

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147363.1 - phase: 0 /pseudo
(427 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
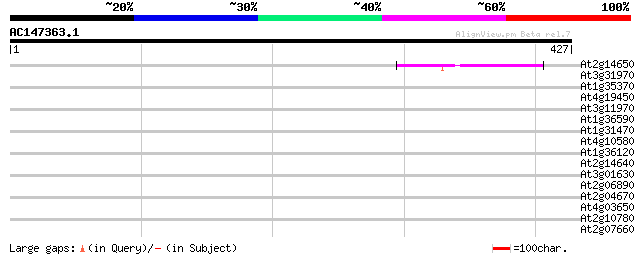
Score E
Sequences producing significant alignments: (bits) Value
At2g14650 putative retroelement pol polyprotein 42 6e-04
At3g31970 hypothetical protein 39 0.007
At1g35370 hypothetical protein 37 0.016
At4g19450 unknown protein 36 0.046
At3g11970 hypothetical protein 35 0.061
At1g36590 hypothetical protein 35 0.061
At1g31470 hypothetical protein 34 0.13
At4g10580 putative reverse-transcriptase -like protein 33 0.30
At1g36120 putative reverse transcriptase gb|AAD22339.1 33 0.30
At2g14640 putative retroelement pol polyprotein 33 0.39
At3g01630 hypothetical protein 32 0.51
At2g06890 putative retroelement integrase 30 2.5
At2g04670 putative retroelement pol polyprotein 29 4.3
At4g03650 putative reverse transcriptase 28 7.4
At2g10780 pseudogene 28 9.7
At2g07660 putative retroelement pol polyprotein 28 9.7
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 42.0 bits (97), Expect = 6e-04
Identities = 33/115 (28%), Positives = 48/115 (41%), Gaps = 6/115 (5%)
Query: 295 IGTCYINNTPLVAIIDIGATHCFIAFDCASTLGL--DMSNMNGEMVVETPAKGSVTTSLV 352
+GT + + D GA+HCFI + AS + D G + V A G L
Sbjct: 303 VGTLLVGGVEAHVLFDSGASHCFITPESASRGNIRGDPGEQLGAVKV---AGGQFVAVLG 359
Query: 353 CLKS-SLSMFGRDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYFSKSVYFSSVE 406
K + + G DL+ P+ VILGM+W + VH++ V F E
Sbjct: 360 RTKGVDIQIAGESMPADLIISPVELYDVILGMDWLDHYRVHLDCHRGRVSFERPE 414
>At3g31970 hypothetical protein
Length = 1329
Score = 38.5 bits (88), Expect = 0.007
Identities = 31/102 (30%), Positives = 45/102 (43%), Gaps = 6/102 (5%)
Query: 308 IIDIGATHCFIAFDCASTLGL--DMSNMNGEMVVETPAKGSVTTSLVCLKS-SLSMFGRD 364
+ D GA+HCFI + AS + D G + V A G L K + + G
Sbjct: 415 LFDSGASHCFITPESASRGNIRGDPGEQLGAVKV---AGGQFLAVLGRAKGVDIQIAGES 471
Query: 365 FEMDLVCLPLSGMYVILGMNWFE*NHVHINYFSKSVYFSSVE 406
+DL+ P+ VILGM+W + VH++ V F E
Sbjct: 472 MPVDLIISPVELYDVILGMDWLDHYRVHLDCHRGRVSFERPE 513
>At1g35370 hypothetical protein
Length = 1447
Score = 37.4 bits (85), Expect = 0.016
Identities = 30/121 (24%), Positives = 57/121 (46%), Gaps = 9/121 (7%)
Query: 268 KQPKKAPTIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDCASTLG 327
++P ++ V ++G +T + GT ++ L +ID G+TH FI A+ LG
Sbjct: 349 QKPMPQISVNAVSGISGYKTMG----VKGT--VDKRDLFILIDSGSTHNFIDSTVAAKLG 402
Query: 328 LDMSNMN-GEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILGMNWF 386
+ + ++ V K +V + L F+ D++ +PL G+ ++LG+ W
Sbjct: 403 CHVESAGLTKVAVADGRKLNVDGQIKGFTWKLQ--STTFQSDILLIPLQGVDMVLGVQWL 460
Query: 387 E 387
E
Sbjct: 461 E 461
>At4g19450 unknown protein
Length = 572
Score = 35.8 bits (81), Expect = 0.046
Identities = 19/47 (40%), Positives = 29/47 (61%), Gaps = 1/47 (2%)
Query: 317 FIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGR 363
+I + C T+GL SN G+ + ++ + S TT+LV L S+ S FGR
Sbjct: 358 YITYFCGGTIGLVYSNNLGQ-IAQSLGQSSNTTTLVTLYSAFSFFGR 403
>At3g11970 hypothetical protein
Length = 1499
Score = 35.4 bits (80), Expect = 0.061
Identities = 24/85 (28%), Positives = 45/85 (52%), Gaps = 11/85 (12%)
Query: 308 IIDIGATHCFIAFDCASTLG--LDMSNMNGEMVVE---TPAKGSVTTSLVCLKSSLSMFG 362
+ID G+TH F+ + A+ LG +D + + V + +G VT L+++
Sbjct: 404 LIDSGSTHNFLDPNTAAKLGCKVDTAGLTRVSVADGRKLRVEGKVTDFSWKLQTTT---- 459
Query: 363 RDFEMDLVCLPLSGMYVILGMNWFE 387
F+ D++ +PL G+ ++LG+ W E
Sbjct: 460 --FQSDILLIPLQGIDMVLGVQWLE 482
>At1g36590 hypothetical protein
Length = 1499
Score = 35.4 bits (80), Expect = 0.061
Identities = 24/85 (28%), Positives = 45/85 (52%), Gaps = 11/85 (12%)
Query: 308 IIDIGATHCFIAFDCASTLG--LDMSNMNGEMVVE---TPAKGSVTTSLVCLKSSLSMFG 362
+ID G+TH F+ + A+ LG +D + + V + +G VT L+++
Sbjct: 404 LIDSGSTHNFLDPNTAAKLGCKVDTAGLTRVSVADGRKLRVEGKVTDFSWKLQTTT---- 459
Query: 363 RDFEMDLVCLPLSGMYVILGMNWFE 387
F+ D++ +PL G+ ++LG+ W E
Sbjct: 460 --FQSDILLIPLQGIDMVLGVQWLE 482
>At1g31470 hypothetical protein
Length = 546
Score = 34.3 bits (77), Expect = 0.13
Identities = 19/47 (40%), Positives = 28/47 (59%), Gaps = 3/47 (6%)
Query: 317 FIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGR 363
+IA+ C T+GL SN G++ + G +T+LV + SS S FGR
Sbjct: 328 YIAYFCGGTIGLVYSNNLGQI---AQSLGQNSTTLVTIYSSFSFFGR 371
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 33.1 bits (74), Expect = 0.30
Identities = 24/86 (27%), Positives = 37/86 (42%), Gaps = 11/86 (12%)
Query: 322 CASTLGLDMSNMNGE-MVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVI 380
C S LD + G+ + V AKG + + G DL+ P+ VI
Sbjct: 331 CFSCGSLDHKDAGGQFLAVLGRAKGV----------DIQIAGESMPADLIISPVELYDVI 380
Query: 381 LGMNWFE*NHVHINYFSKSVYFSSVE 406
LGM+W + VH+++ V+F E
Sbjct: 381 LGMDWLDYYRVHLDWHRGRVFFERPE 406
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 33.1 bits (74), Expect = 0.30
Identities = 30/104 (28%), Positives = 44/104 (41%), Gaps = 10/104 (9%)
Query: 308 IIDIGATHCFIAFDCASTLGLDMSNMNGEMVVETPAK----GSVTTSLVCLKS-SLSMFG 362
+ D GA+HCFI + AS N+ GE + A G T L K ++ +
Sbjct: 352 LFDSGASHCFITPESASR-----DNIRGEPGEQFGAVKIAGGQFLTVLGRAKGVNIQIEE 406
Query: 363 RDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYFSKSVYFSSVE 406
DL+ P+ ILGM+W + VH++ V F E
Sbjct: 407 ESMPADLIINPVVLYDAILGMDWLDHYRVHLDCHRGRVSFERPE 450
>At2g14640 putative retroelement pol polyprotein
Length = 945
Score = 32.7 bits (73), Expect = 0.39
Identities = 31/113 (27%), Positives = 48/113 (42%), Gaps = 16/113 (14%)
Query: 279 VFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDCASTLGLDMSNMNGEMV 338
++AL G T + R+ I+ L+A+I+ G+TH FI + MN +
Sbjct: 306 LYALEGVDTTSTIRV---RATIHRNRLIALINSGSTHNFIGEKA-------VRGMNLKAT 355
Query: 339 VETPAKGSVTTS--LVCLKS----SLSMFGRDFEMDLVCLPLSGMYVILGMNW 385
P V LVC + M G F + L LPL G+ + +G+ W
Sbjct: 356 TTKPFTVRVVNGMPLVCRSRYEAIPVVMGGVVFPVTLYALPLMGLDLAMGVQW 408
>At3g01630 hypothetical protein
Length = 569
Score = 32.3 bits (72), Expect = 0.51
Identities = 17/47 (36%), Positives = 24/47 (50%)
Query: 317 FIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGR 363
++A+ C T+GL SN G++ S SLV L S+ S GR
Sbjct: 357 YVAYFCGGTIGLVYSNNLGQIAQSLGQSSSNAKSLVTLFSAFSFLGR 403
>At2g06890 putative retroelement integrase
Length = 1215
Score = 30.0 bits (66), Expect = 2.5
Identities = 38/177 (21%), Positives = 67/177 (37%), Gaps = 33/177 (18%)
Query: 265 SQCKQPKKAPTIGRVFALTGT--------QTENEDRLIIGTCYINNTPLVAIIDIGATHC 316
S K+ ++ P G + T + E L C+++ IID G+
Sbjct: 197 SSLKENEELPAQGELLVARRTLSVQTKTDEQEQRKNLFHTRCHVHGKVCSLIIDGGSCTN 256
Query: 317 FIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVC--LPL 374
+ LGL N +G+M V K V +V K +E +++C LP+
Sbjct: 257 VASETMVKKLGLKWLNDSGKMRV----KNQVVVPIVIGK---------YEDEILCDVLPM 303
Query: 375 SGMYVILGMNWFE*NHVHINYFS----------KSVYFSSVEEEIGAEFLFTKQLKQ 421
+++LG W V + F+ K++ S E+ + + KQ K+
Sbjct: 304 EAGHILLGRPWQSDRKVMHDGFTNRHSFEFKGGKTILVSMTPHEVYQDQIHLKQKKE 360
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 29.3 bits (64), Expect = 4.3
Identities = 15/49 (30%), Positives = 23/49 (46%)
Query: 358 LSMFGRDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYFSKSVYFSSVE 406
+ + G DL+ P+ VILGM+W + VH++ V F E
Sbjct: 384 IQIAGESMPADLIISPVELYDVILGMDWLDHYRVHLDCHRGRVSFERPE 432
>At4g03650 putative reverse transcriptase
Length = 839
Score = 28.5 bits (62), Expect = 7.4
Identities = 16/55 (29%), Positives = 24/55 (43%)
Query: 358 LSMFGRDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYFSKSVYFSSVEEEIGAE 412
+ + G DL+ + VILGM+W + VH++ V F E G E
Sbjct: 377 IQIAGESLPADLIISHVELYDVILGMDWLDHYRVHLDCHRGRVSFERPEGSAGRE 431
>At2g10780 pseudogene
Length = 1611
Score = 28.1 bits (61), Expect = 9.7
Identities = 16/50 (32%), Positives = 24/50 (48%)
Query: 357 SLSMFGRDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYFSKSVYFSSVE 406
S+ + G + DL+ +PL VILGM+W HI+ + F E
Sbjct: 525 SVVVNGVNMPADLIIVPLKKHDVILGMDWLGKYKAHIDCHRGRIQFERDE 574
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 28.1 bits (61), Expect = 9.7
Identities = 16/50 (32%), Positives = 25/50 (50%)
Query: 357 SLSMFGRDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYFSKSVYFSSVE 406
S+ + G + DL+ +PL VILGM+W HI+ + + F E
Sbjct: 20 SVVVNGVNMPADLIIVPLKKHDVILGMDWLGKYKGHIDCHRERIQFERDE 69
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.343 0.151 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,947,376
Number of Sequences: 26719
Number of extensions: 294858
Number of successful extensions: 925
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 906
Number of HSP's gapped (non-prelim): 17
length of query: 427
length of database: 11,318,596
effective HSP length: 102
effective length of query: 325
effective length of database: 8,593,258
effective search space: 2792808850
effective search space used: 2792808850
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147363.1


