

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147014.15 - phase: 0
(138 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
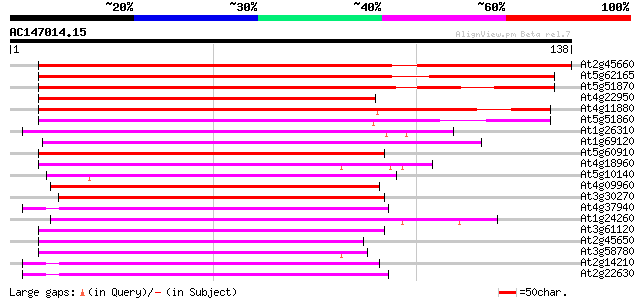
At2g45660 MADS-box protein (AGL20) 134 2e-32
At5g62165 unknown protein 98 1e-21
At5g51870 MADS box transcription factor-like 94 3e-20
At4g22950 putative MADS Box / AGL protein 91 2e-19
At4g11880 MADS-box protein AGL14 87 2e-18
At5g51860 MADS box transcription factor-like protein 85 1e-17
At1g26310 cauliflower (CAL) 74 3e-14
At1g69120 unknown protein 70 3e-13
At5g60910 floral homeotic protein AGL8 70 5e-13
At4g18960 floral homeotic protein agamous 61 2e-10
At5g10140 MADS box protein FLOWERING LOCUS F (FLF) 60 3e-10
At4g09960 MADS-box protein AGL11 59 6e-10
At3g30270 MADS-box transcription factor 59 8e-10
At4g37940 MADS-box protein AGL17 -like protein 59 1e-09
At1g24260 floral homeotic protein, AGL9 57 2e-09
At3g61120 MADS-box protein AGL13 57 3e-09
At2g45650 MADS-box protein (AGL6) 55 9e-09
At3g58780 shatterproof 1 (SHP1)/ agamous -like 1 (AGL1) 55 1e-08
At2g14210 pseudogene 53 4e-08
At2g22630 MADS-box protein AGL17 (AGL17) 53 6e-08
>At2g45660 MADS-box protein (AGL20)
Length = 214
Score = 134 bits (337), Expect = 2e-32
Identities = 73/131 (55%), Positives = 97/131 (73%), Gaps = 6/131 (4%)
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
Q+LK+E A++MKKIE LEASKRKL+GEG+G+CS++ELQQIEQQLEKSV +RARK Q +K
Sbjct: 88 QHLKYEAANMMKKIEQLEASKRKLLGEGIGTCSIEELQQIEQQLEKSVKCIRARKTQVFK 147
Query: 68 HQIDQLKEKEKNLVAENARLSKQVKRNYCPPQPQPQPTTKDHQREDQQPYAESSPSSDVV 127
QI+QLK+KEK L AEN +LS++ + + + + +Q + ESSPSS+V
Sbjct: 148 EQIEQLKQKEKALAAENEKLSEKWGSH------ESEVWSNKNQESTGRGDEESSPSSEVE 201
Query: 128 TELFIGLHRSS 138
T+LFIGL SS
Sbjct: 202 TQLFIGLPCSS 212
>At5g62165 unknown protein
Length = 210
Score = 98.2 bits (243), Expect = 1e-21
Identities = 54/127 (42%), Positives = 79/127 (61%), Gaps = 9/127 (7%)
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
Q LK E + ++ KIELLE KRKL+G+G+ SCSL+ELQ+I+ QL++S+ VR RK Q +K
Sbjct: 88 QQLKQEASHMITKIELLEFHKRKLLGQGIASCSLEELQEIDSQLQRSLGKVRERKAQLFK 147
Query: 68 HQIDQLKEKEKNLVAENARLSKQVKRNYCPPQPQPQPTTKDHQREDQQPYAESSPSSDVV 127
Q+++LK KEK L+ EN +L ++ N P + Q+ Y + +V
Sbjct: 148 EQLEKLKAKEKQLLEENVKLHQKNVIN---------PWRGSSTDQQQEKYKVIDLNLEVE 198
Query: 128 TELFIGL 134
T+LFIGL
Sbjct: 199 TDLFIGL 205
>At5g51870 MADS box transcription factor-like
Length = 207
Score = 93.6 bits (231), Expect = 3e-20
Identities = 54/127 (42%), Positives = 80/127 (62%), Gaps = 13/127 (10%)
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
Q LK E ++KKI+LLE RKL+G+GL SCS+ ELQ+I+ Q+EKS+ +VR+RK + Y
Sbjct: 89 QELKMEIDRMVKKIDLLEVHHRKLLGQGLDSCSVTELQEIDTQIEKSLRIVRSRKAELYA 148
Query: 68 HQIDQLKEKEKNLVAENARLSKQVKRNYCPPQPQPQPTTKDHQREDQQPYAESSPSSDVV 127
Q+ +LKEKE+ L+ E RL ++V ++ + T+ R + SS+V
Sbjct: 149 DQLKKLKEKERELLNERKRLLEEVNMHH-----SSKGNTEGGHR--------TKHSSEVE 195
Query: 128 TELFIGL 134
T+LFIGL
Sbjct: 196 TDLFIGL 202
>At4g22950 putative MADS Box / AGL protein
Length = 219
Score = 90.9 bits (224), Expect = 2e-19
Identities = 43/83 (51%), Positives = 64/83 (76%)
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
Q + ET+ L KKIE LE SKRKL+GEG+ +CS++ELQQ+E QL++S+S +RA+K Q +
Sbjct: 87 QQARDETSGLTKKIEQLEISKRKLLGEGIDACSIEELQQLENQLDRSLSRIRAKKYQLLR 146
Query: 68 HQIDQLKEKEKNLVAENARLSKQ 90
+I++LK +E+NLV EN L ++
Sbjct: 147 EEIEKLKAEERNLVKENKDLKEK 169
>At4g11880 MADS-box protein AGL14
Length = 221
Score = 87.4 bits (215), Expect = 2e-18
Identities = 51/128 (39%), Positives = 79/128 (60%), Gaps = 10/128 (7%)
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
Q K ET L +KIE LE S RK+MGEGL + S++ELQQ+E QL++S+ +RA+K Q +
Sbjct: 88 QQSKDETYGLARKIEHLEISTRKMMGEGLDASSIEELQQLENQLDRSLMKIRAKKYQLLR 147
Query: 68 HQIDQLKEKEKNLVAENARLSK--QVKRNYCPPQPQPQPTTKDHQREDQQPYAESSPSSD 125
+ ++LKEKE+NL+AEN L + +++ + +T + +D + +
Sbjct: 148 EETEKLKEKERNLIAENKMLMEKCEMQGRGIIGRISSSSSTSELDIDDNE--------ME 199
Query: 126 VVTELFIG 133
VVT+LFIG
Sbjct: 200 VVTDLFIG 207
>At5g51860 MADS box transcription factor-like protein
Length = 211
Score = 84.7 bits (208), Expect = 1e-17
Identities = 50/128 (39%), Positives = 72/128 (56%), Gaps = 13/128 (10%)
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
Q LK E +++KKIE+LE RK+MG+ L SCS+ EL +I Q+EKS+ +VR RK + Y+
Sbjct: 89 QGLKKEMVTMVKKIEVLEVHNRKMMGQSLDSCSVKELSEIATQIEKSLHMVRLRKAKLYE 148
Query: 68 HQIDQLKEKEKNLVAENARLS--KQVKRNYCPPQPQPQPTTKDHQREDQQPYAESSPSSD 125
++ +LK KE+ L E RLS K + + C +P + S D
Sbjct: 149 DELQKLKAKERELKDERVRLSLKKTIYTHLCQVGERPMGMP-----------SGSKEKED 197
Query: 126 VVTELFIG 133
V T+LFIG
Sbjct: 198 VETDLFIG 205
>At1g26310 cauliflower (CAL)
Length = 255
Score = 73.6 bits (179), Expect = 3e-14
Identities = 45/122 (36%), Positives = 64/122 (51%), Gaps = 16/122 (13%)
Query: 4 LNIWQNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKN 63
+N N E + L KIELLE ++R +GE L SL +LQ +EQQLE ++ +R+RKN
Sbjct: 87 VNAQTNWSMEYSRLKAKIELLERNQRHYLGEELEPMSLKDLQNLEQQLETALKHIRSRKN 146
Query: 64 QAYKHQIDQLKEKEKNLVAENARLSKQV---------KRNYC-------PPQPQPQPTTK 107
Q ++ L+ KEK + EN+ L+KQ+ K+ C PQPQP
Sbjct: 147 QLMNESLNHLQRKEKEIQEENSMLTKQIKERENILRTKQTQCEQLNRSVDDVPQPQPFQH 206
Query: 108 DH 109
H
Sbjct: 207 PH 208
>At1g69120 unknown protein
Length = 256
Score = 70.5 bits (171), Expect = 3e-13
Identities = 40/108 (37%), Positives = 62/108 (57%)
Query: 9 NLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYKH 68
N E L KIELLE ++R +GE L + S ELQ +EQQL+ ++ +R RKNQ
Sbjct: 90 NWSMEYNRLKAKIELLERNQRHYLGEDLQAMSPKELQNLEQQLDTALKHIRTRKNQLMYE 149
Query: 69 QIDQLKEKEKNLVAENARLSKQVKRNYCPPQPQPQPTTKDHQREDQQP 116
I++L++KEK + +N+ LSKQ+K + Q + + +Q + P
Sbjct: 150 SINELQKKEKAIQEQNSMLSKQIKEREKILRAQQEQWDQQNQGHNMPP 197
>At5g60910 floral homeotic protein AGL8
Length = 242
Score = 69.7 bits (169), Expect = 5e-13
Identities = 37/85 (43%), Positives = 53/85 (61%)
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
+N E A L ++E+LE +KR MGE L S SL ELQ +E QL+ ++ +R+RKNQA
Sbjct: 89 ENWVLEHAKLKARVEVLEKNKRNFMGEDLDSLSLKELQSLEHQLDAAIKSIRSRKNQAMF 148
Query: 68 HQIDQLKEKEKNLVAENARLSKQVK 92
I L++K+K L N L K++K
Sbjct: 149 ESISALQKKDKALQDHNNSLLKKIK 173
>At4g18960 floral homeotic protein agamous
Length = 284
Score = 60.8 bits (146), Expect = 2e-10
Identities = 39/116 (33%), Positives = 66/116 (56%), Gaps = 19/116 (16%)
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
Q + E+A L ++I ++ S R+LMGE +GS S EL+ +E +LE+S++ +R++KN+
Sbjct: 136 QYYQQESAKLRQQIISIQNSNRQLMGETIGSMSPKELRNLEGRLERSITRIRSKKNELLF 195
Query: 68 HQIDQLKEKEKNL----------VAENARLSKQVK-----RNY----CPPQPQPQP 104
+ID ++++E +L +AEN R + + NY PPQ Q QP
Sbjct: 196 SEIDYMQKREVDLHNDNQILRAKIAENERNNPSISLMPGGSNYEQLMPPPQTQSQP 251
>At5g10140 MADS box protein FLOWERING LOCUS F (FLF)
Length = 196
Score = 60.5 bits (145), Expect = 3e-10
Identities = 34/88 (38%), Positives = 53/88 (59%), Gaps = 2/88 (2%)
Query: 10 LKHETASLM--KKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
L H++ +L ELLE KL+G + + S+D L Q+E+ LE ++SV RA+K +
Sbjct: 81 LDHQSKALNYGSHYELLELVDSKLVGSNVKNVSIDALVQLEEHLETALSVTRAKKTELML 140
Query: 68 HQIDQLKEKEKNLVAENARLSKQVKRNY 95
++ LKEKEK L EN L+ Q++ N+
Sbjct: 141 KLVENLKEKEKMLKEENQVLASQMENNH 168
>At4g09960 MADS-box protein AGL11
Length = 230
Score = 59.3 bits (142), Expect = 6e-10
Identities = 29/81 (35%), Positives = 54/81 (65%)
Query: 11 KHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYKHQI 70
+ E+A L ++I+ ++ S R LMG+ L S S+ EL+Q+E +LEK++S +R++K++ +I
Sbjct: 91 QQESAKLRQQIQTIQNSNRNLMGDSLSSLSVKELKQVENRLEKAISRIRSKKHELLLVEI 150
Query: 71 DQLKEKEKNLVAENARLSKQV 91
+ +++E L EN L +V
Sbjct: 151 ENAQKREIELDNENIYLRTKV 171
>At3g30270 MADS-box transcription factor
Length = 125
Score = 58.9 bits (141), Expect = 8e-10
Identities = 31/80 (38%), Positives = 50/80 (61%)
Query: 13 ETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYKHQIDQ 72
E + L++ I++L+ S R L GE + S+ +LQ +E QL+ ++ R+RKNQ I Q
Sbjct: 32 ECSKLLRMIDVLQRSLRHLRGEEVDGLSIRDLQGVEMQLDTALKKTRSRKNQLMVESIAQ 91
Query: 73 LKEKEKNLVAENARLSKQVK 92
L++KEK L +L+K+VK
Sbjct: 92 LQKKEKELKELKKQLTKKVK 111
>At4g37940 MADS-box protein AGL17 -like protein
Length = 228
Score = 58.5 bits (140), Expect = 1e-09
Identities = 31/90 (34%), Positives = 52/90 (57%), Gaps = 3/90 (3%)
Query: 4 LNIWQNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKN 63
+ WQ E A L +++ L+ + R++MGE L S++EL +E Q+E S+ +R RK
Sbjct: 86 VKFWQR---EAAVLRQELHALQENHRQMMGEQLNGLSVNELNSLENQIEISLRGIRMRKE 142
Query: 64 QAYKHQIDQLKEKEKNLVAENARLSKQVKR 93
Q +I +L +K + EN LS++V+R
Sbjct: 143 QLLTQEIQELSQKRNLIHQENLDLSRKVQR 172
>At1g24260 floral homeotic protein, AGL9
Length = 251
Score = 57.4 bits (137), Expect = 2e-09
Identities = 34/114 (29%), Positives = 59/114 (50%), Gaps = 4/114 (3%)
Query: 11 KHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYKHQI 70
+ E L ++ + L+ ++R L+GE LG S EL+ +E+QL+ S+ +RA + Q Q+
Sbjct: 95 QQEYLKLKERYDALQRTQRNLLGEDLGPLSTKELESLERQLDSSLKQIRALRTQFMLDQL 154
Query: 71 DQLKEKEKNLVAENARLSKQVKRNY-CPPQPQPQPTTKDH---QREDQQPYAES 120
+ L+ KE+ L N L ++ Y P Q P DH QQ ++++
Sbjct: 155 NDLQSKERMLTETNKTLRLRLADGYQMPLQLNPNQEEVDHYGRHHHQQQQHSQA 208
>At3g61120 MADS-box protein AGL13
Length = 244
Score = 57.0 bits (136), Expect = 3e-09
Identities = 31/85 (36%), Positives = 48/85 (56%)
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
Q L+ E L K E L + R L+GE L S+ ELQ +E+QLE ++S R +K Q
Sbjct: 86 QGLRQEVTKLKCKYESLLRTHRNLVGEDLEGMSIKELQTLERQLEGALSATRKQKTQVMM 145
Query: 68 HQIDQLKEKEKNLVAENARLSKQVK 92
Q+++L+ KE+ L N +L + +
Sbjct: 146 EQMEELRRKERELGDINNKLKLETE 170
>At2g45650 MADS-box protein (AGL6)
Length = 252
Score = 55.5 bits (132), Expect = 9e-09
Identities = 29/80 (36%), Positives = 45/80 (56%)
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
Q+ E L K E L + R L+GE LG + ELQ +E+QLE +++ R RK Q
Sbjct: 87 QSWCQEVTKLKSKYESLVRTNRNLLGEDLGEMGVKELQALERQLEAALTATRQRKTQVMM 146
Query: 68 HQIDQLKEKEKNLVAENARL 87
+++ L++KE+ L N +L
Sbjct: 147 EEMEDLRKKERQLGDINKQL 166
>At3g58780 shatterproof 1 (SHP1)/ agamous -like 1 (AGL1)
Length = 248
Score = 55.1 bits (131), Expect = 1e-08
Identities = 30/91 (32%), Positives = 54/91 (58%), Gaps = 10/91 (10%)
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
Q + E + L ++I ++ S R ++GE LGS + EL+ +E +LEK +S VR++KN+
Sbjct: 103 QYYQQEASKLRRQIRDIQNSNRHIVGESLGSLNFKELKNLEGRLEKGISRVRSKKNELLV 162
Query: 68 HQIDQLKEKEKNL----------VAENARLS 88
+I+ ++++E L +AE ARL+
Sbjct: 163 AEIEYMQKREMELQHNNMYLRAKIAEGARLN 193
>At2g14210 pseudogene
Length = 234
Score = 53.1 bits (126), Expect = 4e-08
Identities = 30/88 (34%), Positives = 49/88 (55%), Gaps = 3/88 (3%)
Query: 4 LNIWQNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKN 63
+ WQ E ASL ++++ L+ RKL+GE L + ++LQ +E QL S+ VR +K+
Sbjct: 87 IKFWQR---EVASLQQQLQYLQECHRKLVGEELSGMNANDLQNLEDQLVTSLKGVRLKKD 143
Query: 64 QAYKHQIDQLKEKEKNLVAENARLSKQV 91
Q ++I +L K + + EN L V
Sbjct: 144 QLMTNEIRELNRKGQIIQKENHELQNIV 171
>At2g22630 MADS-box protein AGL17 (AGL17)
Length = 227
Score = 52.8 bits (125), Expect = 6e-08
Identities = 30/90 (33%), Positives = 50/90 (55%), Gaps = 3/90 (3%)
Query: 4 LNIWQNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKN 63
+ WQ E +L +++ L+ + R+L G L S+ ELQ IE QLE S+ +R ++
Sbjct: 86 VKFWQR---EAETLRQELHSLQENYRQLTGVELNGLSVKELQNIESQLEMSLRGIRMKRE 142
Query: 64 QAYKHQIDQLKEKEKNLVAENARLSKQVKR 93
Q ++I +L K + EN LS++V+R
Sbjct: 143 QILTNEIKELTRKRNLVHHENLELSRKVQR 172
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.310 0.126 0.343
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,047,589
Number of Sequences: 26719
Number of extensions: 122028
Number of successful extensions: 1099
Number of sequences better than 10.0: 167
Number of HSP's better than 10.0 without gapping: 80
Number of HSP's successfully gapped in prelim test: 87
Number of HSP's that attempted gapping in prelim test: 848
Number of HSP's gapped (non-prelim): 330
length of query: 138
length of database: 11,318,596
effective HSP length: 89
effective length of query: 49
effective length of database: 8,940,605
effective search space: 438089645
effective search space used: 438089645
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC147014.15


