

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147013.10 - phase: 0 /pseudo
(339 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
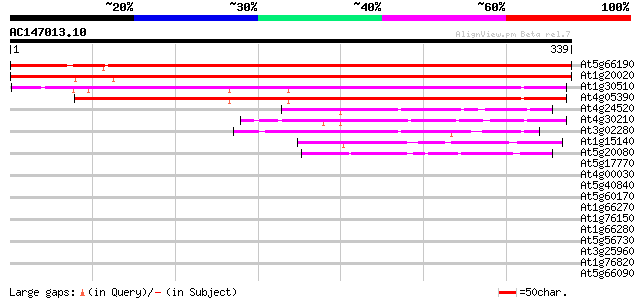
Score E
Sequences producing significant alignments: (bits) Value
At5g66190 ferredoxin-NADP+ reductase 573 e-164
At1g20020 ferredoxin--NADP reductase precursor, putative 543 e-155
At1g30510 ferrodoxin NADP oxidoreductase - like protein 274 4e-74
At4g05390 ferredoxin-NADP+ reductase like protein 272 2e-73
At4g24520 NADPH-ferrihemoprotein reductase ATR1 72 6e-13
At4g30210 NADPH-ferrihemoprotein reductase (ATR2) 67 1e-11
At3g02280 NADPH-ferrihemoprotein reductase like protein 57 1e-08
At1g15140 Unknown protein 52 4e-07
At5g20080 cytochrome-b5 reductase - like protein 42 6e-04
At5g17770 NADH-cytochrome b5 reductase 34 0.100
At4g00030 unknown protein 32 0.50
At5g40840 RAD21-1 variant 2 31 0.85
At5g60170 putative protein 31 1.1
At1g66270 beta-glucosidase 31 1.1
At1g76150 unknown protein 30 1.9
At1g66280 beta-glucosidase, putative 30 2.5
At5g56730 zinc protease PQQL-like protein 29 4.2
At3g25960 putative pyruvate kinase 29 4.2
At1g76820 putative translation initiation factor IF-2 29 4.2
At5g66090 unknown protein 28 5.5
>At5g66190 ferredoxin-NADP+ reductase
Length = 360
Score = 573 bits (1477), Expect = e-164
Identities = 284/342 (83%), Positives = 309/342 (90%), Gaps = 7/342 (2%)
Query: 1 MAAAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVSLNNVSISGRLTVRAEVAT---EA 57
MAAA++AAVS P S S+SL + S +SP R+ KK + V ++V+A+V T EA
Sbjct: 1 MAAAISAAVSLPSSKSSSLLTKISSVSPQRIFLKK---STVCYRRVVSVKAQVTTDTTEA 57
Query: 58 PAPVKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMVFSTEGEVPY 117
P PVKV K SKKQEEGIVVNKFKPK PY GRCLLNTKITGDDAPGETWH+VF+TEGEVPY
Sbjct: 58 P-PVKVVKESKKQEEGIVVNKFKPKNPYTGRCLLNTKITGDDAPGETWHIVFTTEGEVPY 116
Query: 118 REGQSIGIVPDGIDKNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDAGEVVKG 177
REGQSIG++P+GIDKNGKPHKLRLYSIASSA+GDFGDSKTVSLCVKRLVYTND GE+VKG
Sbjct: 117 REGQSIGVIPEGIDKNGKPHKLRLYSIASSAIGDFGDSKTVSLCVKRLVYTNDGGEIVKG 176
Query: 178 VCSNFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHE 237
VCSNFLCDL+PG E +ITGPVGKEMLMPKDPNAT+IMLGTGTGIAPFRSFLWKMFFE+HE
Sbjct: 177 VCSNFLCDLKPGDEAKITGPVGKEMLMPKDPNATIIMLGTGTGIAPFRSFLWKMFFEEHE 236
Query: 238 DYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTR 297
DYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEK P+NFRLDFAVSREQ N+KGEKMYIQTR
Sbjct: 237 DYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKNPDNFRLDFAVSREQTNEKGEKMYIQTR 296
Query: 298 MAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD 339
MA+YAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD
Sbjct: 297 MAEYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD 338
>At1g20020 ferredoxin--NADP reductase precursor, putative
Length = 369
Score = 543 bits (1398), Expect = e-155
Identities = 262/347 (75%), Positives = 300/347 (85%), Gaps = 8/347 (2%)
Query: 1 MAAAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVSL----NNVSISGRL-TVRAEVAT 55
MA + AAVS SNS+S P + I+P+R+ F K + NNV R+ +++A++ T
Sbjct: 1 MATTMNAAVSLTSSNSSSFPATSCAIAPERIRFTKGAFYYKSNNVVTGKRVFSIKAQITT 60
Query: 56 EAPAPV---KVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMVFSTE 112
E P KVEK+SKK EEG++VN+++PK PY G+CLLNTKIT DDAPGETWHMVFS +
Sbjct: 61 ETDTPTPAKKVEKVSKKNEEGVIVNRYRPKEPYTGKCLLNTKITADDAPGETWHMVFSHQ 120
Query: 113 GEVPYREGQSIGIVPDGIDKNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDAG 172
GE+PYREGQS+G++ DGIDKNGKPHK+RLYSIASSALGD G+S+TVSLCVKRLVYTND G
Sbjct: 121 GEIPYREGQSVGVIADGIDKNGKPHKVRLYSIASSALGDLGNSETVSLCVKRLVYTNDQG 180
Query: 173 EVVKGVCSNFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMF 232
E VKGVCSNFLCDL PGS+V++TGPVGKEMLMPKDPNATVIML TGTGIAPFRSFLWKMF
Sbjct: 181 ETVKGVCSNFLCDLAPGSDVKLTGPVGKEMLMPKDPNATVIMLATGTGIAPFRSFLWKMF 240
Query: 233 FEKHEDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKM 292
FEKH+DYKFNGLAWLFLGVPT+SSLLY+EEF+KMK KAPENFR+D+A+SREQ NDKGEKM
Sbjct: 241 FEKHDDYKFNGLAWLFLGVPTTSSLLYQEEFDKMKAKAPENFRVDYAISREQANDKGEKM 300
Query: 293 YIQTRMAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD 339
YIQTRMAQYA ELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAA D
Sbjct: 301 YIQTRMAQYAAELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAND 347
>At1g30510 ferrodoxin NADP oxidoreductase - like protein
Length = 381
Score = 274 bits (701), Expect = 4e-74
Identities = 157/355 (44%), Positives = 211/355 (59%), Gaps = 23/355 (6%)
Query: 2 AAAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVS------LNNVSISGR-------LT 48
A + AVS N SL R SV + + F S LN + S R T
Sbjct: 5 AVSQAGAVSVSIENQRSL--RRSVFKNNSISFNSKSWSSSLALNQKTTSIRDGKRYPSTT 62
Query: 49 VRAEVATEAPAPVKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMV 108
+ V + + V V I + + +N +KPK Y + + ++ G APGET H+V
Sbjct: 63 ICMSVQQTSSSKVTVSPIELEDPKDPPLNLYKPKESYTAKIVSVERVVGPKAPGETCHIV 122
Query: 109 FSTEGEVPYREGQSIGIVPDGID--KNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLV 166
+G +PY EGQS G++P G + K G PH +RLYSIAS+ GDF D KT SLCV+R V
Sbjct: 123 IDHDGNLPYWEGQSYGVIPPGENPKKPGAPHNVRLYSIASTRYGDFFDGKTASLCVRRAV 182
Query: 167 Y----TNDAGEVVKGVCSNFLCDLRPGSEVQITGPVGKEMLMPK-DPNATVIMLGTGTGI 221
Y T GVCSNFLCD +PG ++QITGP GK ML+P+ DPNAT IM+ TGTG+
Sbjct: 183 YYDPETGKEDPSKNGVCSNFLCDSKPGDKIQITGPSGKVMLLPESDPNATHIMIATGTGV 242
Query: 222 APFRSFLWKMFFEKHEDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVS 281
AP+R +L +MF E + F+GLAWLFLGV + SLLY EEF K + P+NFR D A+S
Sbjct: 243 APYRGYLRRMFMENVPNKTFSGLAWLFLGVANTDSLLYDEEFTKYLKDHPDNFRFDKALS 302
Query: 282 REQVNDKGEKMYIQTRMAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLA 336
RE+ N KG KMY+Q ++ +Y++E+++LL + +Y CGLKGM GI D + +A
Sbjct: 303 REEKNKKGGKMYVQDKIEEYSDEIFKLL-DNGAHIYFCGLKGMMPGIQDTLKRVA 356
>At4g05390 ferredoxin-NADP+ reductase like protein
Length = 378
Score = 272 bits (695), Expect = 2e-73
Identities = 141/304 (46%), Positives = 195/304 (63%), Gaps = 8/304 (2%)
Query: 40 NVSISGRLTVRAEVATEAPAPVKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDD 99
++ + R T+ + + + V V + + + +N F+PK PY + +I G
Sbjct: 51 SLGVKKRSTICMSLQQSSKSKVLVTPLELEDPKETPLNLFRPKEPYTATIVSVERIVGPQ 110
Query: 100 APGETWHMVFSTEGEVPYREGQSIGIVPDGID--KNGKPHKLRLYSIASSALGDFGDSKT 157
APGET H+V +G VPY EGQS G++P G + K G PH +RLYSIAS+ GD D KT
Sbjct: 111 APGETCHIVIDHDGNVPYWEGQSYGVIPPGENPKKPGAPHNVRLYSIASTRYGDSFDGKT 170
Query: 158 VSLCVKRLVY----TNDAGEVVKGVCSNFLCDLRPGSEVQITGPVGKEMLMPKD-PNATV 212
SLCV+R +Y T GVCSNFLC+ +PG +V+ITGP GK ML+P+D P AT
Sbjct: 171 ASLCVRRAIYYDPETGKEDPSKAGVCSNFLCNAKPGDKVKITGPSGKVMLLPEDDPKATH 230
Query: 213 IMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPE 272
IM+ TGTG+AP+R +L +MF E ++KF+GLAWLFLGV S SLLY EEF ++ PE
Sbjct: 231 IMIATGTGVAPYRGYLRRMFMENVPNFKFDGLAWLFLGVANSDSLLYDEEFAGYRKDYPE 290
Query: 273 NFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIM 332
NFR D A+SRE+ N KG KMY+Q ++ +Y++E+++LL + +Y CGLKGM GI D +
Sbjct: 291 NFRYDKALSREEKNKKGGKMYVQDKIEEYSDEIFKLL-DNGAHIYFCGLKGMMPGIQDTL 349
Query: 333 VSLA 336
+A
Sbjct: 350 KRVA 353
>At4g24520 NADPH-ferrihemoprotein reductase ATR1
Length = 692
Score = 71.6 bits (174), Expect = 6e-13
Identities = 53/172 (30%), Positives = 92/172 (52%), Gaps = 15/172 (8%)
Query: 165 LVY-TNDAGEVVKGVCSNFLCDLRPGSEV-QITG-PV---GKEMLMPKDPNATVIMLGTG 218
LVY G + KGVCS ++ + P + + +G P+ +P +P+ ++M+G G
Sbjct: 491 LVYGPTPTGRIHKGVCSTWMKNAVPAEKSHECSGAPIFIRASNFKLPSNPSTPIVMVGPG 550
Query: 219 TGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPT-SSSLLYKEEFEKMKEKAPENFRLD 277
TG+APFR FL + K ED + G + LF G +Y++E ++ + L
Sbjct: 551 TGLAPFRGFLQERMALK-EDGEELGSSLLFFGCRNRQMDFIYEDELNNFVDQGVIS-ELI 608
Query: 278 FAVSREQVNDKGEKMYIQTRMAQYAEELWELLKKDNTFVYMCG-LKGMEKGI 328
A SRE +K Y+Q +M + A ++W+L+K++ ++Y+CG KGM + +
Sbjct: 609 MAFSRE----GAQKEYVQHKMMEKAAQVWDLIKEEG-YLYVCGDAKGMARDV 655
>At4g30210 NADPH-ferrihemoprotein reductase (ATR2)
Length = 711
Score = 67.4 bits (163), Expect = 1e-11
Identities = 60/205 (29%), Positives = 96/205 (46%), Gaps = 19/205 (9%)
Query: 140 RLYSIASSALGDFGDSKTVSLCVKRLVYTN-DAGEVVKGVCSNFLCDLRP--GSEVQITG 196
R YSI+SS +++ C LVY G + KGVCS ++ + P SE +
Sbjct: 489 RFYSISSSP--KIAETRIHVTCA--LVYEKMPTGRIHKGVCSTWMKNAVPYEKSENCSSA 544
Query: 197 PV---GKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPT 253
P+ +P D +IM+G GTG+APFR FL + + G + LF G
Sbjct: 545 PIFVRQSNFKLPSDSKVPIIMIGPGTGLAPFRGFLQERLALVESGVEL-GPSVLFFGCRN 603
Query: 254 -SSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLKKD 312
+Y+EE ++ E L A SRE K Y+Q +M A ++W ++ +
Sbjct: 604 RRMDFIYEEELQRFVESG-ALAELSVAFSREGPT----KEYVQHKMMDKASDIWNMISQ- 657
Query: 313 NTFVYMCG-LKGMEKGIDDIMVSLA 336
++Y+CG KGM + + + ++A
Sbjct: 658 GAYLYVCGDAKGMARDVHRSLHTIA 682
>At3g02280 NADPH-ferrihemoprotein reductase like protein
Length = 623
Score = 57.4 bits (137), Expect = 1e-08
Identities = 51/192 (26%), Positives = 86/192 (44%), Gaps = 18/192 (9%)
Query: 136 PHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDAGEVVKGVCSNFLCDLRPGSEVQIT 195
P K R +SI+SS L V L V + + KG+CS++L L P EV I
Sbjct: 395 PLKPRAFSISSSPLAH---PAAVHLTVSIVSWITPYKRTRKGLCSSWLASLAPEQEVNIP 451
Query: 196 GPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPTSS 255
K L + +I++G GTG APFR F+ + + + + + F +
Sbjct: 452 VWFHKGSLPAPSQSLPLILVGPGTGCAPFRGFIAERAVQA-QSSPVAPVMFFFGCRNKDT 510
Query: 256 SLLYKEEFEK-------MKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWEL 308
LY++ +E + E F F+ D+ +K+Y+Q ++ + ++ +W+L
Sbjct: 511 DFLYRDFWESHAREGGMLSEGKGGGFYTAFS------RDQPKKVYVQHKIREMSKRVWDL 564
Query: 309 LKKDNTFVYMCG 320
L D VY+ G
Sbjct: 565 L-CDGAAVYVAG 575
>At1g15140 Unknown protein
Length = 295
Score = 52.4 bits (124), Expect = 4e-07
Identities = 41/165 (24%), Positives = 77/165 (45%), Gaps = 18/165 (10%)
Query: 175 VKGVCSNFLCDLRPGSEVQITGPVGK----EMLMPKDPNATVIMLGTGTGIAPFRSFLWK 230
+ G + LC L+ G V+++ +G +++ P + TV++ TG+GI+P RS +
Sbjct: 132 IAGSTAEILCGLKKGETVELSSVMGNGFNIDLIDPPEEYPTVLIFATGSGISPIRSLIES 191
Query: 231 MF-FEKHEDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKG 289
F ++ D + L+ G + + Y+E+F KE ++ +S+ KG
Sbjct: 192 GFGADRRSDVR------LYYGARNLNRMAYQEKF---KEWESAGVKVVPVLSQPDDGWKG 242
Query: 290 EKMYIQTRMAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVS 334
E Y+Q A+ +L T +CG K M + I ++V+
Sbjct: 243 ETGYVQAAFARAK----QLSAPKATGAVLCGQKQMAEEITSMLVA 283
>At5g20080 cytochrome-b5 reductase - like protein
Length = 328
Score = 41.6 bits (96), Expect = 6e-04
Identities = 37/152 (24%), Positives = 67/152 (43%), Gaps = 10/152 (6%)
Query: 177 GVCSNFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKH 236
G S L+PG +++ GPV K P + + M+ G+GI P + +
Sbjct: 157 GKMSQHFASLKPGDVLEVKGPVEKFKYSP-NMKKHIGMIAGGSGITPMLQVIDAIVKNPE 215
Query: 237 EDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQT 296
++ + ++ L+ V + +L K++ + ++ P N ++ + V N KG YI
Sbjct: 216 DNTQ---ISLLYANV-SPDDILLKQKLDVLQANHP-NLKIFYTVDNPTKNWKGGVGYISK 270
Query: 297 RMAQYAEELWELLKKDNTFVYMCGLKGMEKGI 328
MA L D+T + +CG GM + I
Sbjct: 271 DMALKGLP----LPTDDTLILVCGPPGMMEHI 298
>At5g17770 NADH-cytochrome b5 reductase
Length = 281
Score = 34.3 bits (77), Expect = 0.100
Identities = 33/135 (24%), Positives = 57/135 (41%), Gaps = 10/135 (7%)
Query: 176 KGVCSNFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEK 235
+G S+ ++R G + + GP G+ P A ++ G G+GI P +
Sbjct: 121 QGRMSHHFREMRVGDHLAVKGPKGRFKYQPGQFRAFGMLAG-GSGITPMFQVARAILENP 179
Query: 236 HEDYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSR-EQVNDKG----E 290
+ K + L T +L KEE E + PE F++ + +++ +V D G
Sbjct: 180 TDKTKVH----LIYANVTYDDILLKEELEGLTTNYPEQFKIFYVLNQPPEVWDGGVGFVS 235
Query: 291 KMYIQTRMAQYAEEL 305
K IQT A ++
Sbjct: 236 KEMIQTHCPAPASDI 250
>At4g00030 unknown protein
Length = 212
Score = 32.0 bits (71), Expect = 0.50
Identities = 21/63 (33%), Positives = 36/63 (56%), Gaps = 7/63 (11%)
Query: 52 EVATEAPAPVKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMVFST 111
E+ +E ++ +K KK++E +VN + T IGR + IT DD+ TW ++++T
Sbjct: 42 ELISEEDRGLRTQKDPKKRDE--IVNAIESMT-VIGR----SSITTDDSLSATWRLLWTT 94
Query: 112 EGE 114
E E
Sbjct: 95 EKE 97
>At5g40840 RAD21-1 variant 2
Length = 809
Score = 31.2 bits (69), Expect = 0.85
Identities = 29/125 (23%), Positives = 52/125 (41%), Gaps = 24/125 (19%)
Query: 2 AAAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVSLNNVSISGRLTVRAEVATEAPAPV 61
A++ T+ + N+ P++ SVI+P+ V + S V V +V PV
Sbjct: 608 ASSTTSGTAHQTENAAETPVKPSVIAPETPV--RTSEQTVIAPETPVVSEQVEIAPETPV 665
Query: 62 KVEKISKK--QEEGIVVNKFKPKTPY----------------IGRCLLNTKITGD---DA 100
+ E +SK+ ++ G K +P +P+ + L+N ++ D D
Sbjct: 666 R-ESMSKRFFKDPGTCYKKSRPASPFTSFEEHPSVYYVENRDLDTILMNDEVNADERQDL 724
Query: 101 PGETW 105
ETW
Sbjct: 725 QQETW 729
>At5g60170 putative protein
Length = 944
Score = 30.8 bits (68), Expect = 1.1
Identities = 26/78 (33%), Positives = 35/78 (44%), Gaps = 10/78 (12%)
Query: 14 SNSTSLPIRTSVISPDRLVFKKVS------LNNVSISGRLTVRAEVATEAPAPVKVEKI- 66
S ST+LP S + L S + S++G L A VA A PV I
Sbjct: 256 SRSTALPAAASWGTHQSLATSVTSNGSSDIQRSTSVNGTLPFSAVVANAAHGPVSSNDIL 315
Query: 67 ---SKKQEEGIVVNKFKP 81
S+K+E IV++K KP
Sbjct: 316 KRPSRKEESQIVMDKVKP 333
>At1g66270 beta-glucosidase
Length = 524
Score = 30.8 bits (68), Expect = 1.1
Identities = 20/52 (38%), Positives = 27/52 (51%)
Query: 259 YKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLK 310
YKE+ + MK + FRL A SR + + EK Q + Y E + ELLK
Sbjct: 96 YKEDIQLMKNLNTDAFRLSIAWSRIFPHGRKEKGVSQAGVQFYHELIDELLK 147
>At1g76150 unknown protein
Length = 309
Score = 30.0 bits (66), Expect = 1.9
Identities = 19/69 (27%), Positives = 32/69 (45%)
Query: 255 SSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLKKDNT 314
S LL+ +++ ++ P L VS + DKG+ ++ Y E ELL + T
Sbjct: 91 SLLLHGQQYIEIYRPLPSKASLINKVSLAGLQDKGKAAILELETRSYEEGSGELLCMNRT 150
Query: 315 FVYMCGLKG 323
V++ G G
Sbjct: 151 TVFLRGAGG 159
>At1g66280 beta-glucosidase, putative
Length = 524
Score = 29.6 bits (65), Expect = 2.5
Identities = 19/52 (36%), Positives = 27/52 (51%)
Query: 259 YKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLK 310
YKE+ + MK + FRL A SR + + EK Q + Y + + ELLK
Sbjct: 96 YKEDIQLMKNLNTDAFRLSIAWSRIFPHGRKEKGVSQAGVKFYHDLIDELLK 147
>At5g56730 zinc protease PQQL-like protein
Length = 956
Score = 28.9 bits (63), Expect = 4.2
Identities = 32/127 (25%), Positives = 53/127 (41%), Gaps = 24/127 (18%)
Query: 43 ISGRLTVRAEVATEAPAPVKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPG 102
+S +++ AE ++E ++V K ++E G V+ +++ GR
Sbjct: 141 LSQAISILAEFSSE----IRVSKEDLEKERGAVMEEYRGNRNATGRM------------- 183
Query: 103 ETWHMVFSTEGEVPYREGQSIGI------VPDGIDKNGKPHKLRLYSIASSALGDFGDSK 156
+ H EG Y E IG+ VP K L ++A A+GDF D+K
Sbjct: 184 QDSHWQLMMEGS-KYAERLPIGLEKVIRSVPAATVKQFYQKWYHLCNMAVVAVGDFPDTK 242
Query: 157 TVSLCVK 163
TV +K
Sbjct: 243 TVVDLIK 249
>At3g25960 putative pyruvate kinase
Length = 497
Score = 28.9 bits (63), Expect = 4.2
Identities = 37/135 (27%), Positives = 58/135 (42%), Gaps = 20/135 (14%)
Query: 53 VATEAPAPVKVEKISKKQEEGIVVNKFK---PKTPYIGRCLLNTKITGDDAPGETWHMVF 109
V T PA VE I K + G+ V +F Y L N + T D G ++
Sbjct: 21 VCTLGPASRSVEMIEKLLKAGMNVARFNFSHGSHSYHQETLDNLR-TAMDNTGILCAVML 79
Query: 110 STEGEVPYREGQSIGIVPDGIDKNGKPHKLRLYSIASSALGDF---GDSKTVSLCVKRLV 166
T+ V + G K GKP +L+ + ++ D+ GDS T+S+ K+L
Sbjct: 80 DTKSPV----------IRTGFLKEGKPIQLKQGQEITISI-DYKIQGDSNTISMSYKKLA 128
Query: 167 YTNDAGEVVKGVCSN 181
G+V+ +CS+
Sbjct: 129 EDLKPGDVI--LCSD 141
>At1g76820 putative translation initiation factor IF-2
Length = 1146
Score = 28.9 bits (63), Expect = 4.2
Identities = 19/61 (31%), Positives = 32/61 (52%), Gaps = 4/61 (6%)
Query: 261 EEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLKKDNTFVYMCG 320
++ E MKE A E+ ++ +SR ++ GE +Y+QT E L E LK + + G
Sbjct: 895 DDIEAMKESAMED--MESVLSR--IDKSGEGVYVQTSTLGSLEALLEFLKTPAVNIPVSG 950
Query: 321 L 321
+
Sbjct: 951 I 951
>At5g66090 unknown protein
Length = 210
Score = 28.5 bits (62), Expect = 5.5
Identities = 20/62 (32%), Positives = 37/62 (59%), Gaps = 2/62 (3%)
Query: 2 AAAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVSLNNVSISGRLTVRAEVATEAPAPV 61
AA+ A + S+S++L +S+ S R + +SL +V++ GR+ +A+ A +P+PV
Sbjct: 15 AASKQATLPSSSSSSSALYPLSSLRSKPRQ--QLLSLRSVALKGRVKAQAKEAEASPSPV 72
Query: 62 KV 63
V
Sbjct: 73 GV 74
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.135 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,820,479
Number of Sequences: 26719
Number of extensions: 355509
Number of successful extensions: 922
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 895
Number of HSP's gapped (non-prelim): 26
length of query: 339
length of database: 11,318,596
effective HSP length: 100
effective length of query: 239
effective length of database: 8,646,696
effective search space: 2066560344
effective search space used: 2066560344
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147013.10


