

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.5 - phase: 0
(706 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
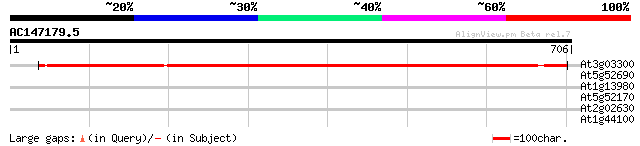
Score E
Sequences producing significant alignments: (bits) Value
At3g03300 unknown protein 835 0.0
At5g52690 unknown protein 32 1.2
At1g13980 putative pattern formation protein EMB30 31 2.7
At5g52170 homeodomain transcription factor-like 30 6.1
At2g02630 hypothetical protein 29 7.9
At1g44100 amino acid permease AAP5 29 7.9
>At3g03300 unknown protein
Length = 2042
Score = 835 bits (2157), Expect = 0.0
Identities = 409/668 (61%), Positives = 496/668 (74%), Gaps = 12/668 (1%)
Query: 37 NGETVIEPKGTPDSAVWVVQLSDLHLSVHHPNRALDFTNLVGHALSFINPSLVLITGDLT 96
+G VIE + T +WVVQLSDLH SVHHP RA+DF N+VG AL+ INPSLVLITGDLT
Sbjct: 45 SGRRVIEAR-TGQDLIWVVQLSDLHFSVHHPERAIDFKNIVGPALALINPSLVLITGDLT 103
Query: 97 DGKSKDLLTMKQNEDEWVEYRNVMDGVIERSGLHKSLFFDLRGNHDSFGVPVVGGSFDFF 156
DGKSKD+LTMK NEDEW+EY +VM V++RSGL+KS+F+DLRGNHD+FGVP VG S DFF
Sbjct: 104 DGKSKDMLTMKLNEDEWLEYESVMQDVVKRSGLNKSIFYDLRGNHDNFGVPSVGSSVDFF 163
Query: 157 SKYSVNGQLGRNRSVNSVTLETKERKHLFVGIDTTMSAGAGLRGPTNVFGHPTDQLLKDL 216
SKYS+NGQ+GR +VN++T+ET ERKHLFVGIDTTM G LRGPTN+FGHPTD+LL L
Sbjct: 164 SKYSINGQMGRKGNVNTITVETSERKHLFVGIDTTMHIG--LRGPTNLFGHPTDELLSSL 221
Query: 217 DLELSHWDSQSEKPVTKISFGHFPLSFSAPSSSGRTLKEVFLKHSISAYLCGHLHSRFGK 276
D LS WD+QS KPV KISFGHFPLSF+A S S ++LK+VFLKHSISAYLCGHLHSRFGK
Sbjct: 222 DSHLSQWDNQSAKPVAKISFGHFPLSFTALSHSQKSLKDVFLKHSISAYLCGHLHSRFGK 281
Query: 277 NLKRHHQLSNRFLSLQNFFQFNVHQNSFESTVNCSIGA-PPQEFWEWEIGDWRKSRAIRI 335
NLKRHH LS + FQ N+ Q+ EST NCS GA P EFWEWE+GDWRK+RA+RI
Sbjct: 282 NLKRHHHSGGISLSDNDLFQLNMRQSGAESTSNCSFGALPAAEFWEWEMGDWRKNRAMRI 341
Query: 336 LAIDRGHVSYVDLDFKSGAKHAIILPTFPLDSRFMQTSSWHHNYECQSVASSSYETIRAL 395
+AIDRGHVSYVDLDFKS + IILPTFPLDSRFM TS H YECQ + SSSY+ IRA+
Sbjct: 342 MAIDRGHVSYVDLDFKSKPQKTIILPTFPLDSRFMSTSFARHKYECQHMISSSYDAIRAI 401
Query: 396 VFSASPVESVVARVYDSRYGDLVLVVEAHMTKRADENSRGN--LYVAPWNYKAFEDTSPD 453
VFS S V VVARVYDS G LV+EA M K ++S + PWNY+AFED PD
Sbjct: 402 VFSHSLVVEVVARVYDSSPGFDNLVMEAPMRKHGGDSSSSGATFFSLPWNYRAFEDPLPD 461
Query: 454 RFWFQIESNDIMGRSTLTELRPFSINGHSFRLSWSWKEFFVMGCQWASLYYPLLWSALGF 513
RFW QIE DI GR TL+E+RPFSING S ++SW+W EF VMGCQWA+LYYP+LW AL
Sbjct: 462 RFWLQIEVTDIKGRLTLSEMRPFSINGLSSKVSWTWNEFRVMGCQWAALYYPILWPALYS 521
Query: 514 MFSFLLVPKALLFFQMNLYTYRNFIANKGVVNGALWILQELCRVPTLWFGWIGYLFYLIL 573
+F L+PK ++ YT + FIA KG + LWILQ+LCR+P +WFG++ YLFYLI
Sbjct: 522 LFLVFLIPKCIIIVFKKQYTLKKFIAKKGPITLVLWILQDLCRMPVVWFGYMAYLFYLIF 581
Query: 574 FPWFMGQVFTEGKSTVYMTYMGWAVETSNGKGKFEWVGFPDIMVLVLPHILFVVLPAILV 633
FPWF G+VF + YMT MGW V +S K E++G PD+MV+V+PH++FVV+P++LV
Sbjct: 582 FPWFSGEVFADSGDRAYMTIMGWVVTSSGADRKHEYIGQPDVMVVVIPHVVFVVIPSVLV 641
Query: 634 TGALTAERAIYRERVLALSGKKKDDIDLNSRRPLLNGNHNSTISTLHLGKRWIRKLLCVV 693
L AER IY++ + +SGKK+DD D ++ + S L +R RK + +
Sbjct: 642 VCCLVAEREIYKDHIRTVSGKKEDDHDRGRKK------RSQRRSLLFSNRRLFRKSVLLA 695
Query: 694 CLAICWKH 701
LA+ WKH
Sbjct: 696 SLALYWKH 703
>At5g52690 unknown protein
Length = 177
Score = 32.0 bits (71), Expect = 1.2
Identities = 30/125 (24%), Positives = 62/125 (49%), Gaps = 10/125 (8%)
Query: 59 DLHLSVHHPNRALDFTNLVGHALSFINPSLVLITGDLTDGKSKDLLTMKQNEDEWVEYRN 118
D+ +H ++++D N G+ + P + LT K ++++ K N + R+
Sbjct: 52 DMGSKLHEKDKSVDIINPNGNRGQYRIPGYTTVYNYLTTPKIQEIVVFKFNVLDEKIKRD 111
Query: 119 VMDGVIERSGLHKSLFFDLRGNHDSFGVPVVGGSFDFFSKYSVNGQLGR-NRSVNSVTLE 177
VM+ + E SG+ +L+ +H + V GG F+K +++ +L R + SV+ +
Sbjct: 112 VMEVIWEFSGITS---VELKRDHQ---LEVKGGE---FNKVAMSTKLKRIDESVSLIINY 162
Query: 178 TKERK 182
T+ +K
Sbjct: 163 TRTKK 167
>At1g13980 putative pattern formation protein EMB30
Length = 1451
Score = 30.8 bits (68), Expect = 2.7
Identities = 16/32 (50%), Positives = 20/32 (62%), Gaps = 3/32 (9%)
Query: 127 SGLHKSLFFDLRGNHDSFGVPVV---GGSFDF 155
+GL K+L D GNHD F V V+ G+FDF
Sbjct: 606 AGLDKNLVGDFLGNHDEFCVQVLNEFAGTFDF 637
>At5g52170 homeodomain transcription factor-like
Length = 682
Score = 29.6 bits (65), Expect = 6.1
Identities = 16/31 (51%), Positives = 19/31 (60%), Gaps = 4/31 (12%)
Query: 210 DQLLKDLDLELSHWDSQSEKPVTKISFGHFP 240
D+LLK +LE S W S+SEK S HFP
Sbjct: 230 DELLKLAELETSLWSSKSEKG----SMNHFP 256
>At2g02630 hypothetical protein
Length = 617
Score = 29.3 bits (64), Expect = 7.9
Identities = 13/49 (26%), Positives = 25/49 (50%)
Query: 91 ITGDLTDGKSKDLLTMKQNEDEWVEYRNVMDGVIERSGLHKSLFFDLRG 139
+ D+ DGK + + + ED+ ++ + DGVI L+F++ G
Sbjct: 297 LRSDIWDGKELEGVPEEVEEDDVESFKRIADGVILHFSHRHHLYFEISG 345
>At1g44100 amino acid permease AAP5
Length = 480
Score = 29.3 bits (64), Expect = 7.9
Identities = 25/89 (28%), Positives = 41/89 (45%), Gaps = 11/89 (12%)
Query: 474 RPFSINGHSFRLSWSWKEFFVMGCQWASLYYPLLWSALGFMFSFLLVPKALLFFQMNLYT 533
+PF++N FRL W + FFVM S+ P +G + + P ++F + +Y
Sbjct: 375 KPFNLN--LFRLVW--RTFFVMTTTLISMLMPFFNDVVGLLGAIGFWP-LTVYFPVEMY- 428
Query: 534 YRNFIANKGVVN-GALWILQELCRVPTLW 561
IA K V G W+ ++ V L+
Sbjct: 429 ----IAQKNVPRWGTKWVCLQVLSVTCLF 453
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.138 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,660,923
Number of Sequences: 26719
Number of extensions: 737822
Number of successful extensions: 2138
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 2129
Number of HSP's gapped (non-prelim): 9
length of query: 706
length of database: 11,318,596
effective HSP length: 106
effective length of query: 600
effective length of database: 8,486,382
effective search space: 5091829200
effective search space used: 5091829200
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 64 (29.3 bits)
Medicago: description of AC147179.5


