDSAD 453
Query: 481 MLNDGAAFEAKR 492
L + A E K+
Sbjct: 454 TLKEAALLEVKK 465
>At2g17170 hypothetical protein
Length = 328
Score = 233 bits (593), Expect = 2e-61
Identities = 136/328 (41%), Positives = 203/328 (61%), Gaps = 29/328 (8%)
Query: 13 SPEDPPTALPDSKKTDPGDDGGDGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEK 72
+P D P + PG+ DG V E+ L+++ AT+ FS++ I+SE+ E+
Sbjct: 8 TPLDNNQQPPQQPQIPPGERKNDGFVNWPVT---EFFLSDITTATNNFSSDEIISENAEE 64
Query: 73 APNVVYKG-KLENNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDE 131
+ NVVYKG LEN VAVKRF W D F +A G+++HKR+V L+G C E DE
Sbjct: 65 SSNVVYKGCLLENLGFVAVKRFKNTPWDDPDYFTEDAKTAGELKHKRLVKLLGYCCEEDE 124
Query: 132 RLLVAEYMPNDTLSKHLFHWDKQPLPWEMRVRVAYHVAQALDHCSMENRKIYHDLNAYRI 191
LLVAE+MPNDTL+K LF ++ + W MR+RVAYH+ +ALD+C+ Y +L+AY +
Sbjct: 125 GLLVAEFMPNDTLAKRLF--QEKNMEWSMRLRVAYHIVEALDYCNSVRFDEYDNLSAYTV 182
Query: 192 LFDEDGDPRLSSFGLMK-----NSRDGKSYSTNLAYTPPEFLRTGRIIAESVIYSYGTVL 246
LFD+DGD LSSFGL+K N R+G + +RTG +V Y +GT+L
Sbjct: 183 LFDKDGDACLSSFGLVKEIIRYNRREGGN------------VRTG-----NVTYRFGTIL 225
Query: 247 LDLLSGKHIPPSHALDLIRGKNALLLMDSSLEGQYANDDATKLVELASKCLQFEAR-ERP 305
++LL+G I PSHA ++I GK+ LMD +L+G+++ ++AT +++LAS+CLQ++ E
Sbjct: 226 VNLLTGLQISPSHAPEMINGKDVTELMDPNLKGKFSTEEATIVLKLASECLQWKGYIENG 285
Query: 306 DIKFLLTAVTPLQKQKEVASHVLMGLTK 333
K L+ + LQ +KE++S + +TK
Sbjct: 286 ITKKLVATLKALQAKKEISSSEMHEVTK 313
>At1g69790 protein kinase like protein
Length = 387
Score = 164 bits (415), Expect = 1e-40
Identities = 106/325 (32%), Positives = 163/325 (49%), Gaps = 37/325 (11%)
Query: 43 PVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKG----------KLENNRLVAVKR 92
P K + NEL+ AT F ++ E G VYKG K + +VAVK+
Sbjct: 67 PTLKAFTFNELKTATRNFKPNSMIGEGGF---GCVYKGWIGERSLSPSKPGSGMVVAVKK 123
Query: 93 FSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWD 152
+ + +++L E +G++ H +V LIG C EG++RLLV EYMP +L HLF
Sbjct: 124 LKSEGFQGHKEWLTEVHYLGRLHHMNLVKLIGYCLEGEKRLLVYEYMPKGSLENHLFRRG 183
Query: 153 KQPLPWEMRVRVAYHVAQALDHCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNSRD 212
+P+PW+ R++VA+ A+ L E + IY D A IL D D + +LS FGL K
Sbjct: 184 AEPIPWKTRMKVAFSAARGLSFLH-EAKVIYRDFKASNILLDVDFNAKLSDFGLAKAGPT 242
Query: 213 G-KSYST-----NLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALD---- 262
G +++ T Y PE++ TGR+ ++S +YS+G VLL+LLSG+ +
Sbjct: 243 GDRTHVTTQVIGTQGYAAPEYIATGRLTSKSDVYSFGVVLLELLSGRPTLDKSKVGVERN 302
Query: 263 --------LIRGKNALLLMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAV 314
L+ + +MD+ L GQY + A +A +CL E + RPD+ +L+ +
Sbjct: 303 LVDWAIPYLVDRRKVFRIMDTKLGGQYPHKGACAAANIALRCLNTEPKLRPDMADVLSTL 362
Query: 315 TPLQKQKEVASHVLMGLTKTPAVLP 339
L+ S MG T+ + P
Sbjct: 363 QQLE-----TSSKKMGSTQNIVMSP 382
>At2g28930 protein kinase like protein
Length = 412
Score = 161 bits (407), Expect = 9e-40
Identities = 112/330 (33%), Positives = 173/330 (51%), Gaps = 38/330 (11%)
Query: 24 SKKTDPGDDGGDGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLE 83
S +T+P +G + Q P K + EL+ AT F + ++ E G + V+KG ++
Sbjct: 37 SIRTNPRTEG----EILQSPNLKSFTFAELKAATRNFRPDSVLGEGGFGS---VFKGWID 89
Query: 84 NNRL----------VAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERL 133
L +AVK+ ++ W Q++LAE +G+ H +V LIG C E + RL
Sbjct: 90 EQTLTASKPGTGVVIAVKKLNQDGWQGHQEWLAEVNYLGQFSHPNLVKLIGYCLEDEHRL 149
Query: 134 LVAEYMPNDTLSKHLFHWDK--QPLPWEMRVRVAYHVAQALDHC-SMENRKIYHDLNAYR 190
LV E+MP +L HLF QPL W +R++VA A+ L + E IY D
Sbjct: 150 LVYEFMPRGSLENHLFRRGSYFQPLSWTLRLKVALGAAKGLAFLHNAETSVIYRDFKTSN 209
Query: 191 ILFDEDGDPRLSSFGLMKNSRDG-KSY-STNL----AYTPPEFLRTGRIIAESVIYSYGT 244
IL D + + +LS FGL K+ G KS+ ST + Y PE+L TG + +S +YSYG
Sbjct: 210 ILLDSEYNAKLSDFGLAKDGPTGDKSHVSTRIMGTYGYAAPEYLATGHLTTKSDVYSYGV 269
Query: 245 VLLDLLSG-----KHIPPSH------ALDLIRGKNALL-LMDSSLEGQYANDDATKLVEL 292
VLL++LSG K+ PP A L+ K L ++D+ L+ QY+ ++A K+ L
Sbjct: 270 VLLEVLSGRRAVDKNRPPGEQKLVEWARPLLANKRKLFRVIDNRLQDQYSMEEACKVATL 329
Query: 293 ASKCLQFEARERPDIKFLLTAVTPLQKQKE 322
A +CL FE + RP++ +++ + +Q E
Sbjct: 330 ALRCLTFEIKLRPNMNEVVSHLEHIQTLNE 359
>At1g26970 protein kinase like protein
Length = 412
Score = 158 bits (400), Expect = 6e-39
Identities = 104/305 (34%), Positives = 155/305 (50%), Gaps = 33/305 (10%)
Query: 43 PVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRL----------VAVKR 92
P K + NEL+ AT F + ++ E G VYKG ++ L VAVK+
Sbjct: 66 PTLKAFTFNELKTATRNFRPDSVIGEGGF---GYVYKGWIDERTLSPSKPGSGMVVAVKK 122
Query: 93 FSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDE-RLLVAEYMPNDTLSKHLFHW 151
++ + +Q+LAE +G++ H +V LIG C++GD RLLV EYMP +L HLF
Sbjct: 123 LKEEGFQGHRQWLAEVDCLGRLHHMNLVKLIGYCSKGDHIRLLVYEYMPKGSLENHLFRR 182
Query: 152 DKQPLPWEMRVRVAYHVAQALDHCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMK--N 209
+P+PW R++VA A+ L E + IY D A IL D + + +LS FGL K
Sbjct: 183 GAEPIPWRTRIKVAIGAARGLAFLH-EAQVIYRDFKASNILLDSEFNAKLSDFGLAKVGP 241
Query: 210 SRDGKSYSTNL----AYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIR 265
+ D ST + Y PE++ TGRI A+S +YS+G VLL+LLSG+ + + R
Sbjct: 242 TGDRTHVSTQVMGTQGYAAPEYVATGRITAKSDVYSFGVVLLELLSGRLTVDKTKVGVER 301
Query: 266 G------------KNALLLMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTA 313
+ +MD+ L GQY + A A +CL E + RP + +L+
Sbjct: 302 NLVDWAIPYLGDKRKVFRIMDTKLGGQYPHKGACLTANTALQCLNQEPKLRPKMSDVLST 361
Query: 314 VTPLQ 318
+ L+
Sbjct: 362 LEELE 366
>At1g07570 protein kinase APK1A
Length = 410
Score = 155 bits (393), Expect = 4e-38
Identities = 104/309 (33%), Positives = 163/309 (52%), Gaps = 34/309 (11%)
Query: 41 QVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRL----------VAV 90
Q P K + EL+ AT F + ++ E G V+KG ++ L +AV
Sbjct: 49 QSPNLKSFSFAELKSATRNFRPDSVLGEGGF---GCVFKGWIDEKSLTASRPGTGLVIAV 105
Query: 91 KRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFH 150
K+ ++ W Q++LAE +G+ H+ +V LIG C E + RLLV E+MP +L HLF
Sbjct: 106 KKLNQDGWQGHQEWLAEVNYLGQFSHRHLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFR 165
Query: 151 WDK--QPLPWEMRVRVAYHVAQALDHC-SMENRKIYHDLNAYRILFDEDGDPRLSSFGLM 207
QPL W++R++VA A+ L S E R IY D IL D + + +LS FGL
Sbjct: 166 RGLYFQPLSWKLRLKVALGAAKGLAFLHSSETRVIYRDFKTSNILLDSEYNAKLSDFGLA 225
Query: 208 KNSRDG-KSY-STNL----AYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHI----PP 257
K+ G KS+ ST + Y PE+L TG + +S +YS+G VLL+LLSG+ P
Sbjct: 226 KDGPIGDKSHVSTRVMGTHGYAAPEYLATGHLTTKSDVYSFGVVLLELLSGRRAVDKNRP 285
Query: 258 SHALDLIRGKNALL--------LMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKF 309
S +L+ L ++D+ L+ QY+ ++A K+ L+ +CL E + RP++
Sbjct: 286 SGERNLVEWAKPYLVNKRKIFRVIDNRLQDQYSMEEACKVATLSLRCLTTEIKLRPNMSE 345
Query: 310 LLTAVTPLQ 318
+++ + +Q
Sbjct: 346 VVSHLEHIQ 354
>At3g07070 putative protein kinase
Length = 414
Score = 155 bits (391), Expect = 6e-38
Identities = 108/324 (33%), Positives = 160/324 (49%), Gaps = 32/324 (9%)
Query: 14 PEDPPTALPDSKKTDPGDDGGDGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKA 73
PE+P T +K D D + + + + + EL AT F E ++ E G
Sbjct: 39 PENPKTVNEQNKNNDE-----DKEVTNNIAA-QTFSFRELATATKNFRQECLIGEGGFGR 92
Query: 74 PNVVYKGKLENN-RLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDER 132
VYKGKLE +VAVK+ + ++F+ E + + HK +VNLIG CA+GD+R
Sbjct: 93 ---VYKGKLEKTGMIVAVKQLDRNGLQGNKEFIVEVLMLSLLHHKHLVNLIGYCADGDQR 149
Query: 133 LLVAEYMPNDTLSKHLFHW--DKQPLPWEMRVRVAYHVAQALD--HCSMENRKIYHDLNA 188
LLV EYM +L HL D+ PL W+ R+R+A A L+ H IY DL A
Sbjct: 150 LLVYEYMSRGSLEDHLLDLTPDQIPLDWDTRIRIALGAAMGLEYLHDKANPPVIYRDLKA 209
Query: 189 YRILFDEDGDPRLSSFGLMKNSRDGKSYSTN------LAYTPPEFLRTGRIIAESVIYSY 242
IL D + + +LS FGL K G + Y PE+ RTG++ +S +YS+
Sbjct: 210 ANILLDGEFNAKLSDFGLAKLGPVGDKQHVSSRVMGTYGYCAPEYQRTGQLTTKSDVYSF 269
Query: 243 GTVLLDLLSGKHI----PPSHALDLIRGKNALL--------LMDSSLEGQYANDDATKLV 290
G VLL+L++G+ + P +L+ + L D SLEG + + V
Sbjct: 270 GVVLLELITGRRVIDTTRPKDEQNLVTWAQPVFKEPSRFPELADPSLEGVFPEKALNQAV 329
Query: 291 ELASKCLQFEARERPDIKFLLTAV 314
+A+ CLQ EA RP + ++TA+
Sbjct: 330 AVAAMCLQEEATVRPLMSDVVTAL 353
>At2g39660 putative protein kinase
Length = 395
Score = 155 bits (391), Expect = 6e-38
Identities = 105/315 (33%), Positives = 169/315 (53%), Gaps = 35/315 (11%)
Query: 35 DGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRL------- 87
+G+ S PV K + NEL+ AT F + ++ E G V+KG L+ + L
Sbjct: 43 EGEILSSTPV-KSFTFNELKLATRNFRPDSVIGEGGF---GCVFKGWLDESTLTPTKPGT 98
Query: 88 ---VAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTL 144
+AVK+ +++ + +++L E +G++ H +V LIG C E + RLLV E+M +L
Sbjct: 99 GLVIAVKKLNQEGFQGHREWLTEINYLGQLSHPNLVKLIGYCLEDEHRLLVYEFMQKGSL 158
Query: 145 SKHLFHWDK--QPLPWEMRVRVAYHVAQALDHCSMENRK-IYHDLNAYRILFDEDGDPRL 201
HLF +PLPW +RV VA A+ L + K IY D+ A IL D D + +L
Sbjct: 159 ENHLFRRGAYFKPLPWFLRVNVALDAAKGLAFLHSDPVKVIYRDIKASNILLDADYNAKL 218
Query: 202 SSFGLMKNSRDGK-SY-STNL----AYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGK-- 253
S FGL ++ G SY ST + Y PE++ +G + A S +YS+G +LL++LSGK
Sbjct: 219 SDFGLARDGPMGDLSYVSTRVMGTYGYAAPEYMSSGHLNARSDVYSFGVLLLEILSGKRA 278
Query: 254 --HIPPSHALDLI--------RGKNALLLMDSSLEGQYANDDATKLVELASKCLQFEARE 303
H P+ +L+ + LL++D+ L+ QY ++A ++ +A +CL FE +
Sbjct: 279 LDHNRPAKEENLVDWARPYLTSKRKVLLIVDNRLDTQYLPEEAVRMASVAVQCLSFEPKS 338
Query: 304 RPDIKFLLTAVTPLQ 318
RP + ++ A+ LQ
Sbjct: 339 RPTMDQVVRALQQLQ 353
>At3g55450 serine/threonine-specific protein kinase -like
Length = 389
Score = 154 bits (388), Expect = 1e-37
Identities = 101/305 (33%), Positives = 160/305 (52%), Gaps = 35/305 (11%)
Query: 46 KEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRL----------VAVKRFSK 95
K + NEL+ AT F ++ +V E G V++G L+ L +AVKR +
Sbjct: 47 KSFSFNELKLATRNFRSDSVVGEGGF---GCVFRGWLDETTLTPTKSSSGLVIAVKRLNP 103
Query: 96 QSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLF---HWD 152
+ +++L E +G++ H +V LIG C E ++RLLV E+M +L HLF + D
Sbjct: 104 DGFQGHREWLTEINYLGQLSHPNLVKLIGYCLEDEQRLLVYEFMHKGSLENHLFANGNKD 163
Query: 153 KQPLPWEMRVRVAYHVAQALDHCSMENRK-IYHDLNAYRILFDEDGDPRLSSFGLMKNSR 211
+PL W +R++VA A+ L + K IY D+ A IL D D + +LS FGL ++
Sbjct: 164 FKPLSWILRIKVALDAAKGLAFLHSDPVKVIYRDIKASNILLDSDFNAKLSDFGLARDGP 223
Query: 212 DG-KSYST-----NLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGK----HIPPSHAL 261
G +SY + Y PE++ TG + A S +YS+G VLL+LL G+ H P+
Sbjct: 224 MGEQSYVSTRVMGTFGYAAPEYVSTGHLNARSDVYSFGVVLLELLCGRQALDHNRPAKEQ 283
Query: 262 DLI--------RGKNALLLMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTA 313
+L+ + LL++D+ L QY + A +L +A +CL FE + RP + ++ A
Sbjct: 284 NLVDWARPYLTSRRKVLLIVDTRLNSQYKPEGAVRLASIAVQCLSFEPKSRPTMDQVVRA 343
Query: 314 VTPLQ 318
+ LQ
Sbjct: 344 LVQLQ 348
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.134 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,266,965
Number of Sequences: 26719
Number of extensions: 482161
Number of successful extensions: 3656
Number of sequences better than 10.0: 757
Number of HSP's better than 10.0 without gapping: 626
Number of HSP's successfully gapped in prelim test: 131
Number of HSP's that attempted gapping in prelim test: 1398
Number of HSP's gapped (non-prelim): 828
length of query: 498
length of database: 11,318,596
effective HSP length: 103
effective length of query: 395
effective length of database: 8,566,539
effective search space: 3383782905
effective search space used: 3383782905
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC147404.5


