

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.9 + phase: 1 /partial
(375 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
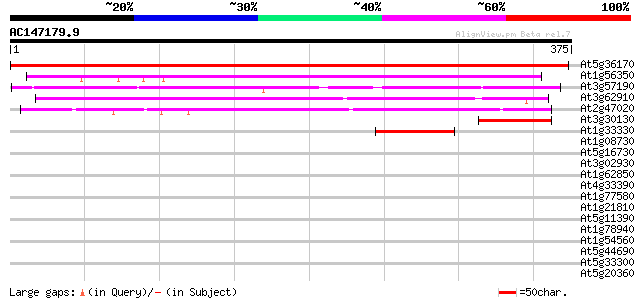
Score E
Sequences producing significant alignments: (bits) Value
At5g36170 translation releasing factor RF-2 613 e-176
At1g56350 hypothetical protein 230 1e-60
At3g57190 unknown protein 223 2e-58
At3g62910 translation releasing factor RF-1 -like protein 162 2e-40
At2g47020 putative peptide chain release factor 128 4e-30
At3g30130 hypothetical protein 57 2e-08
At1g33330 peptide chain release factor, putative 54 1e-07
At1g08730 myosin MYA1, class V (Z28389) like protein 38 0.010
At5g16730 putative protein 37 0.018
At3g02930 unknown protein 35 0.087
At1g62850 hypothetical protein 34 0.11
At4g33390 putative protein 34 0.15
At1g77580 unknown protein 34 0.15
At1g21810 myosin-like protein 33 0.19
At5g11390 unknown protein 32 0.43
At1g78940 unknown protein 32 0.57
At1g54560 32 0.74
At5g44690 putative protein 31 0.97
At5g33300 putative protein 31 1.3
At5g20360 tetratricopeptide repeat protein 30 2.2
>At5g36170 translation releasing factor RF-2
Length = 456
Score = 613 bits (1581), Expect = e-176
Identities = 300/373 (80%), Positives = 344/373 (91%)
Query: 1 FYSLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADV 60
FY+LRKDVEIAS RV+EIR S+ LQ LEQE+ LE +A+ +SFWDDR KAQ+TLS+L D+
Sbjct: 81 FYTLRKDVEIASARVEEIRASANLQQLEQEITNLESKATDTSFWDDRTKAQETLSSLNDL 140
Query: 61 KEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSG 120
K++++LL+++KT VEDAETIV LTEEM+S D L EEA +IKELNKS+D+FELTQLLSG
Sbjct: 141 KDRMRLLSEFKTMVEDAETIVKLTEEMDSTDVSLLEEAMGIIKELNKSLDKFELTQLLSG 200
Query: 121 PYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATI 180
PYDKEGAV+ ITAGAGGTDAQDWADMLLRMY+RWGEKQRYKT+VVE S GEEAGIKSAT+
Sbjct: 201 PYDKEGAVVYITAGAGGTDAQDWADMLLRMYMRWGEKQRYKTKVVEMSNGEEAGIKSATL 260
Query: 181 EVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLE 240
E+EGRYAYGY+SGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEE++ +EIPEEDL+
Sbjct: 261 EIEGRYAYGYISGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEEAVGIEIPEEDLD 320
Query: 241 ISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAE 300
ISF+RAGGKGGQNVNKVETAVRITHIPTGV +RCTEERSQLANK +AL RLKAKL+VIAE
Sbjct: 321 ISFTRAGGKGGQNVNKVETAVRITHIPTGVAVRCTEERSQLANKTRALIRLKAKLMVIAE 380
Query: 301 EQRATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKS 360
EQRATE K+IRGD VKAEWGQQIRNYVFHPYKLVKDVRTGHET DITSV+DG+LDPFIK+
Sbjct: 381 EQRATEIKEIRGDAVKAEWGQQIRNYVFHPYKLVKDVRTGHETSDITSVMDGDLDPFIKA 440
Query: 361 YLKHKYSMTMSTS 373
YLKHKY++ M+++
Sbjct: 441 YLKHKYTLAMASA 453
>At1g56350 hypothetical protein
Length = 524
Score = 230 bits (586), Expect = 1e-60
Identities = 133/370 (35%), Positives = 204/370 (54%), Gaps = 26/370 (7%)
Query: 12 SQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDD--------------RAKAQQTLSTL 57
+Q ++ I+ + L L L E + S WDD K + ++
Sbjct: 120 AQSIQVIKRRLQWKKLLVRLKVLSAELNKSDLWDDPTHAGKISREHGSLTGKMKGVMTFE 179
Query: 58 ADVKEKIKLLNDYK----TQVEDAETIVMLTEEME------SVDKGLYEEASSL--IKEL 105
++ E I +L K +++E + ++++ + SV G+ +L + ++
Sbjct: 180 RELLEHIDMLKLAKEENDSELESVGVLALISQTLGMFSFSVSVSLGMINPKETLKALIDM 239
Query: 106 NKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVV 165
+ EL LLS D I + AGAGGT++ DWA M++ MY W +++++ VV
Sbjct: 240 RRVSKEKELEALLSADNDPCSCYIEVQAGAGGTESNDWAAMVMEMYKTWAQRRKFSVTVV 299
Query: 166 EKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLL 225
+++ GE AGIK ATI+V G YAYGY E G HR+VR SPF+S R TSF+ V V+P+L
Sbjct: 300 DEAPGEIAGIKRATIKVNGEYAYGYAKAEVGVHRLVRISPFDSGKRRHTSFAAVAVIPIL 359
Query: 226 PEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKI 285
+ S VEI + DL I R+GG GGQ+ N ++AVRI HIPTG+T C ERSQ +NK
Sbjct: 360 GDGSTRVEINDSDLRIERFRSGGAGGQHANTTDSAVRIVHIPTGITATCQNERSQHSNKA 419
Query: 286 KALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPD 345
A++ L+++L + ++ Q + + WG QIR YV HPY++VKD+RT +E D
Sbjct: 420 SAMAVLQSRLDQLEMARQTAMNAQHTQSLTEISWGNQIRTYVLHPYRMVKDLRTNYEVSD 479
Query: 346 ITSVIDGELD 355
SV++G+LD
Sbjct: 480 PDSVLEGDLD 489
>At3g57190 unknown protein
Length = 406
Score = 223 bits (567), Expect = 2e-58
Identities = 131/369 (35%), Positives = 218/369 (58%), Gaps = 15/369 (4%)
Query: 2 YSLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVK 61
+SL+K ++ + E+ L+L E++ K EE WDD AK+ + L LAD
Sbjct: 48 FSLKKKIKDVVLKA-EMFAPDALELEEEQWIKQEETMRYFDLWDDPAKSDEILLKLADRA 106
Query: 62 EKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGP 121
+ + L D K + E+A+ I+ L E M+++D L+E+A ++++S+ +E+++LL
Sbjct: 107 KAVDSLKDLKYKAEEAKLIIQLGE-MDAIDYSLFEQAYDSSLDVSRSLHHYEMSKLLRDQ 165
Query: 122 YDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKS--LGEEAGIKSAT 179
YD EGA + I +G+ G +Q W + ++ MY++W E+ RV EK L ++G+ SAT
Sbjct: 166 YDAEGACMIIKSGSPGAKSQIWTEQVVSMYIKWAERLGQNARVAEKCSLLSNKSGVSSAT 225
Query: 180 IEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDL 239
IE E +AYGYL GE+G HR++ S N + + V+++PL S + E+ E DL
Sbjct: 226 IEFEFEFAYGYLLGERGVHRLIISSTSN-----EECSATVDIIPLFLRASPDFEVKEGDL 280
Query: 240 EISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIA 299
+S+ ++ E V I HIP+GVTL+ + ER++ AN+IKAL+RLKAKLLVIA
Sbjct: 281 IVSY-----PAKEDHKIAENMVCIHHIPSGVTLQSSGERNRFANRIKALNRLKAKLLVIA 335
Query: 300 EEQRATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIK 359
+EQ+ ++ +I + E ++ R+YV +K+V D +TG E D+ SV+DG + P +
Sbjct: 336 KEQKVSDVNKIDSKNI-LEPREETRSYVSKGHKMVVDRKTGLEILDLKSVLDGNIGPLLG 394
Query: 360 SYLKHKYSM 368
+++ + S+
Sbjct: 395 AHISMRRSI 403
>At3g62910 translation releasing factor RF-1 -like protein
Length = 422
Score = 162 bits (411), Expect = 2e-40
Identities = 101/346 (29%), Positives = 177/346 (50%), Gaps = 9/346 (2%)
Query: 18 IRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDA 77
IR+ ++ +EL+ + S + K Q++S L +V + D + Q+ ++
Sbjct: 58 IRKMESVEKTWKELSVKLADPDVVSNQSEYQKLAQSMSELDEVVTVFRRFKDCEKQLLES 117
Query: 78 ETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGG 137
+ + + E + + + E +SL KE+ + + ++ L S P D ++ + AG GG
Sbjct: 118 KVLAKEAGDDEDMAEMIGSEINSLTKEIEELEKQLKVLLLPSDPLDARNILLEVRAGTGG 177
Query: 138 TDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGT 197
+A W L+RMY R+ E+ +K +V S E G K+ +E++G Y L E G
Sbjct: 178 DEAAIWTGDLVRMYQRYSERSSWKFSMVSCSEAEHGGYKTCVMEIKGNRVYSKLKYESGV 237
Query: 198 HRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKV 257
HR+ R ++G TS + V +MP + + V I +D+E++ +R+GG GGQNVNKV
Sbjct: 238 HRVQRVPQTETQGRVHTSTATVAIMP--EADEVEVVIDPKDIELTSARSGGAGGQNVNKV 295
Query: 258 ETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKA 317
ETA+ + H P+G+ + CTEER+Q+ NK +A L+AKL I ++ + + R K+
Sbjct: 296 ETAIDLFHKPSGIRIFCTEERTQIRNKARAFQLLRAKLYEIKVREQQEKIRNER----KS 351
Query: 318 EWGQQIRNYVFHPYKLVKDVRTGHETP---DITSVIDGELDPFIKS 360
+ G R+ Y T H +T+ +DG L+ +++
Sbjct: 352 QVGTGARSEKIRTYNYKDSRVTDHRLKMNFALTTFLDGALEDAVQA 397
>At2g47020 putative peptide chain release factor
Length = 395
Score = 128 bits (322), Expect = 4e-30
Identities = 109/377 (28%), Positives = 180/377 (46%), Gaps = 28/377 (7%)
Query: 8 VEIASQRVKEIRESSG-LQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKL 66
++I QR+ I + LQ L + EE S ++ + K + ++ + D++ K K+
Sbjct: 10 IKIMDQRLSAIEHRNAVLQKLINQPEYSPEEFSRAN--KELRKLRDSMLLINDLRAKQKV 67
Query: 67 LN-----DYKTQVEDAETIVMLTEEMESVDKGLYEEASS-------LIKELNKSIDRFEL 114
+N Y Q +E++ + K L E+S + EL+++++ +
Sbjct: 68 VNLLYRIIYLFQRSSLIRFTFCWQEIDGL-KSLVSESSDDKDMLDLAVGELDEAVEEEKR 126
Query: 115 TQLL-------SGPYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEK 167
Q L D+ ++ + AG GG +A +A + RMY R+ +K+ +K +V+
Sbjct: 127 LQTLLLKSLLPKDEADERDCILEVRAGTGGEEASLFAMDIFRMYERYSQKKGWKFDIVDI 186
Query: 168 SLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPE 227
+ + G K A+ + G YG L E G HR+ R G TS V ++P E
Sbjct: 187 TESDMKGYKEASAAICGASVYGKLKFESGIHRVQRIPITEKSGRIHTSAISVAILPQADE 246
Query: 228 ESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKA 287
++V++ EDL I R+GG GGQ+ N +AVRI H+PTG+ + +ERSQ N+ KA
Sbjct: 247 --VDVQLRNEDLRIDTYRSGGSGGQHANTTNSAVRIIHLPTGMMVSIQDERSQHMNRAKA 304
Query: 288 LSRLKAKLLVIAEEQRATEFKQIRGDVV-KAEWGQQIRNYVFHPYKLVKDVRTGHETPDI 346
L L A+L I + + ++R D + + +IR Y F P V D R G I
Sbjct: 305 LKVLCARLYEIERLRIQSSRSKLRSDQIGSGDRSGRIRTYNF-PQGRVTDHRVGITHHAI 363
Query: 347 TSVIDGE-LDPFIKSYL 362
++ GE LD FI + L
Sbjct: 364 EDMMQGENLDMFIDALL 380
>At3g30130 hypothetical protein
Length = 82
Score = 57.0 bits (136), Expect = 2e-08
Identities = 24/49 (48%), Positives = 33/49 (66%)
Query: 314 VVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYL 362
+ + WG QIR YV H Y++VKD+RT +E D SV++G+LD FI L
Sbjct: 27 LTEISWGNQIRTYVLHQYRMVKDLRTNYEVSDPDSVLEGDLDGFISKLL 75
>At1g33330 peptide chain release factor, putative
Length = 257
Score = 54.3 bits (129), Expect = 1e-07
Identities = 24/53 (45%), Positives = 36/53 (67%)
Query: 245 RAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLV 297
R G GGQ+ NK ++AVR+ H+PTG+ + E+RSQ N+ AL+RL+ L +
Sbjct: 104 RVSGPGGQHRNKRDSAVRLKHLPTGIVAQAVEDRSQHKNRASALNRLRTLLAI 156
>At1g08730 myosin MYA1, class V (Z28389) like protein
Length = 1572
Score = 37.7 bits (86), Expect = 0.010
Identities = 37/133 (27%), Positives = 61/133 (45%), Gaps = 28/133 (21%)
Query: 4 LRKDVEIASQRVKEIRESSG--LQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVK 61
L K VE + RV+ + S G + QE+ KL+ SSF + R K +T + L +
Sbjct: 936 LEKKVEELTYRVQLEKRSRGDLEEAKTQEILKLK-----SSFEEMRKKVDETNALLLKER 990
Query: 62 EKIK--------LLNDYKTQVEDAETIVMLTEEMESVDKGL-------------YEEASS 100
E K ++ + + VED + I ++TEE+ESV L +EEA
Sbjct: 991 EAAKKAAEEAPPVIKETQILVEDTKKIELMTEELESVKVTLENEKQRADDAVRKFEEAQE 1050
Query: 101 LIKELNKSIDRFE 113
+++ K ++ E
Sbjct: 1051 SLEDKKKKLEETE 1063
>At5g16730 putative protein
Length = 853
Score = 37.0 bits (84), Expect = 0.018
Identities = 29/114 (25%), Positives = 57/114 (49%), Gaps = 4/114 (3%)
Query: 3 SLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKE 62
+L+K+ + A+ RV+ + E L + E +K EEE S + + + S ++KE
Sbjct: 427 ALKKEQD-ATSRVQRLSEEKSKLLSDLESSKEEEEKSKKAMESLASALHEVSSEGRELKE 485
Query: 63 KIKLLND--YKTQVEDAETIVMLT-EEMESVDKGLYEEASSLIKELNKSIDRFE 113
K+ D Y+TQ++D + ++ T E+ E++ E L+ + ++ FE
Sbjct: 486 KLLSQGDHEYETQIDDLKLVIKATNEKYENMLDEARHEIDVLVSAVEQTKKHFE 539
Score = 34.3 bits (77), Expect = 0.11
Identities = 29/98 (29%), Positives = 51/98 (51%), Gaps = 8/98 (8%)
Query: 5 RKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRA--KAQQTLSTLADV-K 61
++D+E++ QR+ + E E+E+ KL+ E +RA K Q S + + +
Sbjct: 386 KEDLEVSEQRLGSVEEEVSKN--EKEVEKLKSELETVKEEKNRALKKEQDATSRVQRLSE 443
Query: 62 EKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEAS 99
EK KLL+D ++ E+ E + MES+ L+E +S
Sbjct: 444 EKSKLLSDLESSKEEEE---KSKKAMESLASALHEVSS 478
>At3g02930 unknown protein
Length = 806
Score = 34.7 bits (78), Expect = 0.087
Identities = 29/114 (25%), Positives = 56/114 (48%), Gaps = 4/114 (3%)
Query: 3 SLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKE 62
+L+K+ + A+ V+ + E L E E +K EEE S + + + S ++KE
Sbjct: 416 ALKKEQD-ATSSVQRLLEEKKKILSELESSKEEEEKSKKAMESLASALHEVSSESRELKE 474
Query: 63 KIKLLND--YKTQVEDAETIVMLT-EEMESVDKGLYEEASSLIKELNKSIDRFE 113
K+ D Y+TQ+ED + ++ T + E++ E L+ + ++ +FE
Sbjct: 475 KLLSRGDQNYETQIEDLKLVIKATNNKYENMLDEARHEIDVLVNAVEQTKKQFE 528
Score = 30.0 bits (66), Expect = 2.2
Identities = 29/113 (25%), Positives = 57/113 (49%), Gaps = 14/113 (12%)
Query: 7 DVEIASQRVKEIRESSGLQLLEQELAKLEEEA-----SCSSFWDDRAKA-----QQTLST 56
++ +ASQ+V + L + E+E +K E+EA + +++ +A T S
Sbjct: 368 EMTVASQKVDLEKSEQKLGIAEEESSKSEKEAEKLKNELETVNEEKTQALKKEQDATSSV 427
Query: 57 LADVKEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSI 109
++EK K+L++ ++ E+ E + MES+ L+ E SS +EL + +
Sbjct: 428 QRLLEEKKKILSELESSKEEEE---KSKKAMESLASALH-EVSSESRELKEKL 476
>At1g62850 hypothetical protein
Length = 143
Score = 34.3 bits (77), Expect = 0.11
Identities = 13/25 (52%), Positives = 21/25 (84%)
Query: 237 EDLEISFSRAGGKGGQNVNKVETAV 261
+++ ++F+R+GG GGQNVNK+ T V
Sbjct: 9 DNVTLNFARSGGPGGQNVNKLNTKV 33
>At4g33390 putative protein
Length = 779
Score = 33.9 bits (76), Expect = 0.15
Identities = 28/100 (28%), Positives = 48/100 (48%), Gaps = 3/100 (3%)
Query: 9 EIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQT--LSTLADVKEKIKL 66
E Q+ K+ E + L++ E E +E + S + A+A+ T +S L VKE+++
Sbjct: 242 ETEEQQAKQDSELAKLRVQEMEQGIADEASVASKAQLEVAQARHTSAISELESVKEELQT 301
Query: 67 L-NDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKEL 105
L N+Y V++ + V EE K + + L EL
Sbjct: 302 LQNEYDALVKEKDLAVKEAEEAVIASKEVERKVEELTIEL 341
>At1g77580 unknown protein
Length = 779
Score = 33.9 bits (76), Expect = 0.15
Identities = 30/121 (24%), Positives = 59/121 (47%), Gaps = 18/121 (14%)
Query: 3 SLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRA-----KAQQTLSTL 57
SL ++E+ + R+KE+ E L+ LE E +LE E C+ + A ++ S
Sbjct: 340 SLASEIEVLTSRIKELEEK--LEKLEAEKHELENEVKCNR---EEAVVHIENSEVLTSRT 394
Query: 58 ADVKEKIKLLNDYKTQVE-----DAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRF 112
+++EK++ L K +++ + E V+ E + + E +S KEL + +++
Sbjct: 395 KELEEKLEKLEAEKEELKSEVKCNREKAVVHVENSLAAE---IEVLTSRTKELEEQLEKL 451
Query: 113 E 113
E
Sbjct: 452 E 452
Score = 28.5 bits (62), Expect = 6.3
Identities = 27/119 (22%), Positives = 61/119 (50%), Gaps = 12/119 (10%)
Query: 3 SLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASC--SSFWDDRAKAQQTLSTLADV 60
+LR ++E + E+ L+ LE E A+L+ + + + Q+ + L ++
Sbjct: 518 TLRFELEAIACEKMELENK--LEKLEVEKAELQISFDIIKDKYEESQVCLQEIETKLGEI 575
Query: 61 KEKIKLLNDYKTQVEDAETIVMLTE------EMESVDKGLYEEASSLIKELNKSIDRFE 113
+ ++KL+N+ K +VE ++TI M + ++ES+++ + +E + EL + + E
Sbjct: 576 QTEMKLVNELKAEVE-SQTIAMEADAKTKSAKIESLEEDMRKERFA-FDELRRKCEALE 632
>At1g21810 myosin-like protein
Length = 628
Score = 33.5 bits (75), Expect = 0.19
Identities = 26/94 (27%), Positives = 42/94 (44%), Gaps = 10/94 (10%)
Query: 27 LEQELAKLEEEAS---------CSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDA 77
+E EL K+E E + + + R Q+ L +K ++KL N+ KTQ E
Sbjct: 391 MEDELEKMEAEKAELKISFDVIKDQYQESRVCFQEVEMKLEAMKRELKLANESKTQAESR 450
Query: 78 ETIVMLTEEMES-VDKGLYEEASSLIKELNKSID 110
T + E V GL E+ + +EL + I+
Sbjct: 451 VTRMEAEVRKERIVSDGLKEKCETFEEELRREIE 484
>At5g11390 unknown protein
Length = 703
Score = 32.3 bits (72), Expect = 0.43
Identities = 35/165 (21%), Positives = 71/165 (42%), Gaps = 23/165 (13%)
Query: 35 EEEASCSSFWDDRAKAQQTLS-------TLADVKEKIKLLNDYKTQVE-----------D 76
+EE S+ DD A++ L ++VKE LL + +++ D
Sbjct: 93 DEEEPSSNVDDDDDSAEKALEFDLLSSILNSEVKELESLLGFLQNEIQSARVMISPFQHD 152
Query: 77 AETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAG 136
E + L ++ ++ L + ++ E+ K F Q LS D++G+
Sbjct: 153 GEAFLDLEGKLNDTEQSLGQLMEQVV-EMKKQSSNF---QRLSSGLDEQGSWSGGQTSVS 208
Query: 137 GTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIE 181
D + + D+ ++ ++ ++QR R++EKSL +E ++ E
Sbjct: 209 QNDGE-FGDLSAKINMQTADQQRNVLRMLEKSLAKEMELEKKLSE 252
>At1g78940 unknown protein
Length = 740
Score = 32.0 bits (71), Expect = 0.57
Identities = 29/117 (24%), Positives = 50/117 (41%), Gaps = 17/117 (14%)
Query: 6 KDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLST------LAD 59
K+ A Q+ E+++ + E AK EEA+ S +RAKA+ L LA+
Sbjct: 336 KEALSARQQATELQKLRTEEERRLEEAKSSEEAAMSIVEKERAKAKAALEAAEAAKRLAE 395
Query: 60 VKEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQ 116
V+ K +L + KT +E +S +G + E+ ++ F +Q
Sbjct: 396 VESKRRLTAEMKTM-----------KESDSFSRGFVRYRKYTVDEIEEATSNFAESQ 441
>At1g54560
Length = 1529
Score = 31.6 bits (70), Expect = 0.74
Identities = 31/105 (29%), Positives = 49/105 (46%), Gaps = 17/105 (16%)
Query: 4 LRKDVEIASQRVKEIRESSGLQLLE---QELAKLEEEASCSSFWDDRAKAQQTLSTLADV 60
L K VE + R ++ + S + L E QE+ KL+ SS + R K +T L
Sbjct: 897 LEKKVEELTYRA-QLEKRSRVDLEEEKNQEIKKLQ-----SSLEEMRKKVDETNGLLVKE 950
Query: 61 KEKIK--------LLNDYKTQVEDAETIVMLTEEMESVDKGLYEE 97
+E K ++ + + VED + I LTEE+E + L +E
Sbjct: 951 REAAKKAIEEAPPVVTETQVLVEDTQKIEALTEEVEGLKANLEQE 995
>At5g44690 putative protein
Length = 684
Score = 31.2 bits (69), Expect = 0.97
Identities = 20/94 (21%), Positives = 45/94 (47%), Gaps = 4/94 (4%)
Query: 19 RESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAE 78
R G L+E+E E CS + + +T + + + + +++ D K Q++ +
Sbjct: 540 RSEIGSNLVEEEEGNEETLVECSERYQGNVEENETKKLILEPETETQMICDLKQQIKQLQ 599
Query: 79 TIVMLTEEMESVDKGLYEEASSL-IKELNKSIDR 111
++ E++S+ K + SL + L+ S++R
Sbjct: 600 RDIL---ELQSLVKSCVDFQKSLEFESLSDSLER 630
>At5g33300 putative protein
Length = 439
Score = 30.8 bits (68), Expect = 1.3
Identities = 22/80 (27%), Positives = 43/80 (53%), Gaps = 3/80 (3%)
Query: 13 QRVKEIRESSGLQLLEQELAKLE---EEASCSSFWDDRAKAQQTLSTLADVKEKIKLLND 69
+++KE+ E LE L K E E++SCS+ D++ S L ++KE+ L+
Sbjct: 224 RQMKEMAEEIKRFSLEAGLLKAEFEGEQSSCSASCDNQINHTPMDSELKELKEEFNKLST 283
Query: 70 YKTQVEDAETIVMLTEEMES 89
+Q+E ++ T++++S
Sbjct: 284 LVSQMEMTKSQFSETDKVQS 303
>At5g20360 tetratricopeptide repeat protein
Length = 809
Score = 30.0 bits (66), Expect = 2.2
Identities = 18/67 (26%), Positives = 34/67 (49%)
Query: 22 SGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAETIV 81
S ++ L+ ++K+E C S + A + TL D K+K L+D ++VE+ + V
Sbjct: 34 SKVETLDDCVSKVETLDDCVSKAETLADCVSKVETLDDCVSKVKTLDDCVSKVENLDDCV 93
Query: 82 MLTEEME 88
E ++
Sbjct: 94 PKVETLD 100
Score = 28.5 bits (62), Expect = 6.3
Identities = 25/96 (26%), Positives = 44/96 (45%), Gaps = 13/96 (13%)
Query: 22 SGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVE------ 75
S ++ L+ ++K E A C S + + TL D K++ L+D +VE
Sbjct: 44 SKVETLDDCVSKAETLADCVSKVETLDDCVSKVKTLDDCVSKVENLDDCVPKVETLDDCV 103
Query: 76 -DAETIVMLTEEMESVD------KGLYEEASSLIKE 104
ET+ E+E++D +GL EE + L ++
Sbjct: 104 PKVETLDDCVSEVETLDDCVSKAQGLKEEGNKLFQK 139
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.131 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,703,764
Number of Sequences: 26719
Number of extensions: 320363
Number of successful extensions: 986
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 32
Number of HSP's that attempted gapping in prelim test: 956
Number of HSP's gapped (non-prelim): 53
length of query: 375
length of database: 11,318,596
effective HSP length: 101
effective length of query: 274
effective length of database: 8,619,977
effective search space: 2361873698
effective search space used: 2361873698
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147179.9


