

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147404.2 + phase: 0
(599 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
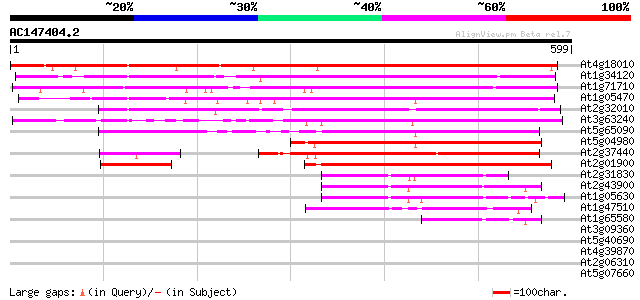
Score E
Sequences producing significant alignments: (bits) Value
At4g18010 putative inositol polyphosphate 5-phosphatase At5P2 608 e-174
At1g34120 putative inositol polyphosphate 5-phosphatase At5P1 416 e-116
At1g71710 RIBOSOMAL PROTEIN, putative (At1g71710) 389 e-108
At1g05470 unknown protein 365 e-101
At2g32010 putative inositol polyphosphate 5'-phosphatase 345 3e-95
At3g63240 inositol-1,4,5-trisphosphate 5-Phosphatase-like protein 315 5e-86
At5g65090 inositol-1, 4, 5-trisphosphate 5-phosphatase-like protein 310 1e-84
At5g04980 putative protein 267 1e-71
At2g37440 unknown protein 256 3e-68
At2g01900 putative inositol polyphosphate-5-phosphatase 251 6e-67
At2g31830 putative inositol polyphosphate 5'-phosphatase 130 3e-30
At2g43900 putative inositol polyphosphate 5'-phosphatase 129 5e-30
At1g05630 putative inositol 1,4,5-trisphosphate 5-phosphatase 126 3e-29
At1g47510 hypothetical protein 90 4e-18
At1g65580 hypothetical protein 83 4e-16
At3g09360 putative transcription factor 32 1.3
At5g40690 unknown protein 31 1.7
At4g39870 unknown protein 31 1.7
At2g06310 putative Athila retroelement ORF1 protein 31 1.7
At5g07660 SMC-like protein 31 2.2
>At4g18010 putative inositol polyphosphate 5-phosphatase At5P2
Length = 646
Score = 608 bits (1569), Expect = e-174
Identities = 334/642 (52%), Positives = 431/642 (67%), Gaps = 61/642 (9%)
Query: 2 KGKRSEVFWPSTVMKKWLNVKQKVYDFSEDEANTETDESEDDV---------KLIEIETN 52
+GKR E FWPS VM KWLN K KVYDFSEDE +TE ESEDDV + E +
Sbjct: 5 RGKRPERFWPSIVMNKWLNRKPKVYDFSEDEIDTEP-ESEDDVCSVKDVPNVHCVTDEDS 63
Query: 53 HTSKPQANTTKPTQLN---TSCKDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWNVAGK 109
H + + ++ S + ++HRR KSETLRAQYINTK+++VT+ TWNVAGK
Sbjct: 64 HNGRRGSEADHGNNISDGGVSVRGGYQRKHRRGKSETLRAQYINTKDIKVTVATWNVAGK 123
Query: 110 HPCNDLEIEGWLCTEEEPSDIYIIGFQEVVPLNAGNVFGAEDSKPIPKWDALIRRTLNKS 169
P +DLEIE WL T+ PSDIYIIGFQEVVPLNAGNVFGAED PIPKW+++IRRTLNKS
Sbjct: 124 RPSDDLEIEDWLSTDN-PSDIYIIGFQEVVPLNAGNVFGAEDRGPIPKWESIIRRTLNKS 182
Query: 170 SEPGTKK-----------KSNSAPPSPI--RRISSFNTQINPLDSALDKKEEIKTIISIE 216
++ +S+SAP SPI + +S + + + D ++ T I+
Sbjct: 183 NKESVYDQSPSCNNNALHRSHSAPSSPILAQEANSIISHVMVENLVADHSLDLATNEFID 242
Query: 217 KNLQLSKIYDIDLQTILDWPELRLDPIHH-VDSSPKMRRVQSTS-----------DSASL 264
L + + +DWPEL LD V S K+RRV S++ AS
Sbjct: 243 AATALPSL-EPQRNPNMDWPELALDSNPQIVGSEGKLRRVFSSNATLGFKLPENPSGASR 301
Query: 265 YGFEMKSLHQSSG--NFSLLWSEKQQEIVPQVFDSHL--DVSDMLSDEDNDTFSELANNE 320
+ E + L +S +L W++ ++EI + S + + ++ D+ +D S + E
Sbjct: 302 FASEARQLKRSRSFETLNLSWNDIKEEIDNRSSSSSEAEEAAKIMHDDSSDGDSSSQDEE 361
Query: 321 DANGIIS--------------VKSHPKYVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQ 366
D + I + VK KYVRIVSKQMVGIYVSVW++R+LRRHV++LKVS
Sbjct: 362 DGDKIRNSYGLPEDLVEECRKVKDSQKYVRIVSKQMVGIYVSVWIRRRLRRHVNNLKVSP 421
Query: 367 VGVGLMGYMGNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDVHEILQRTRFSS 426
VGVGLMGYMGNKGSVS+SM+++QSRMCFVCSHL SG KDGAEQRRN+DV+EI++RTRF+S
Sbjct: 422 VGVGLMGYMGNKGSVSISMTLYQSRMCFVCSHLTSGHKDGAEQRRNADVYEIIRRTRFAS 481
Query: 427 VFDTDQPQKIPSHDKIFWFGDLNYRINMSDGEIRKLVDLKKWNELMKFDQLSNELCKGHV 486
V DTDQP+ IP HD++FWFGDLNYR+NMSDGE+RKLV K+W+EL DQL EL +GHV
Sbjct: 482 VLDTDQPRTIPCHDQVFWFGDLNYRLNMSDGEVRKLVSQKRWDELKNSDQLIRELRRGHV 541
Query: 487 FEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAE 546
F+GW+EG I FPPTYKYEF+SD++ G N +E EK+RAPAWCDRILWLGKGI+Q Y+ +E
Sbjct: 542 FDGWREGPIKFPPTYKYEFDSDRYAGENLREPEKKRAPAWCDRILWLGKGIRQECYKRSE 601
Query: 547 NQLSDHRPVSSIFLVDVEVIDHRKLERAIYF---ASAVVHPD 585
++SDHRPV+SIF V VEV DHRKL+RA++ A++ VHP+
Sbjct: 602 IRMSDHRPVTSIFNVGVEVFDHRKLQRALHVNNAAASAVHPE 643
>At1g34120 putative inositol polyphosphate 5-phosphatase At5P1
Length = 586
Score = 416 bits (1070), Expect = e-116
Identities = 234/587 (39%), Positives = 347/587 (58%), Gaps = 43/587 (7%)
Query: 7 EVFWPSTVMKKWLNVKQKVYDFSEDEANTETDESEDDVKLIEIETNHTSKPQANTTKPTQ 66
+++W V++KW NV D+S D + D S++ + TN P+ +T +
Sbjct: 27 QLYWARIVLRKWFNVSASESDYSADSDDDYEDRSQEFDPISSGVTN----PRVDTDDRPK 82
Query: 67 LNTSCKDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGWLCTEEE 126
L RRR SET R QYI+TK +R+ GTWNV G+ P +DL+I+GWL T E
Sbjct: 83 L------------RRRNSETFRMQYIDTKAIRICAGTWNVGGRVPSSDLDIDGWLDTLE- 129
Query: 127 PSDIYIIGFQEVVPLNAGNVFGAEDSKPIPKWDALIRRTLNKSSEPGTKKKSNSAPPSPI 186
P+DIY++G QE+VPLNAGN+FG ED +P +W+ LIR LN+ K KS+S PPSP
Sbjct: 130 PADIYVLGLQEIVPLNAGNIFGMEDDQPALEWENLIRDALNRVQPRKLKIKSHSDPPSPS 189
Query: 187 RRISSFNTQINPLDSALDKKEEIKTIISIEKNLQLSKIYDIDLQTILDWPELRLDPIHHV 246
+ + D ++ + IS N +L+ + D+ PI
Sbjct: 190 KFKQPEEVPYSVEDMFVETSHDACDGISSMDN-KLNSVESTDV------------PIVSE 236
Query: 247 DSSPKMRRVQSTSDSASLYG--------FEMKSLHQSSGNFSLLWSEKQQEIVPQVFDSH 298
DS + + ST+D+AS F + S + + K++E
Sbjct: 237 DSLTNIDVLGSTNDNASCLPIQEYLQRQFSTPNTPDRSLSMQINSDSKREERFSYTERVG 296
Query: 299 LDVSDMLSDEDNDTFSELANN-EDANGIISVKSHPKYVRIVSKQMVGIYVSVWVQRKLRR 357
L + N SE + + N I+ P YVRIVSKQMVG+++++WV+R LR+
Sbjct: 297 LSWPEPPLRLLNQYVSERRGSFKSVNLTITNLRKPSYVRIVSKQMVGVFLTIWVRRNLRK 356
Query: 358 HVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDVHE 417
H+ +L VS VGVG+MGY+GNKGSVSVSMS++Q+ CF+C+HL+SG+KD +++RN DV E
Sbjct: 357 HISNLCVSTVGVGIMGYIGNKGSVSVSMSIYQTPFCFLCTHLSSGEKDTDQEKRNDDVRE 416
Query: 418 ILQRTRF--SSVFDTDQPQKIPSHDKIFWFGDLNYRINMSDGEIRKLVDLKKWNELMKFD 475
I +RT+F S+ + P+ I +H++I W GDLNYRIN+S + +L+ K+W L+++D
Sbjct: 417 IHRRTQFLPHSLNANELPRSICNHERIIWLGDLNYRINLSYEKTHELIARKEWQRLVEYD 476
Query: 476 QLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRAPAWCDRILWLGK 535
QLS E+ KG++FEGW EG ++F PTYKYE +S+ ++G + + G++R PAWCDRI+W GK
Sbjct: 477 QLSREMTKGNLFEGWSEGTLDFAPTYKYEIDSENYIGDDPESGKRR--PAWCDRIIWNGK 534
Query: 536 GIKQLKYQSAENQLSDHRPVSSIFLVDVEVIDHRKLERAIYFASAVV 582
G+K Y+ E +LSDHRPV++ FL +VEV+ RKL+ A+ A +
Sbjct: 535 GMKLFNYRRNEIKLSDHRPVTATFLAEVEVLSPRKLQHALTLTYAEI 581
>At1g71710 RIBOSOMAL PROTEIN, putative (At1g71710)
Length = 664
Score = 389 bits (998), Expect = e-108
Identities = 231/660 (35%), Positives = 349/660 (52%), Gaps = 96/660 (14%)
Query: 4 KRSEVFWPSTVMKKWLNVKQKVYDFSED--------EANTETDESEDDVKLIEIETNHTS 55
++S+ W VM+KWLN+ + ++ D +A + D+S D + S
Sbjct: 16 QKSQRLWAKLVMRKWLNISGRDPEYGADTDNESENEDAREDNDDSSSDEEGGSGSRGRES 75
Query: 56 KPQANTTKPTQLNTSCKDYEMK-----------RHRRRKSETLRAQYINTKEVRVTIGTW 104
K N ++ D + RRR SETLRAQYIN KE+RV +GTW
Sbjct: 76 KVYENAEDAIAAASAVVDAAAAAAEFISNDAPMKLRRRNSETLRAQYINNKEIRVCVGTW 135
Query: 105 NVAGKHPCNDLEIEGWLCTEEEPSDIYIIGFQEVVPLNAGNVFGAEDSKPIPKWDALIRR 164
NV G P +DL+I+ W+ +P+DIY++G QE+VPLNAGN+ GAED +P+ KW+ +IR
Sbjct: 136 NVGGISPPSDLDIDDWI-EINQPADIYVLGLQEIVPLNAGNILGAEDDRPVAKWEEVIRE 194
Query: 165 TLNKSSEPGTKKKSNSAPPSPIR------------------RISSFNTQINPLDSALDKK 206
LN+ + KS S PPSP R S +I+P+D +++
Sbjct: 195 ALNRVRPKLSGVKSYSDPPSPGRFKPFEETHDIIEEEVAFESDSDAGVEIHPIDEEEEEE 254
Query: 207 E--------------EIKTIIS-------IEKNLQLSKIYDIDLQTILDWPELRLDPIHH 245
E+KT++ +E Q S +D Q L R D
Sbjct: 255 TDRLWALKHDGGVIGEVKTLVDPNTGLPVVEIKRQFSIPKKLDRQLCL-----RADSFKG 309
Query: 246 VDSSPKMRRVQSTSDSASLYGFEMKSLHQSSGNFSLLWSEKQQEIV-PQVFDSHLDVSDM 304
+ S DS + + L W E ++ P V D + +
Sbjct: 310 I----------SDDDSTQTGMKTINRMLSGKERIGLSWPEPPLNMLGPCVLDRQPSIKTV 359
Query: 305 LSDEDNDTF-------SELANNE------------DANGIISVKSHPKYVRIVSKQMVGI 345
S + +F S NN D ++ K P YVR+VSKQMVGI
Sbjct: 360 KSLKTAKSFKAYSSFKSVAGNNNGIPPEVLALAEMDLKLLMERKRRPAYVRLVSKQMVGI 419
Query: 346 YVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMCFVCSHLASGQKD 405
+++WV+R LR+H+ +++VS VGVG+MGY+GNKG+VSVSMS+ Q+ CF+ +HL +G+++
Sbjct: 420 LLTIWVKRSLRKHIQNVRVSTVGVGVMGYIGNKGAVSVSMSINQTFFCFINTHLTAGERE 479
Query: 406 GAEQRRNSDVHEILQRTRFSSVFDTDQPQKIPSHDKIFWFGDLNYRINMSDGEIRKLVDL 465
+ +RN+DVHEI +RT F SV P+ I H++I W GDLNYR++ S + R L+
Sbjct: 480 VDQIKRNADVHEIHKRTVFHSVSALGLPKLIYDHERIIWLGDLNYRLSSSYEKTRDLISK 539
Query: 466 KKWNELMKFDQLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRAPA 525
++W++L+++DQL E KG F+GW EG ++FPPTYKY+ NSD++ + + +R PA
Sbjct: 540 REWSKLLEYDQLVKEYRKGRAFDGWSEGTLHFPPTYKYQANSDEYTANDGK--APKRTPA 597
Query: 526 WCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVIDHRKLERAIYFASAVVHPD 585
WCDR+L GKG++ + Y+ E + SDHRPV++I++ +VEV RKL+RA+ F A + +
Sbjct: 598 WCDRVLSYGKGMRLVHYRRTEQKFSDHRPVTAIYMAEVEVFSARKLQRALTFTDAEIEDE 657
>At1g05470 unknown protein
Length = 585
Score = 365 bits (938), Expect = e-101
Identities = 216/598 (36%), Positives = 311/598 (51%), Gaps = 61/598 (10%)
Query: 10 WPSTVMKKWLNVKQKVYDFSEDEANTETDESEDDVKLIEIETNHTSKPQANTTKPTQLNT 69
W +++KW N+K K +F D+ P+
Sbjct: 5 WSKKMVRKWFNIKSKTEEFQADD-------------------------------PSSAEK 33
Query: 70 SCKDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGWLCTEEEPSD 129
K++E + +RR + + I+ + + + TWNVAG+ P +DL ++ WL + P+D
Sbjct: 34 LSKNWEQQARQRRMNYE-NPRIIDVQNYSIFVATWNVAGRSPPSDLNLDEWLHSSA-PAD 91
Query: 130 IYIIGFQEVVPLNAGNVFGAEDSKPIPKWDALIRRTLNKSSEPGTKKKSNSAPPSPI--- 186
IY++GFQE+VPLNAGNV GAED+ P KW +LIR+TLN + PGT S PSPI
Sbjct: 92 IYVLGFQEIVPLNAGNVLGAEDNGPAQKWLSLIRKTLN--NRPGTSGTSGYHTPSPIPVP 149
Query: 187 --RRISSFNTQINPLDSALDKKEEIKTIISIEKNLQ-----LSKIYDIDLQTILDWPELR 239
+ F+ +S + +T S + L + + + +
Sbjct: 150 MAELDADFSGSTRQKNSTFFHRRSFQTPSSTWNDPSIPQPGLDRRFSVCDRVFFSHRPSD 209
Query: 240 LDPIHHVDSSPKM-----RRVQSTSDSASLYG-----FEMKSLHQSSGNFSL---LWSEK 286
DP SS RR S S Y + + ++S +S
Sbjct: 210 FDPSFRGSSSSHRPSDYSRRPSDYSRRPSDYSRRPSDYSRRPSDSRPSDYSRPSDYYSRP 269
Query: 287 QQEIVPQVFDSHLDVSDMLSDEDNDTFSELANNEDANGIISVKSHPKYVRIVSKQMVGIY 346
P F D + L D + + + NG + +Y + SKQMVG++
Sbjct: 270 SDYSRPSDFSRSSDDDNGLGDSPSTVLYSPGSAANENGYRIPWNSSQYCLVASKQMVGVF 329
Query: 347 VSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMCFVCSHLASGQKDG 406
+++WV+ +LR HV ++KVS VG GLMGY+GNKGS+S+SM + Q+ CFVC+HL SGQK+G
Sbjct: 330 LTIWVKSELREHVKNMKVSCVGRGLMGYLGNKGSISISMLLHQTSFCFVCTHLTSGQKEG 389
Query: 407 AEQRRNSDVHEILQRTRFSSVFDTDQ---PQKIPSHDKIFWFGDLNYRINMSDGEIRKLV 463
E +RNSDV EIL++TRF V +++ P+ I HD++ W GDLNYRI +S + LV
Sbjct: 390 DELKRNSDVMEILKKTRFPRVKSSEEEKSPENILQHDRVIWLGDLNYRIALSYRSAKALV 449
Query: 464 DLKKWNELMKFDQLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRA 523
+++ W L++ DQL E +GHVF+GW EG I FPPTYKY NSD++ G + EKRR
Sbjct: 450 EMQNWRALLENDQLRIEQKRGHVFKGWNEGKIYFPPTYKYSRNSDRYSGDDLHPKEKRRT 509
Query: 524 PAWCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVIDHRKLERAIYFASAV 581
PAWCDRILW G+G+ QL Y E++ SDHRPV IF +VE +R Y AS V
Sbjct: 510 PAWCDRILWFGEGLHQLSYVRGESRFSDHRPVYGIFCAEVESAHNRIKRTTSYSASRV 567
>At2g32010 putative inositol polyphosphate 5'-phosphatase
Length = 501
Score = 345 bits (886), Expect = 3e-95
Identities = 200/504 (39%), Positives = 283/504 (55%), Gaps = 31/504 (6%)
Query: 96 EVRVTIGTWNVAGKHPCNDLEIEGWLCTEEEPSDIYIIGFQEVVPLNAGNVFGAEDSKPI 155
E + + TWNVAG+ P DL ++ WL + P+DIY++GFQE+VPLNAGNV GAED+ P
Sbjct: 6 ENSIFVATWNVAGRSPPEDLNLDEWLHSSA-PADIYVLGFQEIVPLNAGNVLGAEDNGPA 64
Query: 156 PKWDALIRRTLNKSSEPGTKKKSNSAPPSPIRRISS-FNTQINPLDSALDKKEEIKT--I 212
KW +LIR+TLN + + S P PI I + F+ + + +T +
Sbjct: 65 KKWHSLIRKTLNNLPGASSACHTPSPIPVPIAEIDADFSGSSRQKNETFFNRRSFQTPSV 124
Query: 213 ISIEKN------LQLSKIYDIDLQTILDWPELRLDPIHHVDSSPK--MRRVQSTSDSASL 264
S+E+N +L + + + + DP P RR S +
Sbjct: 125 WSMEENDPSISQPRLDRRFSVCDRVFFSHRPSDFDPSFRCGHRPSDYSRRPSDYSRPSDY 184
Query: 265 YGFEMKSLHQSSGNFSLLWSEKQQEIVPQVFDSHLDVSDMLSDEDNDTFSELANNEDANG 324
Y + + S + W + P D S N S L + + NG
Sbjct: 185 YS---RPSNYSRPSDVSRWGSSDDDNGP---------GDSPSTFLNSPGSFLGSAANENG 232
Query: 325 IISVKSHPKYVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSVS 384
+ + +Y + SKQMVGI++++WV+ +LR HV ++KVS VG GLMGY+GNKGS+S+S
Sbjct: 233 YRTPWNSSQYCLVASKQMVGIFLTIWVKSELREHVKNMKVSCVGRGLMGYLGNKGSISIS 292
Query: 385 MSVFQSRMCFVCSHLASGQKDGAEQRRNSDVHEILQRTRFSSVFDTDQPQKIPSHDKIFW 444
M + Q+ CFVC+HL SGQK+G E RRNSDV EIL++TRF V Q +++ W
Sbjct: 293 MLLHQTSFCFVCTHLTSGQKEGDELRRNSDVMEILKKTRFPRV------QSSADENRVIW 346
Query: 445 FGDLNYRINMSDGEIRKLVDLKKWNELMKFDQLSNELCKGHVFEGWKEGLINFPPTYKYE 504
GDLNYRI +S + LV+++ W L++ DQL E +GHVF+GW EG I FPPTYKY
Sbjct: 347 LGDLNYRIALSYRSAKALVEMQNWRALLENDQLRIEQKRGHVFKGWNEGKIYFPPTYKYS 406
Query: 505 FNSDKHVGGNTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVE 564
NSD++ GG+ EKRR PAWCDRILW G+G+ QL Y E++ SDHRPV IF +VE
Sbjct: 407 NNSDRYAGGDLHPKEKRRTPAWCDRILWHGEGLHQLSYVRGESRFSDHRPVYGIFSAEVE 466
Query: 565 VIDHRKLERAIYFASAVVHPDVFL 588
+H++ +R ++A V + L
Sbjct: 467 -SNHKRSKRTNSHSTARVEAEELL 489
>At3g63240 inositol-1,4,5-trisphosphate 5-Phosphatase-like protein
Length = 574
Score = 315 bits (807), Expect = 5e-86
Identities = 217/617 (35%), Positives = 300/617 (48%), Gaps = 116/617 (18%)
Query: 4 KRSEVFWPSTVMKKWLNVKQKVYDFSEDEANTETDESEDDVKLIEIETNHTSKPQANTTK 63
K+S++ WP T++KKWLN+K K DF D+ + + +IE E +A + +
Sbjct: 7 KKSKLSWPKTLVKKWLNIKSKSEDFHADDLDRGEGGGDWRNNVIERE-------EACSVR 59
Query: 64 PTQLNTSCKDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGWLCT 123
++ T K + R + ++ RV TWNVAGK P + L ++ WL T
Sbjct: 60 KSKTETRSKRNSGRARRNKLDVDPPLDHL-----RVFTATWNVAGKSPPSYLNLDDWLHT 114
Query: 124 EEEPSDIYIIGFQEVVPLNAGNVFGAEDSKPIPKWDALIRRTLNKSSEPGTKKKSNSAPP 183
PSDIY V G ++ P LN + GT+
Sbjct: 115 SP-PSDIY--------------VLGFQEIVP-----------LNAGNVLGTEDNG----- 143
Query: 184 SPIRR-ISSFNTQINPLDSALDKKEEIKTIISIEKNLQLSKIYDIDLQTILDWPELRLDP 242
P R+ +S +N L QT P P
Sbjct: 144 -PARKWVSLIRRTLNSLPGG-------------------------SCQTPSPVPH----P 173
Query: 243 IHHVDSSPKMRRVQSTSDSASLYGFEMKSLHQSSGNFSLLWSEKQQEIVPQVFDSHLDVS 302
+ +DS + S + + SL+ +S+ + SL PQ FD V
Sbjct: 174 VAELDSDFEG---DSAAGANSLFYHRSRSMRMDASASSL----------PQQFDRRFSVC 220
Query: 303 D--MLSDEDNDTFSE---LANNEDANGIISVKSH---------------------PKYVR 336
D ML D +D + + ++ED H KY
Sbjct: 221 DRFMLGDTPDDFYDQSFRYCSSEDEPADSPCHDHYSPVSRTGSFVADDRDKGRDKSKYCL 280
Query: 337 IVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMCFVC 396
+ SKQMVGI+++VWV+ LR V++LKVS VG GLMGY+GNKGS+S+SMSV Q+ CFVC
Sbjct: 281 VASKQMVGIFLTVWVKSDLRDSVNNLKVSCVGRGLMGYLGNKGSISISMSVHQTSFCFVC 340
Query: 397 SHLASGQKDGAEQRRNSDVHEILQRTRFSSVF---DTDQPQKIPSHDKIFWFGDLNYRIN 453
SHL SGQK+G E RRNSDV EIL++TRF V D PQ I HD++ W GDLNYRI
Sbjct: 341 SHLTSGQKEGDELRRNSDVLEILRKTRFPRVNNAGDDKSPQMISEHDRVIWLGDLNYRIA 400
Query: 454 MSDGEIRKLVDLKKWNELMKFDQLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGG 513
+S + LV+++ W L++ DQL E KG VFEGWKEG I FPPTYKY NSD + G
Sbjct: 401 LSYRSAKALVEMRDWRALLEKDQLRIEQRKGCVFEGWKEGTIYFPPTYKYSNNSDIYAGD 460
Query: 514 NTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVIDHRKLER 573
+ KRR PAWCDRILW G GI QL Y E++ SDHRPV S+F V++E ++++
Sbjct: 461 DRLPKAKRRTPAWCDRILWHGSGISQLSYVRGESRFSDHRPVYSLFSVEIESAYRNRIKK 520
Query: 574 AIYFASAVVHPDVFLKE 590
+ + S+ + + L +
Sbjct: 521 SSSYTSSRIEVEELLPQ 537
>At5g65090 inositol-1, 4, 5-trisphosphate 5-phosphatase-like protein
Length = 529
Score = 310 bits (795), Expect = 1e-84
Identities = 186/477 (38%), Positives = 270/477 (55%), Gaps = 57/477 (11%)
Query: 95 KEVRVTIGTWNVAGKHPCNDLEIEGWLCTEEEPSDIYIIGFQEVVPLNAGNVFGAEDSKP 154
+E+RV + TWNV G+ P NDL +E +L E +D+YI GFQE+VPL+AGNV ED++P
Sbjct: 71 RELRVFLATWNVGGRTPNNDLNLEDFLLVEGT-ADLYICGFQEIVPLSAGNVLVVEDNEP 129
Query: 155 IPKWDALIRRTLNKSSEPGT-KKKSNSAPPSPIRRISSFNTQINPLDSALD--KKEEIKT 211
KW ALI + LNK + + SA + SS ++ + S+L ++ +K
Sbjct: 130 AAKWLALISQALNKPKQESVYSNAAYSASRTTTCSSSSCGSEESRAPSSLSFFQRPNLKV 189
Query: 212 IISIEKNLQLSKIYDIDLQTILDWPELRLDPIHHVDSSPKMRRVQSTSDSASLYGFEMKS 271
LS+ Y +D + + P+ + RR + SD ++
Sbjct: 190 ---------LSRNYRVDSSLL----KTCNCPVIDTSVGWEARRSKRFSDPST-------- 228
Query: 272 LHQSSGNFSLLWSEKQQEIVPQVFDSHLDVSDMLSDEDNDTFSELANNEDANGIISVKSH 331
SS N + P+ F H +N F ++ G +S
Sbjct: 229 --DSSNN-----------VEPENFRVH----------ENFLFDDVPATTKMPGQMS---- 261
Query: 332 PKYVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSR 391
Y I SKQMVG+++SVW +R+L H+ HL++ VG G+MG +GNKG +++SMS+ Q+
Sbjct: 262 --YRLIASKQMVGLFLSVWARRELIPHISHLRLDSVGRGIMGRLGNKGCIAISMSLHQTS 319
Query: 392 MCFVCSHLASGQKDGAEQRRNSDVHEILQRTRFSSVFDTDQ---PQKIPSHDKIFWFGDL 448
CFVCSHLASG+K+G E RRN+DV EIL+ T+F + P++I HD++ W GDL
Sbjct: 320 FCFVCSHLASGEKEGDELRRNADVAEILKHTQFPKLTKNPNCHAPERIIDHDRVLWLGDL 379
Query: 449 NYRINMSDGEIRKLVDLKKWNELMKFDQLSNELCKGHVFEGWKEGLINFPPTYKYEFNSD 508
NYR+ ++ E R L++ W+ L++ DQL+ E G VF G++EG I F PTYKY NSD
Sbjct: 380 NYRVALTYEETRVLLEDNDWDTLLERDQLNMERGAGRVFSGFQEGQIFFAPTYKYSQNSD 439
Query: 509 KHVGGNTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEV 565
+ G T+ +KRR PAWCDRILW G+GI+QL Y E++ SDHRPV +IF V+V+V
Sbjct: 440 AYAGEMTKSKKKRRTPAWCDRILWKGEGIEQLSYIRGESRFSDHRPVCAIFAVEVDV 496
>At5g04980 putative protein
Length = 366
Score = 267 bits (682), Expect = 1e-71
Identities = 133/276 (48%), Positives = 186/276 (67%), Gaps = 12/276 (4%)
Query: 301 VSDMLSDEDNDTFSELANNEDANG-----IISVKSHPKYVRIVSKQMVGIYVSVWVQRKL 355
+SD DED+D E +ED G ++S + KY + SKQMVGI+++VW++++L
Sbjct: 86 LSDDEEDEDDDDDDE---DEDEGGGKVASLVSNQMTMKYGLVASKQMVGIFLTVWMRKEL 142
Query: 356 RRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDV 415
+HV HL++S V G+MG +GNKG ++VS+ ++++ CF+CSHLASG+++G E+RRN DV
Sbjct: 143 IQHVSHLRISSVTRGIMGCLGNKGCIAVSLQLYKTSFCFICSHLASGEREGDERRRNLDV 202
Query: 416 HEILQRTRFSSVFDTD---QPQKIPSHDKIFWFGDLNYRINMSDGEIRKLVDLKKWNELM 472
EIL+ T F + T P +I HD++ W GDLNYRI +S E + L+D W+ L+
Sbjct: 203 IEILKNTSFPRICRTSFTRVPDRITKHDRVIWLGDLNYRIALSYSETKTLLDKNAWDTLL 262
Query: 473 KFDQLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGE-KRRAPAWCDRIL 531
DQL E G VF+GW EG I F PTYKY +NSD + G ++E + KRR PAWCDRIL
Sbjct: 263 NKDQLKIERDAGRVFKGWHEGKIFFAPTYKYSYNSDAYAGDTSKEKKNKRRTPAWCDRIL 322
Query: 532 WLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVID 567
W G GI+QL Y E++ SDHRPV S+F+VDVEV +
Sbjct: 323 WHGDGIRQLSYVRGESRFSDHRPVCSVFVVDVEVCE 358
>At2g37440 unknown protein
Length = 401
Score = 256 bits (654), Expect = 3e-68
Identities = 141/317 (44%), Positives = 193/317 (60%), Gaps = 25/317 (7%)
Query: 266 GFEMKSLHQSSGNFSLLWSEKQQEIVPQVFDSHLDVSDMLSDEDNDTFSEL--------- 316
G+ ++ F++L+ E Q EIVP L+ ++L EDN ++
Sbjct: 38 GYAYNHINWFYRTFNILFLEFQ-EIVP------LNAGNVLGAEDNGPAAKWLSLIREALN 90
Query: 317 -ANNEDANGI-----ISVKSHPK--YVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVG 368
NN +G+ I S P Y SKQMVGI++ VWV+ LR+ + +LKVS VG
Sbjct: 91 NTNNLSFSGLSDDTPIPCNSTPPRGYSLAASKQMVGIFLCVWVRDDLRKRITNLKVSCVG 150
Query: 369 VGLMGYMGNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDVHEILQRTRFSSVF 428
G+MGY+GNKGSVS+SMS+ ++ +CFVC+HL SG+K+G E RRN DV EI +RTRFS
Sbjct: 151 RGIMGYLGNKGSVSISMSLHETSLCFVCTHLTSGEKEGDELRRNLDVTEIFKRTRFSRSS 210
Query: 429 DTDQPQKIPSHDKIFWFGDLNYRINMSDGEIRKLVDLKKWNELMKFDQLSNELCKGHVFE 488
+P+ I HDK+ W GDLNYR+ S ++ + + W L++ DQL E G +F+
Sbjct: 211 KDSRPETIMDHDKVIWLGDLNYRLRAS-SDLHEQLRNHDWESLLEKDQLKIEQRAGRIFK 269
Query: 489 GWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAENQ 548
GW+EG I F PTYKY NSD +V + EKRR PAWCDRILW G G+KQL Y E++
Sbjct: 270 GWEEGKIYFAPTYKYRINSDNYVVQTEKSKEKRRTPAWCDRILWKGDGMKQLWYVRGESK 329
Query: 549 LSDHRPVSSIFLVDVEV 565
SDHRPV S+F V +++
Sbjct: 330 FSDHRPVQSLFSVHIDL 346
Score = 77.4 bits (189), Expect = 2e-14
Identities = 41/105 (39%), Positives = 58/105 (55%), Gaps = 20/105 (19%)
Query: 97 VRVTIGTWNVAGKHPCNDLEIEGWLCTEEEPSDIYIIG-------------------FQE 137
+R+ +GTWNV GK P L+++ WL + + +DIY++G FQE
Sbjct: 2 IRMFVGTWNVGGKSPHEGLDLKDWLKSPAD-ADIYVLGYAYNHINWFYRTFNILFLEFQE 60
Query: 138 VVPLNAGNVFGAEDSKPIPKWDALIRRTLNKSSEPGTKKKSNSAP 182
+VPLNAGNV GAED+ P KW +LIR LN ++ S+ P
Sbjct: 61 IVPLNAGNVLGAEDNGPAAKWLSLIREALNNTNNLSFSGLSDDTP 105
>At2g01900 putative inositol polyphosphate-5-phosphatase
Length = 417
Score = 251 bits (642), Expect = 6e-67
Identities = 125/265 (47%), Positives = 177/265 (66%), Gaps = 6/265 (2%)
Query: 315 ELANNEDANGIISVKSHPKYVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGY 374
+L+ ++ NGI + I+SKQMVGI ++VWV+ L ++ + VS VG G+MG
Sbjct: 138 DLSESKGINGISQ-----DFRCIISKQMVGILITVWVRGDLWPYIRYPSVSCVGCGIMGC 192
Query: 375 MGNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDVHEILQRTRFSSVFDTDQPQ 434
+GNKGSVSV + ++ CFVCSHLASG +D E++RNSDV+EIL R+ F D P+
Sbjct: 193 LGNKGSVSVRFQLHETTFCFVCSHLASGGRDRDERQRNSDVNEILARSSFPRGSSLDLPK 252
Query: 435 KIPSHDKIFWFGDLNYRINMSDGEIRKLVDLKKWNELMKFDQLSNELCKGHVFEGWKEGL 494
KI HD++ + GDLNYRI++ + + R LV+ KKWN L++ DQL E+ G +F GW+EG+
Sbjct: 253 KILDHDRVIFLGDLNYRISLPEEKTRLLVESKKWNILLENDQLRMEIMNGQIFRGWQEGI 312
Query: 495 INFPPTYKYEFNSDKHVGGNT-QEGEKRRAPAWCDRILWLGKGIKQLKYQSAENQLSDHR 553
+ F PTYKY NSD + G T ++ EK+RAPAWCDRI+W G G+KQ +Y E ++SDHR
Sbjct: 313 VKFAPTYKYVPNSDLYYGCITYKKDEKKRAPAWCDRIIWYGNGLKQHEYTRGETKISDHR 372
Query: 554 PVSSIFLVDVEVIDHRKLERAIYFA 578
PV +IF ++ V K R +F+
Sbjct: 373 PVKAIFTTEITVTRRGKKIRNFFFS 397
Score = 80.1 bits (196), Expect = 3e-15
Identities = 37/75 (49%), Positives = 52/75 (69%)
Query: 98 RVTIGTWNVAGKHPCNDLEIEGWLCTEEEPSDIYIIGFQEVVPLNAGNVFGAEDSKPIPK 157
+V + TWNV G P + L++E L T + P DIY++GFQEVVPL A NV G++++K K
Sbjct: 59 KVFVSTWNVGGIVPDDGLDMEDLLETHKTPCDIYVLGFQEVVPLRASNVLGSDNNKVSTK 118
Query: 158 WDALIRRTLNKSSEP 172
W++LIR LNK + P
Sbjct: 119 WNSLIRDALNKRARP 133
>At2g31830 putative inositol polyphosphate 5'-phosphatase
Length = 1144
Score = 130 bits (326), Expect = 3e-30
Identities = 79/226 (34%), Positives = 122/226 (53%), Gaps = 31/226 (13%)
Query: 334 YVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMC 393
+ R+ S+Q+ G+ +S+WV++ +R HV L V+ V G +GNKG V + + V+ MC
Sbjct: 661 FERMGSRQLAGLLISLWVRKSIRTHVGDLDVAAVPCGFGRAIGNKGGVGLRIRVYDRIMC 720
Query: 394 FVCSHLASGQKDGAEQRRNSDVHEILQRTRFS---SVFDT-------------------- 430
FV HLA+ + A RRN+D + I + FS SV+
Sbjct: 721 FVNCHLAAHLE--AVTRRNADFNHIYRSMVFSKGQSVYTAAAAGASTSAQALKNNPNTNN 778
Query: 431 ---DQPQKIPSHDKIFWFGDLNYRI-NMSDGEIRKLVDLKKWNELMKFDQLSNELCKGHV 486
++ + S D + +FGD NYR+ ++ E R + + ++ L + DQL E+ +G V
Sbjct: 779 STEEEKSHLASADLVAFFGDFNYRLFGITYDEARDFISHRSFDWLREKDQLRQEMNEGKV 838
Query: 487 FEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRAPAWCDRILW 532
F+G +E LI FPPTYK+E N K G GEK+R PAWCDR+++
Sbjct: 839 FQGMREALITFPPTYKFEKN--KPGLGGYDSGEKKRIPAWCDRVIY 882
Score = 29.6 bits (65), Expect = 5.0
Identities = 17/48 (35%), Positives = 23/48 (47%), Gaps = 1/48 (2%)
Query: 91 YINTKEVRVTIGTWNVAGKHPCNDLEIEGWLCTEEEPSDIYIIGFQEV 138
Y V++ IGTWNV G+ + + WL + I IG QEV
Sbjct: 574 YARQDSVKILIGTWNV-GEGRASRGALVSWLGSAVSDVGIVAIGLQEV 620
>At2g43900 putative inositol polyphosphate 5'-phosphatase
Length = 1305
Score = 129 bits (324), Expect = 5e-30
Identities = 86/276 (31%), Positives = 132/276 (47%), Gaps = 46/276 (16%)
Query: 334 YVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMC 393
+ R+ S+Q+ G+ +S+WV++ LR HV + V+ V G +GNKG V + + VF MC
Sbjct: 658 FERMGSRQLAGLLISLWVRKNLRTHVGDIDVAAVPCGFGRAIGNKGGVGLRIRVFDRIMC 717
Query: 394 FVCSHLASGQKDGAEQRRNSDVHEILQRTRF--------------------------SSV 427
F+ HLA+ + A RRN+D I + F ++V
Sbjct: 718 FINCHLAAHLE--AVNRRNADFDHIYKTMSFTRSSNAHNAPAAGVSTGSHTTKSANNANV 775
Query: 428 FDTDQPQKIPSHDKIFWFGDLNYRI-NMSDGEIRKLVDLKKWNELMKFDQLSNELCKGHV 486
+ Q + D + +FGD NYR+ +S E R V + ++ L + DQL E+ G V
Sbjct: 776 NTEETKQDLAEADMVVFFGDFNYRLFGISYDEARDFVSQRSFDWLREKDQLRAEMKAGRV 835
Query: 487 FEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAE 546
F+G +E +I FPPTYK+E + G GEK+R PAWCDR+++ S +
Sbjct: 836 FQGMREAIITFPPTYKFE--RHRPGLGGYDSGEKKRIPAWCDRVIFRDTRTSPESECSLD 893
Query: 547 NQL---------------SDHRPVSSIFLVDVEVID 567
+ SDH+PV F V +E +D
Sbjct: 894 CPVVASIMLYDACMDVTESDHKPVRCKFHVKIEHVD 929
Score = 32.7 bits (73), Expect = 0.59
Identities = 26/89 (29%), Positives = 40/89 (44%), Gaps = 11/89 (12%)
Query: 91 YINTKEVRVTIGTWNVAGKHPCNDLEIEGWLCTEEEPSDIYIIGFQEVVPLNAG------ 144
Y T VR+ G+WNV G+ + + WL + I ++G QE V + AG
Sbjct: 570 YAQTDSVRILTGSWNV-GQGKASHDALMSWLGSVASDVGILVVGLQE-VEMGAGFLAMSA 627
Query: 145 ---NVFGAEDSKPIPKWDALIRRTLNKSS 170
+V G E S W I +TL++ +
Sbjct: 628 AKESVGGNEGSTIGQYWIDTIGKTLDEKA 656
>At1g05630 putative inositol 1,4,5-trisphosphate 5-phosphatase
Length = 1136
Score = 126 bits (317), Expect = 3e-29
Identities = 90/290 (31%), Positives = 144/290 (49%), Gaps = 42/290 (14%)
Query: 334 YVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMC 393
+ R+ S+Q+ G+ +S+W ++ +R HV L V+ V G +GNKG V + + V+ MC
Sbjct: 652 FERMGSRQLAGLLISLWARKDIRTHVGDLDVAAVPCGFGRAIGNKGGVGLRIRVYDRIMC 711
Query: 394 FVCSHLASGQKDGAEQRRNSDVHEILQRTRF--------------SSVFDTDQPQKIPS- 438
FV HLA+ + A RRN+D + I + F S+ T + IPS
Sbjct: 712 FVNCHLAAHLE--AVNRRNADFNHIFRLMVFSRGQNLSNAAAAGVSTSAYTTKSNTIPST 769
Query: 439 -----------HDKIFWFGDLNYRI-NMSDGEIRKLVDLKKWNELMKFDQLSNELCKGHV 486
D + +FGD NYR+ ++ E R + + ++ L + DQL E+ G V
Sbjct: 770 GAEEIKSDLAAADMVAFFGDFNYRLFGITYDEARDFISQRSFDWLRERDQLRAEMKVGKV 829
Query: 487 FEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAE 546
F+G +E LI FPPTYK+E N + G GEK+R PAWCDR+++ + + + +
Sbjct: 830 FQGMREALITFPPTYKFERN--RSGLGGYDSGEKKRIPAWCDRVIY--RDTQSSPFSESN 885
Query: 547 NQLSDHRPVSSIFL----VDVEVIDHRKLERAIYFASAVVHPDVFLKEDE 592
Q VSS+ + +DV DH+ + F + + H D ++ E
Sbjct: 886 LQCP---VVSSVIMYEACMDVTESDHKPVR--CKFHATIAHVDKSVRRQE 930
Score = 32.3 bits (72), Expect = 0.77
Identities = 21/56 (37%), Positives = 28/56 (49%), Gaps = 4/56 (7%)
Query: 83 KSETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGWLCTEEEPSDIYIIGFQEV 138
+ ETL A+ N VR+ IGTWNV +D + WL + I +G QEV
Sbjct: 560 QKETLYARQDN---VRILIGTWNVGQGRASHD-ALMSWLGSVTSDVGIVAVGLQEV 611
>At1g47510 hypothetical protein
Length = 334
Score = 89.7 bits (221), Expect = 4e-18
Identities = 67/250 (26%), Positives = 120/250 (47%), Gaps = 25/250 (10%)
Query: 317 ANNEDANGIISVKSHPKYVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYM- 375
A + + ++ S P + + ++ + + ++ + V LK + VG G +
Sbjct: 92 APKANVDQLLQTASSPTHELLGKAKLQSVQLYLFGPKNSHTLVKELKAERYSVGGCGGLI 151
Query: 376 -GNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDVHEILQRTRFSSVFDTDQPQ 434
KG+V++ ++ +M F+ HL++ K +RN+++ I +S+ D+ +
Sbjct: 152 GRKKGAVAIRINYDDIKMVFISCHLSAHAKK--VDQRNTELRHIA-----NSLLPRDKRK 204
Query: 435 KIPSHDKIFWFGDLNYRI-NMSDGEIRKLVDLKKWNELMKFDQLSNELCKGHVFEGWKEG 493
+ D W GDLNYRI ++S+ +R L+ + L+ DQL E +G +F+G+ EG
Sbjct: 205 R----DLTVWLGDLNYRIQDVSNHPVRSLIQNHLQSVLVSKDQLLQEAERGEIFKGYSEG 260
Query: 494 LINFPPTYKYEFNSDKHVGGNTQEGEKRRAPAWCDRILWLGKGIKQLK-----YQSAENQ 548
+ F PTYKY S + K R PAW DRIL+ + ++ Y S +
Sbjct: 261 TLGFKPTYKYNVGSSDY-----DTSHKIRVPAWTDRILFKIQDTDNIQATLHSYDSIDQV 315
Query: 549 L-SDHRPVSS 557
SDH+PV +
Sbjct: 316 YGSDHKPVKA 325
>At1g65580 hypothetical protein
Length = 993
Score = 83.2 bits (204), Expect = 4e-16
Identities = 52/150 (34%), Positives = 77/150 (50%), Gaps = 30/150 (20%)
Query: 440 DKIFWFGDLNYRIN-MSDGEIRKLVDLKKWNELMKFDQLSNELCKGHVFEGWKEGLINFP 498
D + + GD NYR++ ++ E R + + ++ L + DQL E+ G+VF+G +E +I FP
Sbjct: 641 DMVIFLGDFNYRLDDITYDETRDFISQRCFDWLREKDQLHTEMEAGNVFQGMREAIIRFP 700
Query: 499 PTYKYEFNSDKHVGG--NTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAENQL------- 549
PTYK+E +H G GEK+R PAWCDRIL+ K+ AE L
Sbjct: 701 PTYKFE----RHQAGLAGYDSGEKKRIPAWCDRILYR----DNKKHLGAECSLDCPVVSS 752
Query: 550 ------------SDHRPVSSIFLVDVEVID 567
SDH+PV +F V + +D
Sbjct: 753 ISQYDACMEVTDSDHKPVRCVFSVKIARVD 782
>At3g09360 putative transcription factor
Length = 600
Score = 31.6 bits (70), Expect = 1.3
Identities = 38/165 (23%), Positives = 62/165 (37%), Gaps = 36/165 (21%)
Query: 296 DSHLDVSDMLSDEDNDTFSELANNEDANGIISVKSHPKYVRIVSKQMVGIYVSVWVQRK- 354
+ H D SD ++D FS+++++E NG I+ + Y I +M Y+ ++
Sbjct: 383 EEHADTSD-----ESDNFSDISDDE-VNGYINNEEETHYKTITWTEMNKDYLEEQAAKEA 436
Query: 355 -LRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNS 413
L+ LK S N + F++ Q+ AE+ +N+
Sbjct: 437 ALKAASEALKAS-----------NSNCPEDARKAFEAAKADAAKSRKEKQQKKAEEAKNA 485
Query: 414 D--------VHEILQRTRFSSV---------FDTDQPQKIPSHDK 441
V L + R SSV FDT P+K P K
Sbjct: 486 APPATAVEAVRRTLDKKRLSSVINYDVLESLFDTSAPEKSPKRSK 530
>At5g40690 unknown protein
Length = 210
Score = 31.2 bits (69), Expect = 1.7
Identities = 23/83 (27%), Positives = 36/83 (42%), Gaps = 13/83 (15%)
Query: 54 TSKPQAN-----TTKPTQLN--------TSCKDYEMKRHRRRKSETLRAQYINTKEVRVT 100
TS+P A T+KPT + C D M + K++T + + T +V
Sbjct: 34 TSRPTAGLFTKVTSKPTNHSKFTGKCGQARCLDCHMHPVTKSKAKTKGSSKVRTNDVTYK 93
Query: 101 IGTWNVAGKHPCNDLEIEGWLCT 123
+ TW VA P L++ G+ T
Sbjct: 94 MLTWQVAAGGPRPGLKLSGFSAT 116
>At4g39870 unknown protein
Length = 394
Score = 31.2 bits (69), Expect = 1.7
Identities = 18/41 (43%), Positives = 22/41 (52%), Gaps = 4/41 (9%)
Query: 30 EDEANTETDESEDDVKLIEIETNHTSKPQANTTKPTQLNTS 70
EDE + E DE ED E ET+ TS AN T+ + TS
Sbjct: 93 EDEEDEEEDEEEDS----EAETSDTSSSSANPTRTMKETTS 129
>At2g06310 putative Athila retroelement ORF1 protein
Length = 733
Score = 31.2 bits (69), Expect = 1.7
Identities = 14/53 (26%), Positives = 26/53 (48%)
Query: 29 SEDEANTETDESEDDVKLIEIETNHTSKPQANTTKPTQLNTSCKDYEMKRHRR 81
+ ++ T TD+SED +E+ +H ++P T P T+ K +K +
Sbjct: 384 TREDPKTVTDDSEDQDGPLELPLDHFTRPTTRATSPATSPTAPKPVAVKNKEK 436
>At5g07660 SMC-like protein
Length = 1058
Score = 30.8 bits (68), Expect = 2.2
Identities = 22/77 (28%), Positives = 32/77 (40%), Gaps = 6/77 (7%)
Query: 31 DEANTETDESEDDVKLIEIETNHTSKPQANTTKP--TQLNTSCKDYEMKRHRRRKSETLR 88
++A E E ED++ E E NH + P Q T K+ EMKR K +
Sbjct: 773 EKAEDELKEKEDELHSAETEKNHYEDIMKDKVLPEIKQAETIYKELEMKRQESNK----K 828
Query: 89 AQYINTKEVRVTIGTWN 105
A I + +G W+
Sbjct: 829 ASIICPESEIKALGPWD 845
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.132 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,445,290
Number of Sequences: 26719
Number of extensions: 661420
Number of successful extensions: 2280
Number of sequences better than 10.0: 37
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 2201
Number of HSP's gapped (non-prelim): 52
length of query: 599
length of database: 11,318,596
effective HSP length: 105
effective length of query: 494
effective length of database: 8,513,101
effective search space: 4205471894
effective search space used: 4205471894
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC147404.2


