

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147178.10 + phase: 0
(161 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
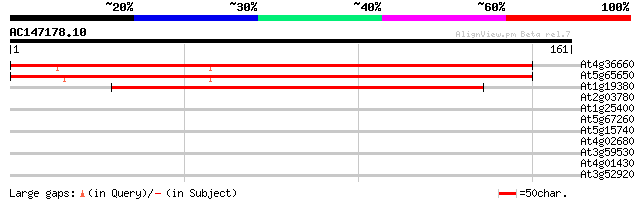
Score E
Sequences producing significant alignments: (bits) Value
At4g36660 hypothetical protein 227 2e-60
At5g65650 putative protein 223 5e-59
At1g19380 unknown protein 120 3e-28
At2g03780 unknown protein 28 1.6
At1g25400 unknown protein 28 2.2
At5g67260 cyclin D3-like protein 27 3.7
At5g15740 unknown protein 27 4.8
At4g02680 unknown protein 27 6.3
At3g59530 unknown protein 27 6.3
At4g01430 unknown protein 26 8.2
At3g52920 unknown protein 26 8.2
>At4g36660 hypothetical protein
Length = 167
Score = 227 bits (579), Expect = 2e-60
Identities = 118/160 (73%), Positives = 130/160 (80%), Gaps = 10/160 (6%)
Query: 1 MKDDDGLPTTTA---------AINKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTV 51
MK+DD LPTTT A +KKE+ DS LFG+GRYKFWA AAILLLAFWSMFTGTV
Sbjct: 1 MKEDDALPTTTTTGTATGTAMANSKKESSDSVLFGRGRYKFWAFAAILLLAFWSMFTGTV 60
Query: 52 SLRWS-GNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFE 110
+LR S GNLN LS DL P +D+LDVLEMEEREKVV+HMWDVYTNSRRI+LPRFWQEAF
Sbjct: 61 TLRLSTGNLNRLSEDLGIPNYDNLDVLEMEEREKVVKHMWDVYTNSRRIKLPRFWQEAFV 120
Query: 111 AAYEELTSDVPGVRDDAITEIAKMSVRSVNYDPPPIQSTV 150
AAYEELTSDVPGVR+ AI EIAKMS RS+ DPPP +S V
Sbjct: 121 AAYEELTSDVPGVREAAIGEIAKMSARSITLDPPPSRSRV 160
>At5g65650 putative protein
Length = 159
Score = 223 bits (567), Expect = 5e-59
Identities = 112/157 (71%), Positives = 127/157 (80%), Gaps = 7/157 (4%)
Query: 1 MKDDDGLPTTTAAI------NKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLR 54
MKD D LP +T+++ KKE S LF KGRYKFWALAAILLLAFWSM TGTV+LR
Sbjct: 1 MKDGDSLPISTSSVAATTVTGKKETGYSALFSKGRYKFWALAAILLLAFWSMLTGTVNLR 60
Query: 55 WS-GNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAY 113
WS GN+N + DL PIH+DLDVLEMEEREKVV+HMWDVY N RRIRLPRFWQEAFEAAY
Sbjct: 61 WSAGNINHFTDDLVFPIHEDLDVLEMEEREKVVKHMWDVYNNGRRIRLPRFWQEAFEAAY 120
Query: 114 EELTSDVPGVRDDAITEIAKMSVRSVNYDPPPIQSTV 150
EELTSDVP V + AI+EIA+MS+RS+ DPPP+ STV
Sbjct: 121 EELTSDVPDVVEAAISEIARMSIRSIVIDPPPLHSTV 157
>At1g19380 unknown protein
Length = 147
Score = 120 bits (301), Expect = 3e-28
Identities = 61/107 (57%), Positives = 77/107 (71%)
Query: 30 YKFWALAAILLLAFWSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHM 89
YK W L A+LLLAF SM TG+VSL+ G ++ DDLDVLE+EEREKVVR M
Sbjct: 25 YKLWVLIAVLLLAFGSMLTGSVSLKGIGLFHSADGVNAFSFGDDLDVLEIEEREKVVRQM 84
Query: 90 WDVYTNSRRIRLPRFWQEAFEAAYEELTSDVPGVRDDAITEIAKMSV 136
WDVY S +++PRFW+EAFEAAYE L SD VR+ A+++IAK+S+
Sbjct: 85 WDVYGRSGGVKVPRFWREAFEAAYEFLISDSAAVRNAAVSDIAKLSL 131
>At2g03780 unknown protein
Length = 287
Score = 28.5 bits (62), Expect = 1.6
Identities = 16/56 (28%), Positives = 30/56 (53%), Gaps = 2/56 (3%)
Query: 80 EEREKVVRHMWDVYTNSRRI--RLPRFWQEAFEAAYEELTSDVPGVRDDAITEIAK 133
E+RE+VV+ D+ NS+++ ++ R ++ E E+ D+ VRD + K
Sbjct: 66 EKRERVVKVSRDITMNSKKVIFQVHRLSKDNKEEVLEKAGKDLEAVRDQHFARLMK 121
>At1g25400 unknown protein
Length = 288
Score = 28.1 bits (61), Expect = 2.2
Identities = 11/19 (57%), Positives = 14/19 (72%)
Query: 30 YKFWALAAILLLAFWSMFT 48
Y FW A+LLLAF++ FT
Sbjct: 41 YSFWKWGALLLLAFFASFT 59
>At5g67260 cyclin D3-like protein
Length = 367
Score = 27.3 bits (59), Expect = 3.7
Identities = 21/75 (28%), Positives = 35/75 (46%), Gaps = 12/75 (16%)
Query: 16 KKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLR----WSGNLNTLSS------- 64
+KE +D L K Y F +L AIL + ++ F ++ L+ W L ++S
Sbjct: 96 RKEALDWVLRVKSHYGFTSLTAILAVNYFDRFMTSIKLQTDKPWMSQLVAVASLSLAAKV 155
Query: 65 -DLDTPIHDDLDVLE 78
++ P+ DL V E
Sbjct: 156 EEIQVPLLLDLQVEE 170
>At5g15740 unknown protein
Length = 508
Score = 26.9 bits (58), Expect = 4.8
Identities = 10/39 (25%), Positives = 19/39 (48%)
Query: 15 NKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSL 53
+K E + + + R W + A+ +L WS F ++L
Sbjct: 24 SKVEKLKNSFVSRPRMSLWMIRAVTVLLLWSCFVHLMAL 62
>At4g02680 unknown protein
Length = 888
Score = 26.6 bits (57), Expect = 6.3
Identities = 15/44 (34%), Positives = 22/44 (49%), Gaps = 5/44 (11%)
Query: 70 IHDDLDVLEMEEREKVVRHMWDVY-----TNSRRIRLPRFWQEA 108
IH++LD ++ER + + V+ T RR L WQEA
Sbjct: 70 IHEELDTCPLQERSILYLLQYQVFRGLGETKLRRRSLQSAWQEA 113
>At3g59530 unknown protein
Length = 403
Score = 26.6 bits (57), Expect = 6.3
Identities = 10/35 (28%), Positives = 20/35 (56%)
Query: 92 VYTNSRRIRLPRFWQEAFEAAYEELTSDVPGVRDD 126
++T + RL ++W E + E+ +D+PG D+
Sbjct: 269 LFTETTNCRLVKYWLEGPKMGEVEVVADLPGFPDN 303
>At4g01430 unknown protein
Length = 365
Score = 26.2 bits (56), Expect = 8.2
Identities = 12/39 (30%), Positives = 20/39 (50%), Gaps = 6/39 (15%)
Query: 26 GKGRYKFWALAAI------LLLAFWSMFTGTVSLRWSGN 58
G + K W L + +LL+ W +F G +S ++ GN
Sbjct: 176 GHDQTKKWLLGCLYLVIGTVLLSLWMLFQGKLSFKYPGN 214
>At3g52920 unknown protein
Length = 180
Score = 26.2 bits (56), Expect = 8.2
Identities = 22/79 (27%), Positives = 36/79 (44%), Gaps = 12/79 (15%)
Query: 72 DDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAYEELTSDVPGVRDDAITEI 131
D+++ +ME RE+V + V +RR+ R E E + + +V VR
Sbjct: 44 DEIEKRKMEVRERVKAQLGRVEEETRRLASIR---EELETMADPMRKEVNWVR------- 93
Query: 132 AKMSVRSVNYDPPPIQSTV 150
+ SVN + P+ STV
Sbjct: 94 --KKIDSVNKELKPLGSTV 110
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.133 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,774,769
Number of Sequences: 26719
Number of extensions: 144360
Number of successful extensions: 408
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 398
Number of HSP's gapped (non-prelim): 11
length of query: 161
length of database: 11,318,596
effective HSP length: 91
effective length of query: 70
effective length of database: 8,887,167
effective search space: 622101690
effective search space used: 622101690
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC147178.10


