

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147013.3 + phase: 0
(380 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
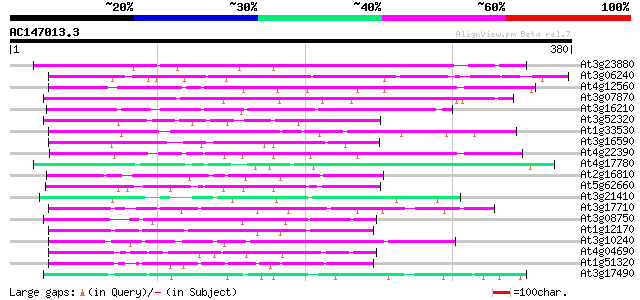
Score E
Sequences producing significant alignments: (bits) Value
At3g23880 unknown protein 155 3e-38
At3g06240 unknown protein 99 3e-21
At4g12560 putative protein 97 1e-20
At3g07870 unknown protein 91 8e-19
At3g16210 hypothetical protein 87 1e-17
At3g52320 putative protein 83 2e-16
At1g33530 hypothetical protein 80 2e-15
At3g16590 hypothetical protein 78 9e-15
At4g22390 unknown protein 77 2e-14
At4g17780 MYB transcription factor like protein 75 5e-14
At2g16810 unknown protein 73 2e-13
At5g62660 putative protein 73 3e-13
At3g21410 hypothetical protein 72 5e-13
At3g17710 unknown protein 71 9e-13
At3g08750 hypothetical protein 71 9e-13
At1g12170 hypothetical protein 71 1e-12
At3g10240 hypothetical protein 70 1e-12
At4g04690 70 2e-12
At1g51320 hypothetical protein 69 6e-12
At3g17490 hypothetical protein 67 1e-11
>At3g23880 unknown protein
Length = 364
Score = 155 bits (393), Expect = 3e-38
Identities = 110/350 (31%), Positives = 178/350 (50%), Gaps = 28/350 (8%)
Query: 17 ETTANKPLPF-LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYP 75
ETT P LP E++ ILLRLPV+SL RFKCVC SW++L S+T FA H LI
Sbjct: 3 ETTREMFSPHNLPLEMMEEILLRLPVKSLTRFKCVCSSWRSLISETLFALKHALILETSK 62
Query: 76 QLVACES---VSAYRTWEIKTYPIESLLENSSTTVIPVSN---TGHQRYTILGSCNGFLC 129
+ +S V + +K+ I SL N+ST + + G Y ++G+C+G +C
Sbjct: 63 ATTSTKSPYGVITTSRYHLKSCCIHSLY-NASTVYVSEHDGELLGRDYYQVVGTCHGLVC 121
Query: 130 LYDNYQRCVRLWNPSINLKSKSSPT------IDRFIYYGFGYDQVNHKYKLLAV--KAFS 181
+ +Y + + LWNP+I L+ + S + + + YGFGYD+ YK++A+ +
Sbjct: 122 FHVDYDKSLYLWNPTIKLQQRLSSSDLETSDDECVVTYGFGYDESEDDYKVVALLQQRHQ 181
Query: 182 RITETMIYTFGENSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRATILSFDIE 241
ET IY+ + ++ ++ G +++GTLNW TI+S+D+
Sbjct: 182 VKIETKIYSTRQKLWRSNTSFPSGVVVADKSRSGIYINGTLNWAATSSSSSWTIISYDMS 241
Query: 242 KETYRQVLLPQ-HGYAVYSPGLYVLSNCICVCTSFLDTRWQLWMMKKYGVAESWTKLMSI 300
++ ++++ P G ++ L L C+ + +W+MK++G SW+KL+SI
Sbjct: 242 RDEFKELPGPVCCGRGCFTMTLGDLRGCLSMVCYCKGANADVWVMKEFGEVYSWSKLLSI 301
Query: 301 PHENLLISNISLWPCVEPLFISENGVVLLMNTSSSQLILYNLNSRGLYFP 350
P L V PL+IS+ VVLL S L LYN ++ ++P
Sbjct: 302 P---------GLTDFVRPLWISDGLVVLL--EFRSGLALYNCSNGRFHYP 340
>At3g06240 unknown protein
Length = 427
Score = 99.4 bits (246), Expect = 3e-21
Identities = 103/411 (25%), Positives = 185/411 (44%), Gaps = 85/411 (20%)
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHF-------LISTVYPQLVA 79
LP E+I ILLRLP +S+ RF+CV K + TL SD FA H + +++ +L+
Sbjct: 36 LPPEIITEILLRLPAKSIGRFRCVSKLFCTLSSDPGFAKIHLDLILRNESVRSLHRKLI- 94
Query: 80 CESVSAYRTWEIK----------------TYPIES---------------LLENSSTTVI 108
VS++ + + YP++ L + ++
Sbjct: 95 ---VSSHNLYSLDFNSIGDGIRDLAAVEHNYPLKDDPSIFSEMIRNYVGDHLYDDRRVML 151
Query: 109 PVSNTGHQR--YTILGSCNGFLCLYDNYQRCVRLWNPSI--------NLKSKSSP-TIDR 157
++ ++R I+GS NG +C+ + V L+NP+ N + KS D
Sbjct: 152 KLNAKSYRRNWVEIVGSSNGLVCISPG-EGAVFLYNPTTGDSKRLPENFRPKSVEYERDN 210
Query: 158 FIYYGFGYDQVNHKYKLLAVKAFSR-ITETMIYTFGENSCKNVEVKDFPRYPPNRKHLGK 216
F YGFG+D + YKL+ + A S I + +Y+ +S + + ++ + G
Sbjct: 211 FQTYGFGFDGLTDDYKLVKLVATSEDILDASVYSLKADSWRRICNLNY-EHNDGSYTSGV 269
Query: 217 FVSGTLNWIVDE-RDGRATILSFDIEKETYRQVLLPQ------HGYAVYSPGLYVLSNCI 269
+G ++W+ E R + +++FDI+ E +R++ +P H ++ + G L+ +
Sbjct: 270 HFNGAIHWVFTESRHNQRVVVAFDIQTEEFREMPVPDEAEDCSHRFSNFVVGS--LNGRL 327
Query: 270 CVCTSFLDTRWQLWMMKKYGVAESWTKLMSIPHENLLISNISLWPCVEPLFISENGVVLL 329
CV S D +W+M +YG A+SW+++ NL L+ ++PL ++N +L
Sbjct: 328 CVVNSCYDVHDDIWVMSEYGEAKSWSRI----RINL------LYRSMKPLCSTKNDEEVL 377
Query: 330 MNTSSSQLILYNLNSRGLYFPRIPPSYLNI--YYIEPKLDLHIYHESLLSP 378
+ L+LYN + S L I + + + Y ESL+SP
Sbjct: 378 LEL-DGDLVLYNFETNA-------SSNLGICGVKLSDGFEANTYVESLISP 420
>At4g12560 putative protein
Length = 408
Score = 97.4 bits (241), Expect = 1e-20
Identities = 86/367 (23%), Positives = 168/367 (45%), Gaps = 49/367 (13%)
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESVSAY 86
+P +++ I LRLP ++L+R + + K L +D F +H ++ L + +
Sbjct: 4 IPMDIVNDIFLRLPAKTLVRCRALSKPCYHLINDPDFIESH-----LHRVLQTGDHLMIL 58
Query: 87 RTWEIKTYPIE-SLLENSSTTVIPVSNTGHQRYTILGSCNGFLCLYDNYQRCVRLWNPSI 145
++ Y ++ L++ S P+ G + GS NG + L N + ++NPS
Sbjct: 59 LRGALRLYSVDLDSLDSVSDVEHPMKRGGPTE--VFGSSNGLIGL-SNSPTDLAVFNPST 115
Query: 146 NLKSKSSPT-IDR--------FIYYGFGYDQVNHKYKLLAVKAF----------SRITET 186
+ P+ ID +++YG GYD V+ YK++ + F S E
Sbjct: 116 RQIHRLPPSSIDLPDGSSTRGYVFYGLGYDSVSDDYKVVRMVQFKIDSEDELGCSFPYEV 175
Query: 187 MIYTFGENSCKNVE--------VKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRAT---I 235
+++ +NS K +E + F + R+ G +L+W++ R G I
Sbjct: 176 KVFSLKKNSWKRIESVASSIQLLFYFYYHLLYRRGYGVLAGNSLHWVLPRRPGLIAFNLI 235
Query: 236 LSFDIEKETYRQVLLPQ---HGYAVYSPGLYVLSNCICVCTSFLDTRWQLWMMKKYGVAE 292
+ FD+ E + V P+ +G + VL C+C+ ++ + +WMMK+Y V +
Sbjct: 236 VRFDLALEEFEIVRFPEAVANGNVDIQMDIGVLDGCLCLMCNYDQSYVDVWMMKEYNVRD 295
Query: 293 SWTKLMSIPHENLLISNISLWPCVEPLFISENGVVLLMNTSSSQLILYNLNSRGLYFPRI 352
SWTK+ ++ ++ + + PL S++ +L+ ++++L+ ++L S+ + RI
Sbjct: 296 SWTKVFTVQKP----KSVKSFSYMRPLVYSKDKKKVLLELNNTKLVWFDLESKKMSTLRI 351
Query: 353 ---PPSY 356
P SY
Sbjct: 352 KDCPSSY 358
>At3g07870 unknown protein
Length = 417
Score = 91.3 bits (225), Expect = 8e-19
Identities = 90/359 (25%), Positives = 156/359 (43%), Gaps = 44/359 (12%)
Query: 24 LPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESV 83
L LPE++I I RLP+ S+ R VC+SW+++ + ++ T L+ C+S
Sbjct: 25 LESLPEDIIADIFSRLPISSIARLMFVCRSWRSVLTQHGRLSSSSSSPTKPCLLLHCDSP 84
Query: 84 SAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFLCLYDN-YQRCVRLWN 142
+ E ++ T+ S+ + ++GSCNG LCL D+ Y + L+N
Sbjct: 85 IRNGLHFLDLSEEEKRIKTKKFTLRFASSM--PEFDVVGSCNGLLCLSDSLYNDSLYLYN 142
Query: 143 P----SINLKSKSSPTIDRFIYYGFGYDQVNHKYKLLAVKAFS----------------- 181
P S+ L S+ D+ + +GFG+ ++ +YK+L + F
Sbjct: 143 PFTTNSLELPECSNKYHDQELVFGFGFHEMTKEYKVLKIVYFRGSSSNNNGIYRGRGRIQ 202
Query: 182 -RITETMIYTFGENSC-KNVEVKDFPRYPPN--RKHLGKFVSGTLNWIVDERD--GRATI 235
+ +E I T + +++ + + P ++ V+G L+++ R
Sbjct: 203 YKQSEVQILTLSSKTTDQSLSWRSLGKAPYKFVKRSSEALVNGRLHFVTRPRRHVPDRKF 262
Query: 236 LSFDIEKETYRQVLLPQ-HGYAVYSPGLYVLSNCICVCTSFLDTRWQLWMMKKYGVAESW 294
+SFD+E E ++++ P G + L L C+C + +W+MK YGV ESW
Sbjct: 263 VSFDLEDEEFKEIPKPDCGGLNRTNHRLVNLKGCLCAVVYGNYGKLDIWVMKTYGVKESW 322
Query: 295 TKLMSIP-------HENL-----LISNISLWPCVEPLFISENGVVLLMNTSSSQLILYN 341
K SI +NL + N V L + ENG +LL S L+ Y+
Sbjct: 323 GKEYSIGTYLPKGLKQNLDRPMWIWKNAENGKVVRVLCLLENGEILL-EYKSRVLVAYD 380
>At3g16210 hypothetical protein
Length = 360
Score = 87.4 bits (215), Expect = 1e-17
Identities = 77/279 (27%), Positives = 127/279 (44%), Gaps = 17/279 (6%)
Query: 26 FLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESVSA 85
FLPEEL + IL+RL ++ L RF+CVCK+W+ L +D F + +S + V+ +
Sbjct: 4 FLPEELAIEILVRLSMKDLARFRCVCKTWRDLINDPGFTETYRDMSPA--KFVSFYDKNF 61
Query: 86 YRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFLCLYDNYQRCVRLWNP-S 144
Y +P+ ++ P+ + T + C+G LC+ + +WNP S
Sbjct: 62 YMLDVEGKHPV-----ITNKLDFPLDQSMIDESTCVLHCDGTLCV-TLKNHTLMVWNPFS 115
Query: 145 INLKSKSSPTI--DRFIYYGFGYDQVNHKYKLLAVKAFSRITETMIYTFGENSCKNVEVK 202
K +P I D I GFGYD V+ YK++ ++ ++ F S
Sbjct: 116 KQFKIVPNPGIYQDSNI-LGFGYDPVHDDYKVVTFIDRLDVSTAHVFEFRTGSWGESLRI 174
Query: 203 DFPRYPPNRKHLGKFVSGTLNWIVDERDGRATILSFDIEKETYRQVLLPQHGYAVYSPGL 262
+P + R G F+ L WI IL F++ YR++ LP + V S L
Sbjct: 175 SYPDW-HYRDRRGTFLDQYLYWIAYRSSADRFILCFNLSTHEYRKLPLPVYNQGVTSSWL 233
Query: 263 YVLSNCICVCT-SFLDTRWQLWMMKKYGVAESWTKLMSI 300
V S +C+ ++ +M+K G SW+K++S+
Sbjct: 234 GVTSQKLCITEYEMCKKEIRISVMEKTG---SWSKIISL 269
>At3g52320 putative protein
Length = 390
Score = 83.2 bits (204), Expect = 2e-16
Identities = 69/252 (27%), Positives = 118/252 (46%), Gaps = 39/252 (15%)
Query: 24 LPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLIST---VYPQLVAC 80
LP +PEE+++ IL+RLP +SL+RFKCV K W +L + +F N F S+ ++ LV
Sbjct: 24 LPEIPEEMLIDILIRLPAKSLMRFKCVSKLWLSLITSRYFTNRFFKPSSPSCLFAYLVDR 83
Query: 81 ESVSAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILG-----SCNGFLCLYDNYQ 135
E+ S Y + + S ++S T+V + H I+G + G LC
Sbjct: 84 ENQSKYLLLQSSS---SSRHDHSDTSVSVIDQ--HSTIPIMGGYLVNAARGLLCYRTG-- 136
Query: 136 RCVRLWNPSIN-------LKSKSSPTIDRFIYYGFGYDQVNHKYKLLAVKAFSRITETMI 188
R V++ NPS ++SK++ ++ FG+D + +YK+L++ F +T+
Sbjct: 137 RRVKVCNPSTRQIVELPIMRSKTN------VWNWFGHDPFHDEYKVLSL--FWEVTKEQT 188
Query: 189 YTFGEN---------SCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRATILSFD 239
E+ S +N + P P + G + G L + R ++SFD
Sbjct: 189 VVRSEHQVLVLGVGASWRNTKSHHTPHRPFHPYSRGMTIDGVLYYSARTDANRCVLMSFD 248
Query: 240 IEKETYRQVLLP 251
+ E + + LP
Sbjct: 249 LSSEEFNLIELP 260
>At1g33530 hypothetical protein
Length = 441
Score = 80.1 bits (196), Expect = 2e-15
Identities = 80/333 (24%), Positives = 139/333 (41%), Gaps = 27/333 (8%)
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHF-LISTVY--PQLVACESV 83
LP+ L+ IL RLPV+ L+R K + K WK+L H A H L+ Y ++
Sbjct: 97 LPDVLVEEILQRLPVKYLVRLKSISKGWKSLIESDHLAEKHLRLLEKKYGLKEIKITVER 156
Query: 84 SAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFLCLYDNYQRCVRLWNP 143
S ++ IK + S + +++ + GSCNG +C+Y+ + L NP
Sbjct: 157 STSKSICIKFFSRRSGMN-------AINSDSDDLLRVPGSCNGLVCVYELDSVYIYLLNP 209
Query: 144 SINLKSKSSPTIDRFIYYGFGYDQVNHKYKLLAVKAFSRITETMIYTFGENSCKNVEVKD 203
+ +P + GFG D V YK++ + F R+ T+++ N + K
Sbjct: 210 MTGVTRTLTPPRGTKLSVGFGIDVVTGTYKVMVLYGFDRV-GTVVFDLDTNKWRQ-RYKT 267
Query: 204 FPRYP----PNRKHLGKFVSGTLNWIVDERDGRATILSFDIEKETYRQVLLPQHGYAVYS 259
P P + FV+G+L W++ + IL D+ E +R + P V
Sbjct: 268 AGPMPLSCIPTPERNPVFVNGSLFWLL--ASDFSEILVMDLHTEKFRTLSQPNDMDDVDV 325
Query: 260 PGLYV----LSNCICVCTSFLDTRWQLWMMKKYGVAESW--TKLMSIPHENLLISNISLW 313
Y+ L + +CV +W++ + ++E W T+ + H +S S W
Sbjct: 326 SSGYIYMWSLEDRLCVSNVRQGLHSYVWVLVQDELSEKWERTRFNLLGHVFPPLSLNSAW 385
Query: 314 ---PCVEPLFISENGVVLLMNTSSSQLILYNLN 343
V P +S + + +S L++ N
Sbjct: 386 FSQTLVSPYQLSSSTCIGSRQRQNSTSALFSRN 418
>At3g16590 hypothetical protein
Length = 374
Score = 77.8 bits (190), Expect = 9e-15
Identities = 74/249 (29%), Positives = 111/249 (43%), Gaps = 40/249 (16%)
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNH-------FLISTVYPQLVA 79
LP EL ILLR+P SL RF+ VCK W TLF+D F NNH F++ T +
Sbjct: 5 LPLELEDEILLRVPPLSLTRFRTVCKRWNTLFNDQRFINNHLACVRPQFILRTEKDSKIY 64
Query: 80 CESVSAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFL---CLYDNYQR 136
++ + E++ +E+ N V Y L C+GFL L D
Sbjct: 65 SIGINIDDSLEVRELNLETQGPNKKLKV----------YRNLFYCDGFLLCPALLDE--- 111
Query: 137 CVRLWNPSINLKSK-SSPTIDRFIYYGFGYDQVNHK--YKLLAV---------KAFSRIT 184
V +WNP + ++K P RF YG GYD + YK+L +++RI
Sbjct: 112 -VAVWNPWLRKQTKWIEPKRSRFNLYGLGYDNRRPEKCYKILGFGYGYSSEINGSYNRIN 170
Query: 185 -ETMIYTFGENSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIV--DERDGRATILSFDIE 241
++ F N+ K+++ F + + + + ++GTL WI E G I SFD
Sbjct: 171 PRVSVFEFETNAWKDLKFGLFDWHLRSPRTV-LSLNGTLYWIAVRCESGGDGFIQSFDFS 229
Query: 242 KETYRQVLL 250
+E + L
Sbjct: 230 REMFEPFCL 238
>At4g22390 unknown protein
Length = 401
Score = 76.6 bits (187), Expect = 2e-14
Identities = 80/352 (22%), Positives = 150/352 (41%), Gaps = 45/352 (12%)
Query: 28 PEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFL--ISTVYPQLVACESVSA 85
P +LI + LRL +L++ + + K +L F ++H + T ++
Sbjct: 5 PTDLINEMFLRLRATTLVKCRVLSKPCFSLIDSPEFVSSHLRRRLETGEHLMILLRGPRL 64
Query: 86 YRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFLCLYDNYQRCVRLWNPS- 144
RT E+ + EN S P+ G + GS NG + L N + ++NPS
Sbjct: 65 LRTVELDSP------ENVSDIPHPLQAGGFTE--VFGSFNGVIGLC-NSPVDLAIFNPST 115
Query: 145 -----INLKSKSSPTID---RFIYYGFGYDQVNHKYKLLAV---------KAFSRITETM 187
+ ++ P D +++YG GYD V +K++ + K F E
Sbjct: 116 RKIHRLPIEPIDFPERDITREYVFYGLGYDSVGDDFKVVRIVQCKLKEGKKKFPCPVEVK 175
Query: 188 IYTFGENSCKNVEVK--------DFPRYPPNRKHLGKFVSGTLNWIVDERDGRAT---IL 236
+++ +NS K V + + + R+ G V+ L+WI+ R G I+
Sbjct: 176 VFSLKKNSWKRVCLMFEFQILWISYYYHLLPRRGYGVVVNNHLHWILPRRQGVIAFNAII 235
Query: 237 SFDIEKETYRQVLLPQHGYAVYSPGLYVLSNCICVCTSFLDTRWQLWMMKKYGVAESWTK 296
+D+ + + PQ Y + + VL C+C+ + +W++K+Y +SWTK
Sbjct: 236 KYDLASDDIGVLSFPQELYIEDNMDIGVLDGCVCLMCYDEYSHVDVWVLKEYEDYKSWTK 295
Query: 297 LMSIPHENLLISNISLWPCVEPLFIS-ENGVVLLMNTSSSQLILYNLNSRGL 347
L +P ++ + PL S + +LL +++ L+ ++L S+ L
Sbjct: 296 LYRVPKP----ESVESVEFIRPLICSKDRSKILLEINNAANLMWFDLESQSL 343
>At4g17780 MYB transcription factor like protein
Length = 745
Score = 75.5 bits (184), Expect = 5e-14
Identities = 81/370 (21%), Positives = 143/370 (37%), Gaps = 38/370 (10%)
Query: 17 ETTANKPLPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQ 76
E N ++ +L+ I LRLP++S+L K V K W+++ F +
Sbjct: 365 EEEKNPSSIYIVADLLEDIFLRLPLKSILISKSVSKRWRSILESKTFVERRMSLQKKRKI 424
Query: 77 LVACESVSAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFLCLYDNYQR 136
L A WE + P S + + V N +T C+G +C+ + R
Sbjct: 425 LAAYNCKCG---WEPRLLPGSSQCKGNEEIVYLHCNAAQPSFT----CDGLVCILE--PR 475
Query: 137 CVRLWNPSINLKSKSSPTIDRFIYYGFGY------DQVNHKYKL--LAVKAFSRITETMI 188
+ + NP + YGFG+ D+V YK+ + + +FS I
Sbjct: 476 WIDVLNPWTRQLRR----------YGFGFGTIFGVDKVTGSYKVVKMCLISFSEICARDP 525
Query: 189 YTFGENSCKNVEVKDF-----PRYPPNRKHLGKFVSGTLNWIVDERDGRATILSFDIEKE 243
E S +VE ++ P Y + +G++ W+ + TIL+ D+ KE
Sbjct: 526 EV--EYSVLDVETGEWRMLSPPPYKVFEVRKSECANGSIYWLHKPTERAWTILALDLHKE 583
Query: 244 TYRQVLLPQHGYAVYSPGLYVLSNCICVCTSFLDTRWQLWMMKKYGVAESWTKLMSIPHE 303
+ +P + + L + + + ++ T W+L + E+WTK SI E
Sbjct: 584 ELHNISVPDMSVTQETFQIVNLEDRLAIANTYTKTEWKLEIWSMDTEVETWTKTYSIDLE 643
Query: 304 NLLISNISLWPCVEPLFISENGVVLLMNTSSSQLILYNLNSRGLYFPR----IPPSYLNI 359
N + S P+ +S+ G ++ + Y + LY I P + N+
Sbjct: 644 NRVASRERRNRWFTPVSVSKQGNIVFYDNHKRLFKYYPRKNEILYLSADTCVISPFFENL 703
Query: 360 YYIEPKLDLH 369
+ K LH
Sbjct: 704 APLPQKSTLH 713
>At2g16810 unknown protein
Length = 295
Score = 73.2 bits (178), Expect = 2e-13
Identities = 67/251 (26%), Positives = 112/251 (43%), Gaps = 35/251 (13%)
Query: 26 FLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESVSA 85
++P++L++ IL RLP +S++RFKCV K W +L S +F N FLI PQ S
Sbjct: 40 YIPQDLLIEILTRLPPKSVMRFKCVSKFWSSLLSSRYFC-NRFLIVPSQPQ------PSL 92
Query: 86 YRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTI-------LGSCNGFLCLYDNYQRCV 138
Y + +SL+ +S+ + P S Q TI L GF+ N +
Sbjct: 93 YMCLLDRYNYSKSLILSSAPSTSPYSFVFDQDLTIRKMGGFFLRILRGFIFFTRNLK--A 150
Query: 139 RLWNPSINLKSKSSPTIDRF---------IYYGFGYDQVNHKYKLLAVKAFSR------- 182
R++NP+ + PTI I Y +D VN +YKLL +++
Sbjct: 151 RIYNPTTR-QLVILPTIKESDIIAGPPYNILYFICHDPVNDRYKLLCTVSYASDNDLQNL 209
Query: 183 ITETMIYTFGENSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRATILSFDIEK 242
+E I+ +FP + P+ HL ++G L ++ ++SFD+
Sbjct: 210 KSELWIFVLEAGGSWKRVANEFPHHVPS--HLDLNMNGVLYFLAWTDPHTCMLVSFDVRS 267
Query: 243 ETYRQVLLPQH 253
E + + +P++
Sbjct: 268 EEFNTMQVPRN 278
>At5g62660 putative protein
Length = 379
Score = 72.8 bits (177), Expect = 3e-13
Identities = 72/258 (27%), Positives = 117/258 (44%), Gaps = 46/258 (17%)
Query: 25 PFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLIST------VYPQLV 78
P +P +L++ IL +LP +SL+RFKCV K W +L F+N + + T +Y LV
Sbjct: 39 PEIPLDLLIEILTKLPAKSLMRFKCVSKLWSSLIRSRFFSNCYLTVKTPRRPPRLYMSLV 98
Query: 79 ---ACESVSAYRTWEIKTYPIESLLENSSTTV------IPVSNTGHQRYTILGSCNGFLC 129
C S+ + YP ES+L +SS++ + ++ G + +L L
Sbjct: 99 DHLLCNSLM------VCHYPCESVLLSSSSSAESLEQNLTIAGMGGRNMVVLRG----LI 148
Query: 130 LYDNYQRCVRLWNP----SINLKSKSSPTIDR-----FIYYGFGYDQVNHKYKLLAV--- 177
LY R ++NP S+ L + S + + + Y FGYD V +YK++
Sbjct: 149 LY-VVCRTASIYNPTTRQSVTLPAVKSNILAQKSHWNSLLYFFGYDPVLDQYKVVCTVAL 207
Query: 178 --KAFSRIT-ETMIYTFGE-NSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRA 233
K RIT E ++ S K +E D P P LG V+G + ++ R
Sbjct: 208 FSKRLKRITSEHWVFVLEPGGSWKRIEF-DQPHLP---TRLGLCVNGVIYYLASTWKRRD 263
Query: 234 TILSFDIEKETYRQVLLP 251
++SFD+ E + + P
Sbjct: 264 IVVSFDVRSEEFSMIQGP 281
>At3g21410 hypothetical protein
Length = 374
Score = 72.0 bits (175), Expect = 5e-13
Identities = 81/310 (26%), Positives = 123/310 (39%), Gaps = 52/310 (16%)
Query: 21 NKPLPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHF------------ 68
+K LP EL IL R+P +SLLR K CK W LF+D F H
Sbjct: 5 SKECSLLPFELFEEILCRVPTKSLLRLKLTCKRWLALFNDKRFIYKHLALVREHIIRTNQ 64
Query: 69 LISTVYPQLVACESVSAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNG-F 127
++ + P + AC S S +++K T++P C+G
Sbjct: 65 MVKIINPVVGACSSFSLPNKFQVK---------GEIYTMVP--------------CDGLL 101
Query: 128 LCLYDNYQRCVRLWNPSINLKS---KSSPTIDRFIYYGFGYDQVNH-KYKLLAVKA--FS 181
LC+++ V WNP +N +P+ YG GYD ++ YK+L A S
Sbjct: 102 LCIFETGSMAV--WNPCLNQVRWIFLLNPSFRGCSCYGIGYDGLSRDSYKILRFYANTGS 159
Query: 182 RITETMIYTFGENSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVD-ERDGRATILSFDI 240
E IY NS K +V + + G + G + WI R I SF+
Sbjct: 160 YKPEVDIYELKSNSWKTFKVS--LDWHVVLRCKGLSLKGNMYWIAKWNRKPDIFIQSFNF 217
Query: 241 EKETYRQVLLPQHGYAVYSP---GLYVLSNCICVCTSFLDTRWQLWMMKKY--GVAESWT 295
ET+ + Y V++ + N + S ++ +W+ K GV+ WT
Sbjct: 218 STETFEPLCSLPVRYDVHNVVALSAFKGDNLSLLHQSKETSKIDVWVTNKVKNGVSILWT 277
Query: 296 KLMSIPHENL 305
KL S+ +L
Sbjct: 278 KLFSVTRPDL 287
>At3g17710 unknown protein
Length = 368
Score = 71.2 bits (173), Expect = 9e-13
Identities = 88/333 (26%), Positives = 139/333 (41%), Gaps = 52/333 (15%)
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESVSAY 86
LP +L IL RLP RSL+RF+ VCK W LFSD F H + PQ + +
Sbjct: 6 LPWDLEEEILSRLPPRSLVRFRTVCKHWNGLFSDKRFVKKHLV--RARPQFI-------F 56
Query: 87 RTWEIKTYPIESLLENSSTTVIPVSNTGH-----QRYTILGSCNGFLCLYDNYQRCVRLW 141
T K Y IE L + V V H + +T + +C+G L D +++ V +W
Sbjct: 57 LTESKKMYSIEIDL-GGTIEVREVPYDFHCQPMKKNFTTIMACDGLL-FRDFWKQGVAVW 114
Query: 142 NPSINLKSKSSPTIDRFIYYGFGYD--QVNHKYKLLA-VKAFSRITETM--------IYT 190
NP + F + G GYD + + YK+L R+++++ IY
Sbjct: 115 NPWLRQVGWIEYEDKGFRFCGVGYDSCKPDKCYKILGYFNCTRRLSDSLQEGYYQAAIYE 174
Query: 191 FGENSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRAT--ILSFDIEKETYRQ- 247
+ K ++ + PN+ L V+G L W+ I +FD E ++
Sbjct: 175 CASQAFKFIDTPNPFNLWPNKDPLS--VNGNLYWLAHNHPETLEYFIETFDFSMEIFKPF 232
Query: 248 VLLP--------QHGYAVYSPGLY-VLSNCICVCTSFLDTRWQLWMMKKYGVAES---WT 295
LLP + AV+ + +L C F T+ ++W+ K E W
Sbjct: 233 CLLPCRKDFGSNELVLAVFKEDRFSLLKQC------FETTKIEIWVTKMKIDREEEVVWI 286
Query: 296 KLMSIPHENLLISNISLWPCVEPLFISENGVVL 328
K M++P NL N+ C FI + +++
Sbjct: 287 KFMTLPTTNL--PNLDDTYCCSSYFIFDKTIIM 317
>At3g08750 hypothetical protein
Length = 369
Score = 71.2 bits (173), Expect = 9e-13
Identities = 66/238 (27%), Positives = 100/238 (41%), Gaps = 29/238 (12%)
Query: 24 LPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESV 83
LP LP ELI IL ++P SL+RFK CK W L ++ F NH +
Sbjct: 9 LPSLPFELIEEILYKIPAESLIRFKSTCKKWYNLITEKRFMYNHL------------DHY 56
Query: 84 SAYRTWEIKTYP---IESLLENSSTTVIPVSNTGHQRYTILGSCNG-FLCLYDNYQRCVR 139
S R I+TY I+ + E S +IP + C+G LC + +
Sbjct: 57 SPERF--IRTYDQQIIDPVTEILSDALIPDEFRDLYPIYSMVHCDGLMLCTCRKWDNSLA 114
Query: 140 LWNPSINLKSKSSPTID--RFIYYGFGYDQVNHKYKLLAVKAFSRITE-------TMIYT 190
+WNP + P++ Y G GYD + +K R+ + IY
Sbjct: 115 VWNPVLREIKWIKPSVCYLHTDYVGIGYDDNVSRDNYKILKLLGRLPKDDDSDPNCEIYE 174
Query: 191 FGENSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRATILSFDIEKETYRQV 248
F +S K + K F R + G V G + WI +++ TI+ FD ET++++
Sbjct: 175 FKSDSWKTLVAK-FDWDIDIRCNNGVSVKGKMYWIAKKKED-FTIIRFDFSTETFKEI 230
>At1g12170 hypothetical protein
Length = 364
Score = 70.9 bits (172), Expect = 1e-12
Identities = 63/233 (27%), Positives = 98/233 (42%), Gaps = 20/233 (8%)
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESVSAY 86
LP EL+ IL R+P SL RFK VCK W TLF F NNH + V PQ + Y
Sbjct: 6 LPWELVEEILYRVPPLSLTRFKIVCKQWNTLFKSKSFVNNHLV--RVRPQFLLWTDSKMY 63
Query: 87 RTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFL-CLYDNYQRCVRLWNPSI 145
+ + + IP N + Y C+G L C ++++ +WNP +
Sbjct: 64 SV-SVNLNDDQKIDMRELPLDIPYLNNFMRTY--FTPCDGLLFCDSWSWRKKAAIWNPWL 120
Query: 146 NLKSKSSPTIDR-FIYYGFGYD--QVNHKYKLLAVKAFSR--------ITETMIYTFGEN 194
+ ++ F + G GYD + + +K++ +++ IYTF N
Sbjct: 121 RQTKWIEYSKEKTFTFRGIGYDSGRPDKGHKIIGSSIYNKRKLIEDPLYRSVEIYTFETN 180
Query: 195 SCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRATIL-SFDIEKETYR 246
K++ F R ++G L W+V + + SFD KE Y+
Sbjct: 181 GWKSMNT--FSEEGEIRPLSDVSLNGNLYWVVSNGETHECFIESFDFSKEIYK 231
>At3g10240 hypothetical protein
Length = 389
Score = 70.5 bits (171), Expect = 1e-12
Identities = 70/296 (23%), Positives = 130/296 (43%), Gaps = 34/296 (11%)
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVAC------ 80
+P +L+ ILLRLP +S+ RF+CV K W ++ ++ +F N L++T P+L+ C
Sbjct: 27 IPLDLVSEILLRLPEKSVARFRCVSKPWSSITTEPYFIN---LLTTRSPRLLLCFKANEK 83
Query: 81 -------ESVSAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFLCLYDN 133
+ + TW K++ L++ N + S NG +C ++
Sbjct: 84 FFVSSIPQHRQTFETWN-KSHSYSQLIDRYHMEFSEEMN----YFPPTESVNGLICFQES 138
Query: 134 YQRCVRLWNPS----INLKSKSSPTIDRFIYYGFGYDQVNHKYKLLAVKAFSRITETMIY 189
+ V WNPS + L + + D I+ GYD V K+K++ ++ + +
Sbjct: 139 ARLIV--WNPSTRQLLILPKPNGNSNDLTIF--LGYDPVEGKHKVMCMEFSATYDTCRVL 194
Query: 190 TFGENSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRATIL-SFDIEKETYRQV 248
T G VK ++ + G+ ++G + I +D +L SFD+ E + +
Sbjct: 195 TLGSAQKLWRTVKTHNKHRSDYYDSGRCINGVVYHIAYVKDMCVWVLMSFDVRSEIFDMI 254
Query: 249 LLPQHGYAVYSPGLYVLSNCI-CVCTSFLDTRW-QLWMMKKYGVAESWTKLMSIPH 302
LP V+ L + + CV ++ +LW+++K+ S L + H
Sbjct: 255 ELPSSD--VHKDVLIDYNGRLACVGREIIEKNGIRLWILEKHNKWSSKDFLAPLVH 308
>At4g04690
Length = 378
Score = 70.1 bits (170), Expect = 2e-12
Identities = 65/235 (27%), Positives = 102/235 (42%), Gaps = 28/235 (11%)
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESVSAY 86
LP EL+ IL + P SL RFK CK W + + F NH S P+ + +
Sbjct: 11 LPFELVEEILKKTPAESLNRFKSTCKQWYGIITSKRFMYNHLDHS---PERFI--RIDDH 65
Query: 87 RTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGF---LCLYDNYQRC----VR 139
+T +I P+ + +S +P + + C+G +C +Y+R +
Sbjct: 66 KTVQIMD-PMTGIFSDSP---VPDVFRSPHSFASMVHCDGLMLCICSDSSYERTREANLA 121
Query: 140 LWNPSINLKSKSSPTIDRFI---YYGFGYDQV-NHKYKLLAVKAFSRI--TETMIYTFGE 193
+WNP + K K +D + Y+G GYD YK++ TE IY F
Sbjct: 122 VWNP-VTKKIKWIEPLDSYYETDYFGIGYDNTCRENYKIVRFSGPMSFDDTECEIYEFKS 180
Query: 194 NSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRATILSFDIEKETYRQV 248
+S + ++ K + Y R G V G + WI D ++ IL FD ET++ V
Sbjct: 181 DSWRTLDTKYWDVYTQCR---GVSVKGNMYWIADTKE--KFILRFDFSMETFKNV 230
>At1g51320 hypothetical protein
Length = 375
Score = 68.6 bits (166), Expect = 6e-12
Identities = 69/231 (29%), Positives = 112/231 (47%), Gaps = 26/231 (11%)
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESVSAY 86
LP EL+ IL R+P +SL++F+ VCK W +LF D F N+HF+ S PQL+ +
Sbjct: 6 LPWELVEEILCRVPPQSLVKFRTVCKQWNSLFDDNKFVNDHFVQS--QPQLI-------F 56
Query: 87 RTWEIKTYPIESLLENSSTTV--IPVSNTGHQ-----RYTILGSCNGFLCLYDNYQRCVR 139
RT E K Y + + V +P++ G + R C+G L +Y V
Sbjct: 57 RT-ESKIYSVAVNFKGPRIEVHELPLAIPGLKSEMPIRLHDYVDCDGLL-FCTSYFNGVL 114
Query: 140 LWNPSINLKSKSSPTIDRF-IYYGFGYDQVNHKYKLL---AVKAFSRITETMIYTFGENS 195
+WNP + +++ P+I R+ + Y GYD +YK+L + S I +++IY G S
Sbjct: 115 IWNPWLR-QTRFFPSIHRYPVTYDIGYDN-KKQYKMLDYYKCEGDSSI-KSIIYEIGLYS 171
Query: 196 CKNVEVKDFPRYPPNRKHLGKFVSGTLNWIVDERDGRATILSFDIEKETYR 246
K +++ + + + ++GTL W + + I SF E R
Sbjct: 172 KKVKDLESDSTWSIFQSN-SVSLNGTLYWAGVDVNNGIFIRSFSFSTEKPR 221
>At3g17490 hypothetical protein
Length = 388
Score = 67.4 bits (163), Expect = 1e-11
Identities = 90/368 (24%), Positives = 147/368 (39%), Gaps = 61/368 (16%)
Query: 24 LPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESV 83
+P L E+L+ IL R+P SL R + CK W LF+D F+ H P+ ++
Sbjct: 3 MPHLSEDLVEEILSRVPAISLKRLRYTCKQWNALFNDQRFSKKH---RDKAPKTYLGLTL 59
Query: 84 SAYRTWEIKTYPIESLLENSSTTVI--------PVSNTGHQRYTILGSCNGFLCLYDNYQ 135
+R + + + + LL N++ ++ +++ + + C+G +
Sbjct: 60 KDFRIYSMSS-NLHGLLHNNNIDLLMEFKGKLSSLNDLNDFEISQIYPCDGLILCSTKRN 118
Query: 136 RCVRLWNPSIN----LKSKSSPTIDRFIYYGFGYDQVN----HKYKLLAV--KAFSRITE 185
+ +WNP +K ++ D F FGYD + YK+L V K + E
Sbjct: 119 TRLVVWNPCTGQTRWIKRRNRRMCDTF---AFGYDNSKSSCLNNYKILRVCEKIKGQQFE 175
Query: 186 TMIYTFGENSCKNVEVKDFPRYPPNRKHLGKFVS--GTLNWIVDERDGRATILSFDIEKE 243
I+ F NS + ++V PN G+ VS G W I FD E
Sbjct: 176 YEIFEFSSNSWRVLDVN------PNCIIEGRSVSVKGNSYWFATITKTHYFIRRFDFSSE 229
Query: 244 TYRQVLLPQHGY---------AVYSPGLYVLSNCICVCTSFLDTRWQLWMMKKYGVAE-- 292
T++++ LP H + A L VL SF + +W+ K
Sbjct: 230 TFQKLPLPFHIFDYNDSRALSAFREEQLSVLHQ------SFDTEKMDIWVTNKIDETTDW 283
Query: 293 SWTKLMSIPHENLLISNISLWPCVEPL--FISENGVVLLM------NTSSSQLILYNLNS 344
SW+K ++ N L IS+ PL FI E ++L NT S +++ +
Sbjct: 284 SWSKFFTVRLINRLDYPISMM-MKSPLSFFIDEKKNIILCYDKHRENTYKSLVLIVGKDK 342
Query: 345 --RGLYFP 350
R YFP
Sbjct: 343 VYREFYFP 350
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.138 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,297,203
Number of Sequences: 26719
Number of extensions: 397088
Number of successful extensions: 1399
Number of sequences better than 10.0: 288
Number of HSP's better than 10.0 without gapping: 252
Number of HSP's successfully gapped in prelim test: 36
Number of HSP's that attempted gapping in prelim test: 1012
Number of HSP's gapped (non-prelim): 336
length of query: 380
length of database: 11,318,596
effective HSP length: 101
effective length of query: 279
effective length of database: 8,619,977
effective search space: 2404973583
effective search space used: 2404973583
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147013.3


