

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147202.5 - phase: 0
(389 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
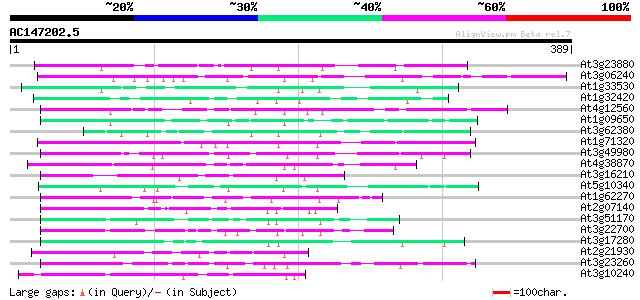
Score E
Sequences producing significant alignments: (bits) Value
At3g23880 unknown protein 107 9e-24
At3g06240 unknown protein 84 2e-16
At1g33530 hypothetical protein 69 3e-12
At1g32420 hypothetical protein 69 4e-12
At4g12560 putative protein 68 1e-11
At1g09650 hypothetical protein 67 2e-11
At3g62380 putative protein 65 8e-11
At1g71320 hypothetical protein 64 1e-10
At3g49980 putative protein 64 2e-10
At4g38870 putative protein 63 2e-10
At3g16210 hypothetical protein 61 9e-10
At5g10340 putative protein 61 1e-09
At1g62270 hypothetical protein 60 2e-09
At2g07140 pseudogene 59 3e-09
At3g51170 putative protein 59 6e-09
At3g22700 hypothetical protein 59 6e-09
At3g17280 hypothetical protein 59 6e-09
At2g21930 unknown protein 58 8e-09
At3g23260 unknown protein 58 1e-08
At3g10240 hypothetical protein 58 1e-08
>At3g23880 unknown protein
Length = 364
Score = 107 bits (268), Expect = 9e-24
Identities = 93/325 (28%), Positives = 146/325 (44%), Gaps = 58/325 (17%)
Query: 18 SPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLH--------LQRSIQN 69
SP LPL+++ E+L LPVKSL +F+CVC SW+ LIS++ F H S ++
Sbjct: 10 SPHNLPLEMMEEILLRLPVKSLTRFKCVCSSWRSLISETLFALKHALILETSKATTSTKS 69
Query: 70 PRLAVTQESRECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLL 129
P +T I H L N S T ++ L +D V G+C+GL+
Sbjct: 70 PYGVITTSRYHLKSCCI-------HSLYNAS--TVYVSEHDGELLGRDYYQVVGTCHGLV 120
Query: 130 CLLGLSEDREMWLRFWNPATRAISDKLGHYPADFTGG---FEVAFGYDNSTDTYKVVYL- 185
C + D+ ++L WNP T + +L + + FGYD S D YKVV L
Sbjct: 121 C-FHVDYDKSLYL--WNP-TIKLQQRLSSSDLETSDDECVVTYGFGYDESEDDYKVVALL 176
Query: 186 -----QKGMTGVFSLGDNVWRNIESFPLGYYLDNR----VHLRDSLNWLGLRSYVDDCDD 236
K T ++S +WR+ SFP G + ++ +++ +LNW
Sbjct: 177 QQRHQVKIETKIYSTRQKLWRSNTSFPSGVVVADKSRSGIYINGTLNWAA---------- 226
Query: 237 YDCEYITSIEQFMIVALDLRTETCKELLLP----RGFDEVPCYEPSLCVLMDCICFSHLV 292
+S + I++ D+ + KEL P RG C+ +L L C+
Sbjct: 227 -----TSSSSSWTIISYDMSRDEFKELPGPVCCGRG-----CFTMTLGDLRGCLSMVCYC 276
Query: 293 KKTHLVIWKMMDYGDDDSWTQLLEI 317
K + +W M ++G+ SW++LL I
Sbjct: 277 KGANADVWVMKEFGEVYSWSKLLSI 301
>At3g06240 unknown protein
Length = 427
Score = 83.6 bits (205), Expect = 2e-16
Identities = 111/424 (26%), Positives = 173/424 (40%), Gaps = 96/424 (22%)
Query: 20 IILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNP------RLA 73
++LP ++ITE+L LP KS+ +FRCV + L SD F K+HL ++N R
Sbjct: 34 LVLPPEIITEILLRLPAKSIGRFRCVSKLFCTLSSDPGFAKIHLDLILRNESVRSLHRKL 93
Query: 74 VTQESRECSMDV------IVSPTSLSH---LLENPSKPTTL----TNDPYYS-------L 113
+ S+D I ++ H L ++PS + + D Y L
Sbjct: 94 IVSSHNLYSLDFNSIGDGIRDLAAVEHNYPLKDDPSIFSEMIRNYVGDHLYDDRRVMLKL 153
Query: 114 NDKDCR----SVAGSCNGLLCLLGLSEDREMWLRFWNPAT-------RAISDKLGHYPAD 162
N K R + GS NGL+C+ E + +NP T K Y D
Sbjct: 154 NAKSYRRNWVEIVGSSNGLVCI----SPGEGAVFLYNPTTGDSKRLPENFRPKSVEYERD 209
Query: 163 FTGGFEV-AFGYDNSTDTYKVVYLQKGM-----TGVFSLGDNVWRNIESFPL----GYYL 212
F+ FG+D TD YK+V L V+SL + WR I + G Y
Sbjct: 210 ---NFQTYGFGFDGLTDDYKLVKLVATSEDILDASVYSLKADSWRRICNLNYEHNDGSYT 266
Query: 213 DNRVHLRDSLNWLGLRSYVDDCDDYDCEYITSIEQFMIVALDLRTETCKELLLPRGFDE- 271
VH +++W+ S + Q ++VA D++TE +E+ +P ++
Sbjct: 267 SG-VHFNGAIHWVFTESRHN--------------QRVVVAFDIQTEEFREMPVPDEAEDC 311
Query: 272 --------VPCYEPSLCVLMDCICFSHLVKKTHLVIWKMMDYGDDDSWTQLLEINLQILK 323
V LCV+ C H IW M +YG+ SW++ + INL
Sbjct: 312 SHRFSNFVVGSLNGRLCVVNSCY-------DVHDDIWVMSEYGEAKSWSR-IRINLL--- 360
Query: 324 KIDEWSAWVPLHLSKNYDTLILGSTLEHEFAVYNLRDGSIEKTRITNGE-SSWIYINDYV 382
+ + PL +KN + ++L L+ + +YN + I + S N YV
Sbjct: 361 ----YRSMKPLCSTKNDEEVLL--ELDGDLVLYNFETNASSNLGICGVKLSDGFEANTYV 414
Query: 383 ESLV 386
ESL+
Sbjct: 415 ESLI 418
>At1g33530 hypothetical protein
Length = 441
Score = 69.3 bits (168), Expect = 3e-12
Identities = 78/318 (24%), Positives = 120/318 (37%), Gaps = 52/318 (16%)
Query: 9 QQPPSNVASSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLH---LQR 65
+ P + + + LP L+ E+L LPVK L++ + + WK LI + H L++
Sbjct: 84 KSPSCSETTLAVELPDVLVEEILQRLPVKYLVRLKSISKGWKSLIESDHLAEKHLRLLEK 143
Query: 66 SIQNPRLAVTQESRECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSC 125
+ +T E R S + + S + +D D V GSC
Sbjct: 144 KYGLKEIKITVE-RSTSKSICIK------FFSRRSGMNAINSD------SDDLLRVPGSC 190
Query: 126 NGLLCLLGLSEDREMWLRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNSTDTYKVVYL 185
NGL+C+ L L TR ++ G V FG D T TYKV+ L
Sbjct: 191 NGLVCVYELDSVYIYLLNPMTGVTRTLTPPRG-------TKLSVGFGIDVVTGTYKVMVL 243
Query: 186 ---QKGMTGVFSLGDNVWR----NIESFPLGYYLD---NRVHLRDSLNWLGLRSYVDDCD 235
+ T VF L N WR PL N V + SL WL + +
Sbjct: 244 YGFDRVGTVVFDLDTNKWRQRYKTAGPMPLSCIPTPERNPVFVNGSLFWLLASDFSE--- 300
Query: 236 DYDCEYITSIEQFMIVALDLRTETCKELLLPRGFDEVPCYEPSLCV--LMDCICFSHLVK 293
I+ +DL TE + L P D+V + + L D +C S++ +
Sbjct: 301 --------------ILVMDLHTEKFRTLSQPNDMDDVDVSSGYIYMWSLEDRLCVSNVRQ 346
Query: 294 KTHLVIWKMMDYGDDDSW 311
H +W ++ + W
Sbjct: 347 GLHSYVWVLVQDELSEKW 364
>At1g32420 hypothetical protein
Length = 302
Score = 68.9 bits (167), Expect = 4e-12
Identities = 76/309 (24%), Positives = 125/309 (39%), Gaps = 65/309 (21%)
Query: 17 SSPIILPLDLITELLSFLPVKSLLQFRCVCMSW-KILISDSFFVKLHLQRSIQNPRLAVT 75
S + +PLDL E+L LP KSLL+F+CV W I+ S F+ + RS+ P
Sbjct: 32 SGNVNIPLDLTVEILKKLPAKSLLRFQCVSKQWLSIISSRRDFIDSIVTRSLTQP----- 86
Query: 76 QESRECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGS----CNGLLCL 131
R+ + H + P + + Y DK+ + S GL+C
Sbjct: 87 -PPRDIKL-------IFHHQVLYPGPHFFIFSSTYPQNTDKESLTTRASSYHYVRGLICC 138
Query: 132 LGLSEDREMWLRFWNPATRAISDKLGHYPADFTGGFEVA----FGYDNSTDTYKVV---- 183
+ +NP TR +Y T ++ FGYD + YKV+
Sbjct: 139 WSHCPTT---VDIYNPTTRQ------YYTVPDTNRYQYIETCFFGYDPVENQYKVMVLPK 189
Query: 184 -YLQKGMTGVFSLGDNV---WRNIESFPLGYYLDNRVHLRDSLNWLGLRSYVDDCDDYDC 239
Y+++ VF++GD + WR+I+ + + L + V + + Y ++Y
Sbjct: 190 YYMEESPCQVFTVGDPIEKPWRDIQGIGVHFLLKDAVCINGVI-------YYQATNEYGS 242
Query: 240 EYITSIEQFMIVALDLRTETCKELLLPRGFDEVPC----YEPSLCVLMDCICFSHLVKKT 295
Y +V+ D+R+E + P+ + PC Y+ L ++M C K
Sbjct: 243 TY-------FLVSFDVRSEKFNHVKAPKILTDHPCTLINYQGKLGLIMCC--------KK 287
Query: 296 HLVIWKMMD 304
L IW M D
Sbjct: 288 GLEIWVMED 296
>At4g12560 putative protein
Length = 408
Score = 67.8 bits (164), Expect = 1e-11
Identities = 93/359 (25%), Positives = 151/359 (41%), Gaps = 68/359 (18%)
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQ---NPRLAVTQES 78
+P+D++ ++ LP K+L++ R + LI+D F++ HL R +Q + + +
Sbjct: 4 IPMDIVNDIFLRLPAKTLVRCRALSKPCYHLINDPDFIESHLHRVLQTGDHLMILLRGAL 63
Query: 79 RECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSEDR 138
R S+D + S S+S +E+P K T V GS NGL+ L D
Sbjct: 64 RLYSVD-LDSLDSVSD-VEHPMKRGGPT-------------EVFGSSNGLIGLSNSPTD- 107
Query: 139 EMWLRFWNPATRAISDKLGHYPADFTGGFEV------AFGYDNSTDTYKVVYLQKGM--- 189
L +NP+TR I +L D G GYD+ +D YKVV + +
Sbjct: 108 ---LAVFNPSTRQI-HRLPPSSIDLPDGSSTRGYVFYGLGYDSVSDDYKVVRMVQFKIDS 163
Query: 190 -----------TGVFSLGDNVWRNIES--------FPLGYYLDNR----VHLRDSLNWLG 226
VFSL N W+ IES F Y+L R V +SL+W+
Sbjct: 164 EDELGCSFPYEVKVFSLKKNSWKRIESVASSIQLLFYFYYHLLYRRGYGVLAGNSLHWVL 223
Query: 227 LRSYVDDCDDYDCEYITSIEQFMIVALDLRTETCKELLLPRGFDEVPCYEPSLCVLMDCI 286
R + + ++E+F IV E + D + + VL C+
Sbjct: 224 PRRPGLIAFNLIVRFDLALEEFEIVRF-------PEAVANGNVD----IQMDIGVLDGCL 272
Query: 287 CFSHLVKKTHLVIWKMMDYGDDDSWTQLLEINLQILKKIDEWSAWVPLHLSKNYDTLIL 345
C ++++ +W M +Y DSWT++ + Q K + +S PL SK+ ++L
Sbjct: 273 CLMCNYDQSYVDVWMMKEYNVRDSWTKVFTV--QKPKSVKSFSYMRPLVYSKDKKKVLL 329
>At1g09650 hypothetical protein
Length = 382
Score = 66.6 bits (161), Expect = 2e-11
Identities = 81/329 (24%), Positives = 129/329 (38%), Gaps = 55/329 (16%)
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQ--NPRLAVTQESR 79
LP +++ +L L LL+F+ V WK I FF + + Q NP + +
Sbjct: 13 LPHEVVECILERLDADPLLRFKAVSKQWKSTIESPFFQRRQFTQRQQSGNPDVLMVSRCA 72
Query: 80 ECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSEDRE 139
+ + ++ T + + PT + DK+ SC+GL+CL D+
Sbjct: 73 DINSEIEALTTLVLGSSSSVKIPTPWEEE---EEEDKEYSVSQDSCDGLVCLF----DKF 125
Query: 140 MWLRFWNPATR---------------AISDKLGHYPADFTGGFEVAFGYDNSTDTYKVVY 184
+NPATR AI D G Y + + FG D T TYK V+
Sbjct: 126 KSGFVFNPATRWYHPLPLCQLQQLLIAIGD--GFYDLGYRVS-RLGFGKDKLTGTYKPVW 182
Query: 185 LQKGM---------TGVFSLGDNVWRNIESFPLGYYLDNRVHLRDSLNWLGLRSYVDDCD 235
L + VF N WR + P Y +G + V C
Sbjct: 183 LYNSIEIGLENATTCEVFDFSTNAWRYVS--PAAPY-----------RIVGCPAPV--CV 227
Query: 236 DYDCEYITSIEQFMIVALDLRTETCKELLLPRGFDEVPCYEPSLCVLMDCICFSHLVKKT 295
D + T E+ I++ DL TET +++ F V ++ +C L + +C S + K
Sbjct: 228 DGSLHWFTECEETKILSFDLHTETF-QVVSKAPFANVDGFDIVMCNLDNRLCVSEM-KLP 285
Query: 296 HLVIWKMMDYGDDDSWTQLLEINLQILKK 324
+ VIW + +W ++ INL I +
Sbjct: 286 NQVIWSF--NSGNKTWHKMCSINLDITSR 312
>At3g62380 putative protein
Length = 325
Score = 64.7 bits (156), Expect = 8e-11
Identities = 73/293 (24%), Positives = 117/293 (39%), Gaps = 46/293 (15%)
Query: 52 LISDSFFVKLHLQRSIQNPRLAVTQ---ESRECSMDVIVSPTSLSH------LLENPSKP 102
+I ++ K L R NP+L V + S CS + + S H + + + P
Sbjct: 1 MIESTYLAKKRLVRH-PNPKLLVLRLEMSSDRCSRTLFLETYSKDHHDHINIFIHSYTSP 59
Query: 103 TTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSEDREMWLRFWNPATRAISDKLGHYPAD 162
P YS D + G CNG++C+ L ++ NPATR + + +
Sbjct: 60 VPY---PKYSTYDT-VGQIMGYCNGIVCIYDLG-----YIYLINPATRKLRILSPEFLRE 110
Query: 163 FTGG--------FEVAFGYDNSTDTYKVVYL---QKGMTGVFSLG-DNVWRNIESFPLGY 210
+TG V FG D T TYKV+ + + +L N R FP+ Y
Sbjct: 111 YTGRTGWSNDLPINVGFGRDMVTGTYKVILMYLFDRNFVKTEALNLKNGERRYVYFPISY 170
Query: 211 -YLDN---RVHLRDSLNWLGLRSYVDDCDDYDCEYITSIEQFMIVALDLRTETCKELLLP 266
L N + SL WL D D Y + I + A+DL TET + + LP
Sbjct: 171 GQLGNDKRSIFANGSLYWLPK----PDVDFYTNQLIK------LAAIDLHTETFRYVPLP 220
Query: 267 RGFDEVPCYEPSLCVLMDCICFSHLVKKTHLVIWKMMDYGDDDSWTQLLEINL 319
+ + C L L D +C S +++ + +W + W ++ +N+
Sbjct: 221 SWYTKY-CKSVYLWSLKDSLCVSDVLQNPSVDVWSLQQEEPSVKWEKMFSVNI 272
>At1g71320 hypothetical protein
Length = 392
Score = 63.9 bits (154), Expect = 1e-10
Identities = 74/346 (21%), Positives = 140/346 (40%), Gaps = 57/346 (16%)
Query: 20 IILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRS-IQNPRLAVTQES 78
I +P D+ + LP+KSL +F+ + W +I ++F L R+ + P + S
Sbjct: 12 IFIPDDIAEGIFHHLPIKSLARFKVLSKKWTSMIESTYFSHKRLIRTGLPTPNMKFLHIS 71
Query: 79 RECSMDVIVS-PTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCL------ 131
+ + + + S++ LLE S+ + +K + V GSC+GL+ L
Sbjct: 72 QHFTANFVEEYSNSITFLLETFSRDDQNNRKTFDESQNKTIQ-VLGSCDGLVLLRIHDDF 130
Query: 132 -------------LGLSEDREMW---LRFWNPAT-----RAISDKLGHYPADFTGGFEVA 170
+ LS + W L + PA R +S + Y +A
Sbjct: 131 RSIYLINPTTKEHMKLSPEFMQWPFTLSYLTPAMANKPWRQLSQSVLDYDISVKRMPSLA 190
Query: 171 -FGYDNSTDTYKVVYL--------QKGMTGVFSLGDNVWRNIESFPLGYYL----DNRVH 217
FG D +YKVV + + V SL + R++ + + + + V+
Sbjct: 191 GFGKDIVKKSYKVVLIYTRYEIHDHEFKAKVLSLDNGEQRDVGFYDIHHCIFCDEQTSVY 250
Query: 218 LRDSLNWLGLRSYVDDCDDYDCEYITSIEQFMIVALDLRTETCKELLLPRGFDEVPCYEP 277
SL WL L+ S + ++A+DL TE + +LLP D
Sbjct: 251 ANGSLFWLTLKKL-------------SQTSYQLLAIDLHTEEFRWILLPE-CDTKYATNI 296
Query: 278 SLCVLMDCICFSHLVKKTHLVIWKMMDYGDDDSWTQLLEINLQILK 323
+ L + +C S +++ ++LV+W + + W ++ I + +++
Sbjct: 297 EMWNLNERLCLSDVLESSNLVVWSLHQEYPTEKWEKIYSIKIDVIR 342
>At3g49980 putative protein
Length = 382
Score = 63.5 bits (153), Expect = 2e-10
Identities = 82/322 (25%), Positives = 134/322 (41%), Gaps = 53/322 (16%)
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESREC 81
LP +L+ E+L +P SL Q R C W L +D F + H ++ + + V +E C
Sbjct: 5 LPQELLEEILCRVPATSLKQLRLTCKEWNRLFNDRTFSRKHFDKAPKQFLITVLEE--RC 62
Query: 82 SMDVIVSPTSLSHLLEN--PSKPTT--LTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSED 137
+ +SLS L + PS+ T L+ Y+S + + C+GL L +
Sbjct: 63 RL------SSLSINLHSGFPSEEFTGELSPIDYHSNSSQVIIMKIFHCDGLFVCTILKDT 116
Query: 138 REMWLRFWNPATR-----AISDKLGHYPADFTGGFEVAFGYDNSTD-TYKVVYLQKGMTG 191
R + WNP T + L DF G+ + + S+D +YK++ + G
Sbjct: 117 R---IVVWNPCTGQKKWIQTGENLDENGQDFVLGY---YQDNKSSDKSYKILSYKGYNYG 170
Query: 192 -----VFSLGDNVWRNIESFPL-GYYL---DNRVHLRDSLNWLGLRSYVDDCDDYDCEYI 242
++ + N WRN++ P+ G Y D RV L+ + W Y
Sbjct: 171 DQEFKIYDIKSNTWRNLDVTPIPGNYFTCSDYRVSLKGNTYWFA--------------YD 216
Query: 243 TSIEQFMIVALDLRTETCKELLLPRGFDEVPCYEPSLCVLMD--CICFSHLVKKTHLVIW 300
EQ +++ D TE + L LP D + ++ SL V+ + L IW
Sbjct: 217 LKDEQLGLISFDYTTERFERLWLPFQCD-ISDHDYSLSVVGEEKLSVVLQLKDAPRREIW 275
Query: 301 ---KMMDYGDDDSWTQLLEINL 319
KM D + SW +L E+ +
Sbjct: 276 ITNKMDDETKEMSWRKLFEVEV 297
>At4g38870 putative protein
Length = 426
Score = 63.2 bits (152), Expect = 2e-10
Identities = 75/291 (25%), Positives = 130/291 (43%), Gaps = 42/291 (14%)
Query: 13 SNVASSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRL 72
+ V+ + +LP+DLI E+L L +K L++F CV W +I D +F+KL L S++ P+
Sbjct: 44 TKVSVNSELLPVDLIMEILKKLSLKPLIRFLCVSKLWASIIRDPYFMKLFLNESLKRPK- 102
Query: 73 AVTQESRECSMDVIVSPTSLSHLLE--NPSKPTTLTNDPYY-SLNDKDCRSVAGSCNGLL 129
++ R S+ I S L E + S ++ ++ Y+ + + +++ S +GL+
Sbjct: 103 SLVFVFRAQSLGSIFSSVHLKSTREISSSSSSSSASSITYHVTCYTQQRMTISPSVHGLI 162
Query: 130 CLLGLSEDREMWLRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNSTDTYKVVYLQKGM 189
C S L +NP TR S L A GYD YKVV + +GM
Sbjct: 163 CYGPPSS-----LVIYNPCTRR-SITLPKIKAG-RRAINQYIGYDPLDGNYKVVCITRGM 215
Query: 190 ------------TGVFSLG--DNVWRNI-ESFPLGYYLDNRVHLRDSLNWLGLRSYVDDC 234
V +LG D+ WR I + P + + + L + R+++
Sbjct: 216 PMLRNRRGLAEEIQVLTLGTRDSSWRMIHDIIPPHSPVSEELCINGVLYY---RAFIG-- 270
Query: 235 DDYDCEYITSIEQFMIVALDLRTETCKELLLP---RGFDEVPCYEPSLCVL 282
T + + I++ D+R+E + +P R F ++ YE L V+
Sbjct: 271 --------TKLNESAIMSFDVRSEKFDLIKVPCNFRSFSKLAKYEGKLAVI 313
>At3g16210 hypothetical protein
Length = 360
Score = 61.2 bits (147), Expect = 9e-10
Identities = 63/221 (28%), Positives = 91/221 (40%), Gaps = 32/221 (14%)
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESREC 81
LP +L E+L L +K L +FRCVC +W+ LI+D F T+ R+
Sbjct: 5 LPEELAIEILVRLSMKDLARFRCVCKTWRDLINDPGF----------------TETYRDM 48
Query: 82 SMDVIVSPTSLS-HLLENPSKPTTLTNDPYYSLNDK--DCRSVAGSCNGLLCLLGLSEDR 138
S VS + ++L+ K +TN + L+ D + C+G LC+ +
Sbjct: 49 SPAKFVSFYDKNFYMLDVEGKHPVITNKLDFPLDQSMIDESTCVLHCDGTLCV----TLK 104
Query: 139 EMWLRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNSTDTYKVV----YLQKGMTGVFS 194
L WNP ++ K+ P + + FGYD D YKVV L VF
Sbjct: 105 NHTLMVWNPFSKQF--KIVPNPGIYQDSNILGFGYDPVHDDYKVVTFIDRLDVSTAHVFE 162
Query: 195 LGDNVWRNI--ESFPLGYYLDNR-VHLRDSLNWLGLRSYVD 232
W S+P +Y D R L L W+ RS D
Sbjct: 163 FRTGSWGESLRISYPDWHYRDRRGTFLDQYLYWIAYRSSAD 203
>At5g10340 putative protein
Length = 445
Score = 60.8 bits (146), Expect = 1e-09
Identities = 84/344 (24%), Positives = 135/344 (38%), Gaps = 68/344 (19%)
Query: 21 ILPLDLIT-ELLSFLPVKSLLQFRCVCMSWKILISDSFFVKL----HLQRSIQNPRLAVT 75
+LP D+I ++ L VK+LL+F+ V W I F + HL +S +P + +
Sbjct: 68 LLPHDVIEYHIMVRLDVKTLLKFKSVSKQWMSTIQSPSFQERQLIHHLSQSSGDPHVLLV 127
Query: 76 QESRECSMDVIVSPTSL----SHLLENPSK----PTTLTNDPYYSLNDKDCRSVAGSCNG 127
C+ S +S + L+E+ + PT + Y+ N SC+G
Sbjct: 128 SLYDPCARQQDPSISSFEALRTFLVESSAASVQIPTPWEDKLYFVCNT--------SCDG 179
Query: 128 LLCLLGLSEDREMWLRFWNPATR------------AISDKLGHYPADFTGGFEVAFGYDN 175
L+CL E + + NP TR +DK + FG D
Sbjct: 180 LICLFSFYELPSIVV---NPTTRWHRTFPKCNYQLVAADKGERHECFKVACPTPGFGKDK 236
Query: 176 STDTYKVVYLQKG----------MTGVFSLGDNVWRNIESFPLGYYL----DNRVHLRDS 221
+ TYK V+L VF N WR + FP +L + V++ S
Sbjct: 237 ISGTYKPVWLYNSAELDLNDKPTTCEVFDFATNAWRYV--FPASPHLILHTQDPVYVDGS 294
Query: 222 LNWLGLRSYVDDCDDYDCEYITSIEQFMIVALDLRTETCKELLLPRGFDEVPCYEPSLCV 281
L+W S+ + M+++LDL +ET + + + Y +C
Sbjct: 295 LHWFTALSHEGET--------------MVLSLDLHSETFQVISKAPFLNVSDEYYIVMCN 340
Query: 282 LMDCICFSHLVKKTHLVIWKMMDYGDDDSWTQLLEINLQILKKI 325
L D +C S K + VIW +D D +W Q+ I+L I +
Sbjct: 341 LGDRLCVSE-QKWPNQVIWS-LDDSDHKTWKQIYSIDLIITSSL 382
>At1g62270 hypothetical protein
Length = 383
Score = 60.5 bits (145), Expect = 2e-09
Identities = 67/255 (26%), Positives = 109/255 (42%), Gaps = 37/255 (14%)
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESREC 81
LP DL+ ++L+ +P SL + R C W L +D F K+H ++ + + + + C
Sbjct: 12 LPWDLVEDILARVPATSLKRLRSTCKQWNFLFNDQIFTKMHFDKAEKQFLVLILRLYTVC 71
Query: 82 SMDVIVSPTSLSHLLEN--PS---KPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSE 136
SM + L L +N PS K DP+ S + K S CNGLL ++
Sbjct: 72 SMSL-----DLRGLHDNIDPSIEVKGELSLIDPHCS-SRKTFVSKVFHCNGLLLCTTMT- 124
Query: 137 DREMWLRFWNPA---TRAISDKLGHYPADFTGGFEVAFGYDNSTDTY-----KVVYLQKG 188
L WNP TR I ++ H D + A GY N Y K + L+
Sbjct: 125 ----GLVVWNPCTDQTRWIKTEVPHNRND-----KYALGYGNYKSCYNYKIMKFLDLESF 175
Query: 189 MTGVFSLGDNVWRNIESFPLGYYLD---NRVHLRDSLNWLG--LRSYVDDCDDYDCEYIT 243
++ + N WR + + + + V LR + W+ R +++ ++ + EY
Sbjct: 176 DLEIYEVNSNSWRVLGTVTPDFTIPLDAEGVSLRGNSYWIASHKREEIEEEEEEENEYF- 234
Query: 244 SIEQFMIVALDLRTE 258
I F+I + D TE
Sbjct: 235 -INDFLI-SFDFTTE 247
>At2g07140 pseudogene
Length = 384
Score = 59.3 bits (142), Expect = 3e-09
Identities = 62/219 (28%), Positives = 100/219 (45%), Gaps = 24/219 (10%)
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSW-KILISDSFFVKLHLQRSI-QNPRLAVTQESR 79
LP DL+ E+LSF+P SL + R C W ++ D F ++H +++ Q L +T+ R
Sbjct: 6 LPKDLVEEILSFVPATSLKRLRSTCKGWNRLFKDDKRFTRIHTEKAAKQFQPLTLTKNYR 65
Query: 80 ECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSV--AGSCNGLLCLLGLSED 137
C ++V + T+ S LE ++ + L DP +S N ++ C+GLL +
Sbjct: 66 ICPINVNLHGTTPS--LEVKNEVSLL--DP-HSKNSAAQFNIDRVFHCDGLLLCTSQKDS 120
Query: 138 REMWLRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNST--DTYKVV---YLQKGMTGV 192
R + WNP T K + G GYDN + +YK + YL K + +
Sbjct: 121 RFV---VWNPLTGV--TKWIELGDRYNEGMAFILGYDNKSCNKSYKAMSFNYLDKD-SEI 174
Query: 193 FSLGDNVWRNIESF--PLGY--YLDNRVHLRDSLNWLGL 227
+ + WR I+ P Y Y L+ + WLG+
Sbjct: 175 YEFSSDSWRVIDDIIKPPHYMDYFRECFSLKGNTYWLGI 213
>At3g51170 putative protein
Length = 346
Score = 58.5 bits (140), Expect = 6e-09
Identities = 71/274 (25%), Positives = 112/274 (39%), Gaps = 57/274 (20%)
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESREC 81
LP +L ++LS L +SL++FR VC W L + F+K H R+ P+ +S
Sbjct: 4 LPWELEEDILSRLAAQSLVRFRSVCKRWNYLFDEKSFIKNHFARAC--PQFIFMTDSNIY 61
Query: 82 SMDVI----VSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSED 137
S+++I V PT +L++ P Y ++ +C+G L +
Sbjct: 62 SIEIIGLDGVDPTIKLRVLDSSGIPYRDWKFAYITIT---------ACDGFLFCNSWASP 112
Query: 138 REMWLRFWNPATRAISDKLGHYPADFTGGFEV-AFGYDNSTD--TYKVV----------- 183
+ FWNP + + K Y GF V GYDN+ D YK++
Sbjct: 113 K--GTAFWNPWLKQV--KWIEYE---DKGFNVCGLGYDNTRDGKVYKILGYLMCRPEDGS 165
Query: 184 -YLQKGMTGVFSLGDNVWRNIESFPLGYYLDN----RVHLRDSLNWLGLRSYVDDCDDYD 238
Y+ ++ + ++ IE ++L + V L +L WL S + + DD
Sbjct: 166 SYVYHQRVAIYDCASHAFKFIEISNKDWFLTDVTRFSVSLNGNLYWL---SPIPNTDD-- 220
Query: 239 CEYITSIEQFMIVALDLRTETCKEL-LLP-RGFD 270
F+I D E CK LLP R FD
Sbjct: 221 ---------FLIQCFDFSREICKPFCLLPCRKFD 245
>At3g22700 hypothetical protein
Length = 338
Score = 58.5 bits (140), Expect = 6e-09
Identities = 70/268 (26%), Positives = 114/268 (42%), Gaps = 55/268 (20%)
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESREC 81
LPLDL+ E+LS +P SL + R C W L+ D F + H +++ P+ ++ +E
Sbjct: 4 LPLDLVEEILSRVPATSLKRLRSTCRQWNALLKDRRFTEKHFRKA---PKESLVLMLKEI 60
Query: 82 SMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSEDREMW 141
S+++ V+P S+ K D + + D V C+GLL L ++R +
Sbjct: 61 SVNLNVTPPSIEF------KDALGLKDSHSNSEQVDIVQVL-HCDGLL-LCTTKDNRHV- 111
Query: 142 LRFWNPA---TRAISDKL--GHYPADFTGGFEVAFGYDNSTDTYKVVY-----------L 185
WNP T I K+ G + F G+ + S +YK+++
Sbjct: 112 --VWNPCLGETHWIQFKVDYGRVYSSFALGY---IQNNESCRSYKILWRWKSNDYKSSPR 166
Query: 186 QKGMTGVFSLGDNVWR-----NIESFPLGYYLDN--RVHLRDSLNWLGLRSYVDDCDDYD 238
Q+G ++ + WR N +S YL RV L+ ++ WL VDD +D
Sbjct: 167 QRGFE-IYEFISDKWRVIDDVNHDSLVNHNYLGRCCRVSLKGNIYWL-----VDDVED-- 218
Query: 239 CEYITSIEQFMIVALDLRTETCKELLLP 266
++ D +TE K L LP
Sbjct: 219 -------NSRSLLMFDFKTERFKRLCLP 239
>At3g17280 hypothetical protein
Length = 386
Score = 58.5 bits (140), Expect = 6e-09
Identities = 71/314 (22%), Positives = 120/314 (37%), Gaps = 43/314 (13%)
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESREC 81
LP DL+ E+LS LP KS+ + + C W L D FV+ L ++ + + E
Sbjct: 7 LPYDLLPEILSRLPTKSIPKLKTTCKKWYALFKDPKFVEKKLGKAARETVFLMNHEVNSI 66
Query: 82 SMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSEDREMW 141
S+D+ P S ++ T +D + + CNGL L
Sbjct: 67 SVDIHGIPKGYSVSMDFTGTLTIPEG------SDLEIFRI-HHCNGLF----LCATMNCR 115
Query: 142 LRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNSTD--TYKVVYL-----QKGMTGVFS 194
L WNP T I+ + D + + G D S+ +YK++ +K ++ ++
Sbjct: 116 LVVWNPCTGQITWIIPRTRYDSDDIYALGCGDDKSSSLHSYKILRCCDDNQKKPVSEIYD 175
Query: 195 LGDNVWRNIESFPLGYYLD-NRVHLRDSLNWLGLRSYVDDCDDYDCEYITSIEQFMIVAL 253
+ WR ++ +++ N V L++S W Y + + + I+
Sbjct: 176 FSSSSWRVLDGVTANCFIECNGVALKESAYW------------YASDKRETPKGKFILRF 223
Query: 254 DLRTETCKELLLPRGFDE---------VPCYEPSLCVLMDCICFSHLVKKTHLVIW---K 301
D TE L LP F E L +L H +K + + IW
Sbjct: 224 DFATERFARLCLPLNFQRDRDNKSVVVSVVGEEKLALLQQFDHRVHGLKYSKIKIWVTDT 283
Query: 302 MMDYGDDDSWTQLL 315
+ G D SW+ +L
Sbjct: 284 KIGEGKDLSWSNIL 297
>At2g21930 unknown protein
Length = 396
Score = 58.2 bits (139), Expect = 8e-09
Identities = 65/212 (30%), Positives = 93/212 (43%), Gaps = 35/212 (16%)
Query: 16 ASSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPR---L 72
+ + + +P DLI E+L LPVK+L +F V + +I + F+K +L S P+
Sbjct: 19 SGNSVQIPFDLIPEILKRLPVKTLARFLSVSKEYTSIIRNRDFMKSYLINSSTRPQSLIF 78
Query: 73 AVTQESRECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLC-- 130
+ C +I S S SKPT L N P+ L ++ A S +GL+C
Sbjct: 79 TIAGGGIHCFFSLIDQGESTS-----SSKPTYLMNCPHLQL-----KTFAPSVHGLICHG 128
Query: 131 ---LLGLSEDREMWLRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNSTDTYKVVYLQK 187
L +S R L NP+TR S L A+ + GYD YKV+ + K
Sbjct: 129 PPSTLIVSSPR---LIVSNPSTRR-SIILPKIDANHECIYH-HMGYDPIDGDYKVLCMMK 183
Query: 188 GM-----------TGVFSL-GDNVWRNIESFP 207
GM VF+L N WR +E FP
Sbjct: 184 GMHVYQRRYLAKELQVFTLRKGNSWRMVEDFP 215
>At3g23260 unknown protein
Length = 362
Score = 57.8 bits (138), Expect = 1e-08
Identities = 82/323 (25%), Positives = 131/323 (40%), Gaps = 56/323 (17%)
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESREC 81
LP++L E+LS +P K L + R W L F K H + + P + + ++SR
Sbjct: 6 LPVELQEEILSRVPAKYLARLRSTSKQWNALSKTGSFAKKHSANATKEPLIIMLKDSR-- 63
Query: 82 SMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSEDREMW 141
V ++ +L + N ++ L + Y L D +V C+GLL L + E+
Sbjct: 64 ---VYLASVNLHGVHNNVAQSFELGSRLY--LKDPHISNVF-HCDGLLLLCSIKENT--- 114
Query: 142 LRFWNPAT---RAISDKLGHY-PADFTGGFEVAFGYDN--STDTYKV--VYLQKGMTG-- 191
L WNP + + I + +Y +DF A GYDN S YKV V Q + G
Sbjct: 115 LEVWNPCSGEAKLIKPRHSYYKESDF-----YALGYDNKSSCKKYKVLRVISQVHVQGDF 169
Query: 192 -----VFSLGDNVWR-NIESFPLGYYLDNRVHLRDSLNWLGLRSYVDDCDDYDCEYITSI 245
++ ++ WR + + L + V ++ S W+ Y + +Y S
Sbjct: 170 KIEYEIYDFTNDSWRVHGATTELSIRQKHPVSVKGSTYWVVRNRY------FPYKYFLS- 222
Query: 246 EQFMIVALDLRTETCKELLLPRGF-----DEVPCYEPSLCVLMDCICFSHLVKKTHLVIW 300
D TE + L LP+ F D E LC+ ++ L +W
Sbjct: 223 -------FDFSTERFQSLSLPQPFPYLVTDLSVVREEQLCLFG---YYNWSTTSEDLNVW 272
Query: 301 KMMDYGDDDSWTQLLEINLQILK 323
G SW++ L I QI+K
Sbjct: 273 VTTSLGSVVSWSKFLTI--QIIK 293
>At3g10240 hypothetical protein
Length = 389
Score = 57.8 bits (138), Expect = 1e-08
Identities = 54/207 (26%), Positives = 92/207 (44%), Gaps = 22/207 (10%)
Query: 7 KSQQPPSNVASSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRS 66
KS++ S +S +PLDL++E+L LP KS+ +FRCV W + ++ +F+ L RS
Sbjct: 16 KSKRSKSGSSS----IPLDLVSEILLRLPEKSVARFRCVSKPWSSITTEPYFINLLTTRS 71
Query: 67 IQNPRLAVTQESRECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCR--SVAGS 124
PRL + ++ E + + N S + D Y+ ++ S
Sbjct: 72 ---PRLLLCFKANEKFFVSSIPQHRQTFETWNKSHSYSQLIDRYHMEFSEEMNYFPPTES 128
Query: 125 CNGLLCLLGLSEDREMWLRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNSTDTYKVVY 184
NGL+C L WNP+TR + + P + + GYD +KV+
Sbjct: 129 VNGLICF-----QESARLIVWNPSTRQL--LILPKPNGNSNDLTIFLGYDPVEGKHKVMC 181
Query: 185 LQKGMT----GVFSLG--DNVWRNIES 205
++ T V +LG +WR +++
Sbjct: 182 MEFSATYDTCRVLTLGSAQKLWRTVKT 208
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.139 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,940,227
Number of Sequences: 26719
Number of extensions: 451495
Number of successful extensions: 1545
Number of sequences better than 10.0: 261
Number of HSP's better than 10.0 without gapping: 236
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 1224
Number of HSP's gapped (non-prelim): 298
length of query: 389
length of database: 11,318,596
effective HSP length: 101
effective length of query: 288
effective length of database: 8,619,977
effective search space: 2482553376
effective search space used: 2482553376
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147202.5


