

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147011.7 + phase: 0 /pseudo
(384 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
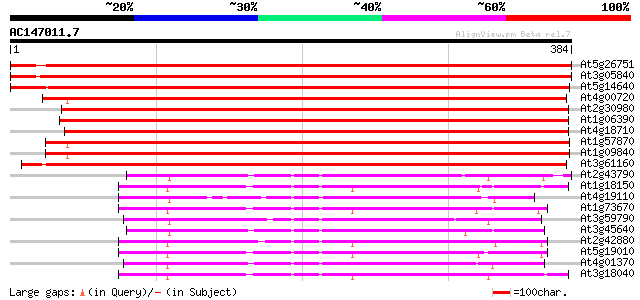
Score E
Sequences producing significant alignments: (bits) Value
At5g26751 shaggy-like kinase alpha 732 0.0
At3g05840 shaggy related protein kinase, ASK-GAMMA 726 0.0
At5g14640 protein kinase MSK-3 - like 679 0.0
At4g00720 Shaggy related protein kinase tetha 623 e-179
At2g30980 putative shaggy-like protein kinase dzeta 605 e-173
At1g06390 putative shaggy kinase iota protein 604 e-173
At4g18710 shaggy-like protein kinase etha (EC 2.7.1.-) 601 e-172
At1g57870 protein kinase, putative 596 e-171
At1g09840 shaggy-like protien kinase, kappa 584 e-167
At3g61160 shaggy-like kinase beta 554 e-158
At2g43790 MAP kinase (ATMPK6) 190 1e-48
At1g18150 unknown protein 183 2e-46
At4g19110 putative serine/threonine-protein kinase 180 1e-45
At1g73670 MAP kinase like protein 180 1e-45
At3g59790 mitogen-activated protein kinase-like protein 179 3e-45
At3g45640 mitogen-activated protein kinase 3 178 4e-45
At2g42880 MAP kinase 4 homolog 176 2e-44
At5g19010 MAP kinase -like protein 175 3e-44
At4g01370 MAP kinase 4 175 4e-44
At3g18040 MAP kinase like protein 175 4e-44
>At5g26751 shaggy-like kinase alpha
Length = 405
Score = 732 bits (1889), Expect = 0.0
Identities = 357/384 (92%), Positives = 365/384 (94%), Gaps = 6/384 (1%)
Query: 1 MASVGVAPTSGFKESLGDGEIGVDDILPEEMSDMKIRDDREMEATVVDGNGTETGHIIVT 60
MASVG+AP G ++S G D LPEEM+DMKIRDD+EMEATVVDGNGTETGHIIVT
Sbjct: 1 MASVGIAPNPGARDSTGV------DKLPEEMNDMKIRDDKEMEATVVDGNGTETGHIIVT 54
Query: 61 TIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKCLETGETVAIKKVLQDKRYKNRELQTM 120
TIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKCLETGETVAIKKVLQD+RYKNRELQTM
Sbjct: 55 TIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKCLETGETVAIKKVLQDRRYKNRELQTM 114
Query: 121 RLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYT 180
RLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHY+KLNQRMP+IYVKLYT
Sbjct: 115 RLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYNKLNQRMPLIYVKLYT 174
Query: 181 YQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRY 240
YQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRY
Sbjct: 175 YQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRY 234
Query: 241 YRAPELIFGATEYTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEI 300
YRAPELIFGATEYTTAIDVWS GCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEI
Sbjct: 235 YRAPELIFGATEYTTAIDVWSAGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEI 294
Query: 301 KCMNPNYTEFKFPQIKAHPWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFD 360
KCMNPNYTEFKFPQIKAHPWHKIFHKRMP EAVDLVSRLLQYSPNLR ALD L HPFFD
Sbjct: 295 KCMNPNYTEFKFPQIKAHPWHKIFHKRMPPEAVDLVSRLLQYSPNLRSAALDTLVHPFFD 354
Query: 361 ELRDPNARLPTGRFLPPLFNFKPH 384
ELRDPNARLP GRFLPPLFNFKPH
Sbjct: 355 ELRDPNARLPNGRFLPPLFNFKPH 378
>At3g05840 shaggy related protein kinase, ASK-GAMMA
Length = 409
Score = 726 bits (1875), Expect = 0.0
Identities = 351/384 (91%), Positives = 366/384 (94%), Gaps = 2/384 (0%)
Query: 1 MASVGVAPTSGFKESLGDGEIGVDDILPEEMSDMKIRDDREMEATVVDGNGTETGHIIVT 60
MASVG+ P++ +ES G+ + D LPEEM DMKI+DD+EMEAT+V+GN TETGHIIVT
Sbjct: 1 MASVGIEPSAAVRESTGN--VTDADRLPEEMKDMKIQDDKEMEATIVNGNVTETGHIIVT 58
Query: 61 TIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKCLETGETVAIKKVLQDKRYKNRELQTM 120
TIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKCLETGETVAIKKVLQD+RYKNRELQTM
Sbjct: 59 TIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKCLETGETVAIKKVLQDRRYKNRELQTM 118
Query: 121 RLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYT 180
RLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHY+KLNQRMP++YVKLYT
Sbjct: 119 RLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYNKLNQRMPLVYVKLYT 178
Query: 181 YQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRY 240
YQIFR+LSYIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRY
Sbjct: 179 YQIFRSLSYIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRY 238
Query: 241 YRAPELIFGATEYTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEI 300
YRAPELIFGATEYTTAIDVWS GCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEI
Sbjct: 239 YRAPELIFGATEYTTAIDVWSAGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEI 298
Query: 301 KCMNPNYTEFKFPQIKAHPWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFD 360
KCMNPNYTEFKFPQIKAHPWHKIFHKRMP EAVDLVSRLLQYSPNLRC ALD L HPFFD
Sbjct: 299 KCMNPNYTEFKFPQIKAHPWHKIFHKRMPPEAVDLVSRLLQYSPNLRCAALDSLVHPFFD 358
Query: 361 ELRDPNARLPTGRFLPPLFNFKPH 384
ELRDPNARLP GRFLPPLFNFKPH
Sbjct: 359 ELRDPNARLPNGRFLPPLFNFKPH 382
>At5g14640 protein kinase MSK-3 - like
Length = 410
Score = 679 bits (1753), Expect = 0.0
Identities = 329/383 (85%), Positives = 348/383 (89%), Gaps = 1/383 (0%)
Query: 1 MASVGVAPTSGFKESLGDGEIGVDDILPEEMSDMKIRDDREMEATVVDGNGTETGHIIVT 60
MASVG P S + I + LPE +++MKI+DD+EMEA VVDGNGTETGHIIVT
Sbjct: 1 MASVGTLPASSMATKQSNASICAEK-LPEGINEMKIKDDKEMEAAVVDGNGTETGHIIVT 59
Query: 61 TIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKCLETGETVAIKKVLQDKRYKNRELQTM 120
TIGG+NGQPKQTISYMAER+VG GSFG+VFQAKCLETGETVAIKKVLQDKRYKNRELQTM
Sbjct: 60 TIGGKNGQPKQTISYMAERIVGQGSFGIVFQAKCLETGETVAIKKVLQDKRYKNRELQTM 119
Query: 121 RLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYT 180
RLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETV+RV KHYS+ NQRMP+IYVKLYT
Sbjct: 120 RLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETVYRVSKHYSRANQRMPIIYVKLYT 179
Query: 181 YQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRY 240
YQI RAL+YIH +GVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRY
Sbjct: 180 YQICRALAYIHGGVGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRY 239
Query: 241 YRAPELIFGATEYTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEI 300
YRAPELIFGATEYTT ID+WS GCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEI
Sbjct: 240 YRAPELIFGATEYTTTIDIWSAGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEI 299
Query: 301 KCMNPNYTEFKFPQIKAHPWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFD 360
KCMNPNYTEFKFPQIKAHPWHKIFHKR P EAVDLVSRLLQYSPNLR A++ + HPFFD
Sbjct: 300 KCMNPNYTEFKFPQIKAHPWHKIFHKRTPPEAVDLVSRLLQYSPNLRSTAMEAIVHPFFD 359
Query: 361 ELRDPNARLPTGRFLPPLFNFKP 383
ELRDPN RLP GR LPPLFNFKP
Sbjct: 360 ELRDPNTRLPNGRALPPLFNFKP 382
>At4g00720 Shaggy related protein kinase tetha
Length = 472
Score = 623 bits (1607), Expect = e-179
Identities = 297/366 (81%), Positives = 329/366 (89%), Gaps = 7/366 (1%)
Query: 23 VDDILPEEMSDMKIRD-------DREMEATVVDGNGTETGHIIVTTIGGRNGQPKQTISY 75
VDD LP+ M +MKIRD D++ME TVV+G+GTETG +I TT+GGR+G+PKQTISY
Sbjct: 79 VDDQLPDVMIEMKIRDERNANREDKDMETTVVNGSGTETGQVITTTVGGRDGKPKQTISY 138
Query: 76 MAERVVGHGSFGVVFQAKCLETGETVAIKKVLQDKRYKNRELQTMRLLDHPNVVSLKHCF 135
MA+RVVG GSFGVVFQAKCLETGE VAIKKVLQDKRYKNRELQ MRL DHPNVV L+H F
Sbjct: 139 MAQRVVGTGSFGVVFQAKCLETGEQVAIKKVLQDKRYKNRELQIMRLQDHPNVVRLRHSF 198
Query: 136 FSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRCIG 195
FSTT+KDELYLNLVLEYVPETV+R KHY+K+NQ MP+I+V+LYTYQI RAL+Y+HR +G
Sbjct: 199 FSTTDKDELYLNLVLEYVPETVYRASKHYTKMNQHMPIIFVQLYTYQICRALNYLHRVVG 258
Query: 196 VCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTT 255
VCHRDIKPQNLLVNP THQ+K+CDFGSAK+LV GEPNISYICSRYYRAPELIFGATEYT
Sbjct: 259 VCHRDIKPQNLLVNPQTHQLKICDFGSAKMLVPGEPNISYICSRYYRAPELIFGATEYTN 318
Query: 256 AIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQI 315
AID+WS GCV+AELLLGQPLFPGESG+DQLVEIIK+LGTPTREEI+CMNPNYTEFKFPQI
Sbjct: 319 AIDMWSGGCVMAELLLGQPLFPGESGIDQLVEIIKILGTPTREEIRCMNPNYTEFKFPQI 378
Query: 316 KAHPWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRDPNARLPTGRFL 375
KAHPWHKIFHKRMP EAVDLVSRLLQYSPNLRC AL+ HPFFD+LRDPN LP GR L
Sbjct: 379 KAHPWHKIFHKRMPPEAVDLVSRLLQYSPNLRCTALEACAHPFFDDLRDPNVSLPNGRAL 438
Query: 376 PPLFNF 381
PPLFNF
Sbjct: 439 PPLFNF 444
>At2g30980 putative shaggy-like protein kinase dzeta
Length = 412
Score = 605 bits (1559), Expect = e-173
Identities = 283/347 (81%), Positives = 317/347 (90%)
Query: 36 IRDDREMEATVVDGNGTETGHIIVTTIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKCL 95
I +D+EM A V++GN TGHII TTIGG+NG+PKQTISYMAERVVG GSFG+VFQAKCL
Sbjct: 33 IDNDKEMSAAVIEGNDAVTGHIISTTIGGKNGEPKQTISYMAERVVGTGSFGIVFQAKCL 92
Query: 96 ETGETVAIKKVLQDKRYKNRELQTMRLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPE 155
ETGE+VAIKKVLQD+RYKNRELQ MRL+DHPNVVSLKHCFFSTT +DEL+LNLV+EYVPE
Sbjct: 93 ETGESVAIKKVLQDRRYKNRELQLMRLMDHPNVVSLKHCFFSTTTRDELFLNLVMEYVPE 152
Query: 156 TVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQV 215
T++RV+KHY+ NQRMP+ YVKLYTYQIFR L+YIH GVCHRD+KPQNLLV+P THQ
Sbjct: 153 TLYRVLKHYTSSNQRMPIFYVKLYTYQIFRGLAYIHTAPGVCHRDVKPQNLLVDPLTHQC 212
Query: 216 KLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTTAIDVWSVGCVLAELLLGQPL 275
KLCDFGSAKVLVKGE NISYICSRYYRAPELIFGATEYT++ID+WS GCVLAELLLGQPL
Sbjct: 213 KLCDFGSAKVLVKGEANISYICSRYYRAPELIFGATEYTSSIDIWSAGCVLAELLLGQPL 272
Query: 276 FPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFHKRMPAEAVDL 335
FPGE+ VDQLVEIIKVLGTPTREEI+CMNPNYT+F+FPQIKAHPWHK+FHKRMP EA+DL
Sbjct: 273 FPGENSVDQLVEIIKVLGTPTREEIRCMNPNYTDFRFPQIKAHPWHKVFHKRMPPEAIDL 332
Query: 336 VSRLLQYSPNLRCQALDCLTHPFFDELRDPNARLPTGRFLPPLFNFK 382
SRLLQYSP+LRC AL+ HPFF+ELR+PNARLP GR LPPLFNFK
Sbjct: 333 ASRLLQYSPSLRCTALEACAHPFFNELREPNARLPNGRPLPPLFNFK 379
>At1g06390 putative shaggy kinase iota protein
Length = 407
Score = 604 bits (1558), Expect = e-173
Identities = 281/348 (80%), Positives = 317/348 (90%)
Query: 35 KIRDDREMEATVVDGNGTETGHIIVTTIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKC 94
++ D+EM A V++GN TGHII TTIGG+NG+PKQTISYMAERVVG GSFG+VFQAKC
Sbjct: 30 ELDSDKEMSAAVIEGNDAVTGHIISTTIGGKNGEPKQTISYMAERVVGTGSFGIVFQAKC 89
Query: 95 LETGETVAIKKVLQDKRYKNRELQTMRLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVP 154
LETGE+VAIKKVLQD+RYKNRELQ MR +DHPNV+SLKHCFFSTT +DEL+LNLV+EYVP
Sbjct: 90 LETGESVAIKKVLQDRRYKNRELQLMRPMDHPNVISLKHCFFSTTSRDELFLNLVMEYVP 149
Query: 155 ETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQ 214
ET++RV++HY+ NQRMP+ YVKLYTYQIFR L+YIH GVCHRD+KPQNLLV+P THQ
Sbjct: 150 ETLYRVLRHYTSSNQRMPIFYVKLYTYQIFRGLAYIHTVPGVCHRDVKPQNLLVDPLTHQ 209
Query: 215 VKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTTAIDVWSVGCVLAELLLGQP 274
VKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYT +ID+WS GCVLAELLLGQP
Sbjct: 210 VKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTASIDIWSAGCVLAELLLGQP 269
Query: 275 LFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFHKRMPAEAVD 334
LFPGE+ VDQLVEIIKVLGTPTREEI+CMNPNYT+F+FPQIKAHPWHK+FHKRMP EA+D
Sbjct: 270 LFPGENSVDQLVEIIKVLGTPTREEIRCMNPNYTDFRFPQIKAHPWHKVFHKRMPPEAID 329
Query: 335 LVSRLLQYSPNLRCQALDCLTHPFFDELRDPNARLPTGRFLPPLFNFK 382
L SRLLQYSP+LRC AL+ HPFF+ELR+PNARLP GR LPPLFNFK
Sbjct: 330 LASRLLQYSPSLRCTALEACAHPFFNELREPNARLPNGRPLPPLFNFK 377
>At4g18710 shaggy-like protein kinase etha (EC 2.7.1.-)
Length = 380
Score = 601 bits (1549), Expect = e-172
Identities = 281/345 (81%), Positives = 313/345 (90%)
Query: 38 DDREMEATVVDGNGTETGHIIVTTIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKCLET 97
DD+EM A VVDG+ TGHII TTIGG+NG+PKQTISYMAERVVG GSFG+VFQAKCLET
Sbjct: 3 DDKEMPAAVVDGHDQVTGHIISTTIGGKNGEPKQTISYMAERVVGTGSFGIVFQAKCLET 62
Query: 98 GETVAIKKVLQDKRYKNRELQTMRLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETV 157
GETVAIKKVLQD+RYKNRELQ MR++DHPNVV LKHCFFSTT KDEL+LNLV+EYVPE++
Sbjct: 63 GETVAIKKVLQDRRYKNRELQLMRVMDHPNVVCLKHCFFSTTSKDELFLNLVMEYVPESL 122
Query: 158 HRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQVKL 217
+RV+KHYS NQRMP++YVKLY YQIFR L+YIH GVCHRD+KPQNLLV+P THQVK+
Sbjct: 123 YRVLKHYSSANQRMPLVYVKLYMYQIFRGLAYIHNVAGVCHRDLKPQNLLVDPLTHQVKI 182
Query: 218 CDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTTAIDVWSVGCVLAELLLGQPLFP 277
CDFGSAK LVKGE NISYICSR+YRAPELIFGATEYTT+ID+WS GCVLAELLLGQPLFP
Sbjct: 183 CDFGSAKQLVKGEANISYICSRFYRAPELIFGATEYTTSIDIWSAGCVLAELLLGQPLFP 242
Query: 278 GESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFHKRMPAEAVDLVS 337
GE+ VDQLVEIIKVLGTPTREEI+CMNP+YT+F+FPQIKAHPWHKIFHKRMP EA+D S
Sbjct: 243 GENAVDQLVEIIKVLGTPTREEIRCMNPHYTDFRFPQIKAHPWHKIFHKRMPPEAIDFAS 302
Query: 338 RLLQYSPNLRCQALDCLTHPFFDELRDPNARLPTGRFLPPLFNFK 382
RLLQYSP+LRC AL+ HPFFDELR+PNARLP GR PPLFNFK
Sbjct: 303 RLLQYSPSLRCTALEACAHPFFDELREPNARLPNGRPFPPLFNFK 347
>At1g57870 protein kinase, putative
Length = 420
Score = 596 bits (1536), Expect = e-171
Identities = 281/365 (76%), Positives = 316/365 (85%), Gaps = 6/365 (1%)
Query: 25 DILPEEMSDMKIRD------DREMEATVVDGNGTETGHIIVTTIGGRNGQPKQTISYMAE 78
D L +M +MKIRD +R+ E ++DG G E GH+I TT+ GRNGQ +QT+SY+AE
Sbjct: 26 DWLSRDMLEMKIRDKTEADEERDSEPDIIDGVGAEPGHVITTTLPGRNGQSRQTVSYIAE 85
Query: 79 RVVGHGSFGVVFQAKCLETGETVAIKKVLQDKRYKNRELQTMRLLDHPNVVSLKHCFFST 138
VVG GSFG+VFQAKC ETGE VAIKKVLQDKRYKNRELQ M++LDHPNVV LKH F+S
Sbjct: 86 HVVGTGSFGMVFQAKCRETGEVVAIKKVLQDKRYKNRELQIMQMLDHPNVVCLKHSFYSR 145
Query: 139 TEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRCIGVCH 198
TE +E+YLNLVLE+VPETV+R + YS++NQ MP+IYVKLYTYQI R L+Y+H C G+CH
Sbjct: 146 TENEEVYLNLVLEFVPETVNRTARSYSRMNQLMPLIYVKLYTYQICRGLAYLHNCCGLCH 205
Query: 199 RDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTTAID 258
RDIKPQNLLVNPHTHQ+K+CDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTTAID
Sbjct: 206 RDIKPQNLLVNPHTHQLKICDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTTAID 265
Query: 259 VWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAH 318
+WS GCV+AELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIK H
Sbjct: 266 IWSTGCVMAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKPH 325
Query: 319 PWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRDPNARLPTGRFLPPL 378
PWHK+F KR+P EAVDL+ R QYSPNLRC A++ HPFFDELRDPNARLP GR LPPL
Sbjct: 326 PWHKVFQKRLPPEAVDLLCRFFQYSPNLRCTAVEACIHPFFDELRDPNARLPNGRPLPPL 385
Query: 379 FNFKP 383
FNFKP
Sbjct: 386 FNFKP 390
>At1g09840 shaggy-like protien kinase, kappa
Length = 421
Score = 584 bits (1506), Expect = e-167
Identities = 275/365 (75%), Positives = 314/365 (85%), Gaps = 6/365 (1%)
Query: 25 DILPEEMSDMKIRD------DREMEATVVDGNGTETGHIIVTTIGGRNGQPKQTISYMAE 78
D L ++++ +IRD +R+ E ++DG G E GH+I TT+ GRNGQ +QT+SY++E
Sbjct: 27 DWLTRDLAETRIRDKVETDDERDSEPDIIDGAGAEPGHVIRTTLRGRNGQSRQTVSYISE 86
Query: 79 RVVGHGSFGVVFQAKCLETGETVAIKKVLQDKRYKNRELQTMRLLDHPNVVSLKHCFFST 138
VVG GSFG+VFQAKC ETGE VAIKKVLQDKRYKNRELQ M++LDHPN V+LKH FFS
Sbjct: 87 HVVGTGSFGMVFQAKCRETGEVVAIKKVLQDKRYKNRELQIMQMLDHPNAVALKHSFFSR 146
Query: 139 TEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRCIGVCH 198
T+ +E+YLNLVLE+VPETV+RV + YS+ NQ MP+IYVKLYTYQI RAL+YIH G+CH
Sbjct: 147 TDNEEVYLNLVLEFVPETVNRVARSYSRTNQLMPLIYVKLYTYQICRALAYIHNSFGLCH 206
Query: 199 RDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTTAID 258
RDIKPQNLLVNPHTHQ+K+CDFGSAKVLVKGEPN+SYICSRYYRAPELIFGA+EYTTAID
Sbjct: 207 RDIKPQNLLVNPHTHQLKICDFGSAKVLVKGEPNVSYICSRYYRAPELIFGASEYTTAID 266
Query: 259 VWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAH 318
+WS GCV+AELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIK H
Sbjct: 267 IWSTGCVMAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKPH 326
Query: 319 PWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRDPNARLPTGRFLPPL 378
PWHK+F KR+P EAVDL+ R QYSPNLRC AL+ HP FDELRDPN RLP GR LPPL
Sbjct: 327 PWHKVFQKRLPPEAVDLLCRFFQYSPNLRCTALEACIHPLFDELRDPNTRLPNGRPLPPL 386
Query: 379 FNFKP 383
FNFKP
Sbjct: 387 FNFKP 391
>At3g61160 shaggy-like kinase beta
Length = 438
Score = 554 bits (1427), Expect = e-158
Identities = 265/373 (71%), Positives = 311/373 (83%), Gaps = 2/373 (0%)
Query: 9 TSGFKESLGDGEIGVDDILPEEMSDMKIRDDREMEATVVDGNGTETGHIIVTTIGGRNGQ 68
TS K+S+ E D LP+E+ ++ + DD++M+ ++ GNGTE+G II T G N Q
Sbjct: 45 TSYEKDSVSSSENS--DHLPKEIREVGLGDDKDMDCGIIKGNGTESGRIITTKKKGLNDQ 102
Query: 69 PKQTISYMAERVVGHGSFGVVFQAKCLETGETVAIKKVLQDKRYKNRELQTMRLLDHPNV 128
+TISY AE V+G GSFGVVFQAKCLET E VAIKKVLQDKRYKNRELQ MR+LDHPNV
Sbjct: 103 KDKTISYRAEHVIGTGSFGVVFQAKCLETEEKVAIKKVLQDKRYKNRELQIMRMLDHPNV 162
Query: 129 VSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALS 188
V LKH FFSTTEKDELYLNLVLEYVPET++R + Y+K+NQ MP+IY++LYTYQI RA++
Sbjct: 163 VELKHSFFSTTEKDELYLNLVLEYVPETIYRASRSYTKMNQHMPLIYIQLYTYQICRAMN 222
Query: 189 YIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRYYRAPELIF 248
Y+H+ +GVCHRDIKPQNLLVN TH+VK+CDFGSAK+L+ GEPNISYICSRYYRAPELIF
Sbjct: 223 YLHQVVGVCHRDIKPQNLLVNNVTHEVKICDFGSAKMLIPGEPNISYICSRYYRAPELIF 282
Query: 249 GATEYTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYT 308
GATEYT+AID+WSVGCV+AEL LG PLFPGE+ VDQLVEIIK+LGTP REEIK MNP Y
Sbjct: 283 GATEYTSAIDMWSVGCVMAELFLGHPLFPGETSVDQLVEIIKILGTPAREEIKNMNPRYN 342
Query: 309 EFKFPQIKAHPWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRDPNAR 368
+FKFPQIKA PWHKIF +++ EA+DL SRLLQYSPNLRC AL+ HPFFD+LRDP A
Sbjct: 343 DFKFPQIKAQPWHKIFRRQVSPEAMDLASRLLQYSPNLRCTALEACAHPFFDDLRDPRAS 402
Query: 369 LPTGRFLPPLFNF 381
LP GR LPPLF+F
Sbjct: 403 LPNGRALPPLFDF 415
>At2g43790 MAP kinase (ATMPK6)
Length = 395
Score = 190 bits (482), Expect = 1e-48
Identities = 109/316 (34%), Positives = 176/316 (55%), Gaps = 25/316 (7%)
Query: 81 VGHGSFGVVFQAKCLETGETVAIKKVLQ------DKRYKNRELQTMRLLDHPNVVSLKHC 134
+G G++G+V A ET E+VAIKK+ D + RE++ +R +DH N+V+++
Sbjct: 69 IGKGAYGIVCSAMNSETNESVAIKKIANAFDNKIDAKRTLREIKLLRHMDHENIVAIRDI 128
Query: 135 FFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRCI 194
+ + E + +H++I+ NQ + + + + YQI R L YIH
Sbjct: 129 IPPPLRNAFNDVYIAYELMDTDLHQIIRS----NQALSEEHCQYFLYQILRGLKYIHSA- 183
Query: 195 GVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYT 254
V HRD+KP NLL+N + +K+CDFG A+V + + Y+ +R+YRAPEL+ +++YT
Sbjct: 184 NVLHRDLKPSNLLLNANC-DLKICDFGLARVTSESDFMTEYVVTRWYRAPELLLNSSDYT 242
Query: 255 TAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQ 314
AIDVWSVGC+ EL+ +PLFPG V QL +++++GTP+ EE++ +N N + Q
Sbjct: 243 AAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEEELEFLNENAKRY-IRQ 301
Query: 315 IKAHPWHKIFHK--RMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRD----PNAR 368
+ +P I K + A+DL+ ++L + P R LD L HP+ + L D P
Sbjct: 302 LPPYPRQSITDKFPTVHPLAIDLIEKMLTFDPRRRITVLDALAHPYLNSLHDISDEPECT 361
Query: 369 LPTGRFLPPLFNFKPH 384
+P F+F+ H
Sbjct: 362 IPFN------FDFENH 371
>At1g18150 unknown protein
Length = 589
Score = 183 bits (464), Expect = 2e-46
Identities = 124/324 (38%), Positives = 178/324 (54%), Gaps = 25/324 (7%)
Query: 75 YMAERVVGHGSFGVVFQAKCLETGETVAIKKV------LQDKRYKNRELQTMRLLDHPNV 128
Y + VVG GS+GVV A TGE VAIKK+ + D RE++ +RLL HP+V
Sbjct: 104 YQIQEVVGKGSYGVVASAVDSHTGERVAIKKINDVFEHVSDATRILREIKLLRLLRHPDV 163
Query: 129 VSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALS 188
V +KH + ++ + +V E + +H+VIK N + + + + YQ+ R L
Sbjct: 164 VEIKHIMLPPSRREFRDIYVVFELMESDLHQVIK----ANDDLTPEHYQFFLYQLLRGLK 219
Query: 189 YIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNI----SYICSRYYRAP 244
Y+H V HRD+KP+N+L N ++K+CDFG A+V P Y+ +R+YRAP
Sbjct: 220 YVHAA-NVFHRDLKPKNILANADC-KLKICDFGLARVSFNDAPTAIFWTDYVATRWYRAP 277
Query: 245 ELIFGA-TEYTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEI-KC 302
EL ++YT AID+WSVGC+ AE+LLG+PLFPG++ V QL + LGTP E I +
Sbjct: 278 ELCGSFFSKYTPAIDIWSVGCIFAEMLLGKPLFPGKNVVHQLDLMTDFLGTPPPESISRI 337
Query: 303 MNPNYTEFKFPQIKAHP---WHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFF 359
N + K P HK F K P A+ L+ RLL + P R A D L P+F
Sbjct: 338 RNEKARRYLSSMRKKQPVPFSHK-FPKADPL-ALRLLERLLAFDPKDRASAEDALADPYF 395
Query: 360 DELRDPNARLPTGRFLPPL-FNFK 382
L + + R PT + + L F+F+
Sbjct: 396 SGLSN-SEREPTTQPISKLEFDFE 418
>At4g19110 putative serine/threonine-protein kinase
Length = 461
Score = 180 bits (456), Expect = 1e-45
Identities = 108/295 (36%), Positives = 177/295 (59%), Gaps = 25/295 (8%)
Query: 75 YMAERVVGHGSFGVVFQAKCLETGETVAIKKVLQ-----DKRYKNRELQTMRLLDHPNVV 129
Y + VG G+FG V++A +TGE VAIKK+ + D+ RE++++R ++HPN+V
Sbjct: 4 YKLIKEVGDGTFGSVWRAINKQTGEVVAIKKMKKKYYSWDECINLREVKSLRRMNHPNIV 63
Query: 130 SLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSY 189
LK E D LY V EY+ +++++K KL +K + +Q+F+ LSY
Sbjct: 64 KLKEVI---RENDILYF--VFEYMECNLYQLMKDRQKLFAEAD---IKNWCFQVFQGLSY 115
Query: 190 IHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFG 249
+H+ G HRD+KP+NLLV+ +K+ DFG A+ + P Y+ +R+YRAPE++
Sbjct: 116 MHQR-GYFHRDLKPENLLVSKDI--IKIADFGLAREVNSSPPFTEYVSTRWYRAPEVLLQ 172
Query: 250 ATEYTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREE-IKCMN-PNY 307
+ YT+ +D+W++G ++AELL +P+FPG S D++ +I V+GTPT E ++ +N N
Sbjct: 173 SYVYTSKVDMWAMGAIMAELLSLRPIFPGASEADEIYKICSVIGTPTEETWLEGLNLANT 232
Query: 308 TEFKFPQIKAHPWHKIFHKRMPA---EAVDLVSRLLQYSPNLRCQALDCLTHPFF 359
++FPQ+ P + MP+ +A++L+ RL + P+ R A + L HPFF
Sbjct: 233 INYQFPQLPGVPLSSL----MPSASEDAINLIERLCSWDPSSRPTAAEVLQHPFF 283
>At1g73670 MAP kinase like protein
Length = 576
Score = 180 bits (456), Expect = 1e-45
Identities = 117/312 (37%), Positives = 174/312 (55%), Gaps = 25/312 (8%)
Query: 75 YMAERVVGHGSFGVVFQAKCLETGETVAIKKV------LQDKRYKNRELQTMRLLDHPNV 128
Y + VVG GS+GVV A TGE VAIKK+ + D RE++ +RLL HP+V
Sbjct: 90 YQIQEVVGKGSYGVVGSAIDTHTGERVAIKKINDVFDHISDATRILREIKLLRLLLHPDV 149
Query: 129 VSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALS 188
V +KH + ++ + +V E + +H+VIK N + + + + YQ+ R L
Sbjct: 150 VEIKHIMLPPSRREFRDVYVVFELMESDLHQVIK----ANDDLTPEHHQFFLYQLLRGLK 205
Query: 189 YIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNI----SYICSRYYRAP 244
Y+H V HRD+KP+N+L N ++K+CDFG A+V P Y+ +R+YRAP
Sbjct: 206 YVHAA-NVFHRDLKPKNILANADC-KLKICDFGLARVSFNDAPTAIFWTDYVATRWYRAP 263
Query: 245 ELIFGA-TEYTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEI-KC 302
EL ++YT AID+WSVGC+ AE+LLG+PLFPG++ V QL + LGTP E I K
Sbjct: 264 ELCGSFFSKYTPAIDIWSVGCIFAEMLLGKPLFPGKNVVHQLDIMTDFLGTPPPEAISKI 323
Query: 303 MNPNYTEFKFPQIKAH--PWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFD 360
N + K P+ K F K P+ A+ L+ RL+ + P R A + L P+F+
Sbjct: 324 RNDKARRYLGNMRKKQPVPFSKKFPKADPS-ALRLLERLIAFDPKDRPSAEEALADPYFN 382
Query: 361 ----ELRDPNAR 368
++R+P+ +
Sbjct: 383 GLSSKVREPSTQ 394
>At3g59790 mitogen-activated protein kinase-like protein
Length = 393
Score = 179 bits (453), Expect = 3e-45
Identities = 101/294 (34%), Positives = 162/294 (54%), Gaps = 15/294 (5%)
Query: 79 RVVGHGSFGVVFQAKCLETGETVAIKKVLQ------DKRYKNRELQTMRLLDHPNVVSLK 132
R +G G+ G+V A ET E VAIKK+ Q + + RE++ +R DH N+V+++
Sbjct: 64 RPIGRGACGIVCSAVDSETNEKVAIKKITQVFDNTIEAKRTLREIKLLRHFDHENIVAIR 123
Query: 133 HCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHR 192
+ +V E + ++R +K +L + M ++ YQI R L YIH
Sbjct: 124 DVILPPQRDSFEDVYIVNELMEFDLYRTLKSDQELTKDHGMYFM----YQILRGLKYIHS 179
Query: 193 CIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATE 252
V HRD+KP NLL++ +K+CDFG A+ + Y+ +R+YRAPEL+ G+++
Sbjct: 180 A-NVLHRDLKPSNLLLSTQC-DLKICDFGLARATPESNLMTEYVVTRWYRAPELLLGSSD 237
Query: 253 YTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKF 312
YT AIDVWSVGC+ E++ +PLFPG+ V+QL +++++GTP+ EE+ ++ Y +
Sbjct: 238 YTAAIDVWSVGCIFMEIMNREPLFPGKDQVNQLRLLLELIGTPSEEELGSLS-EYAKRYI 296
Query: 313 PQIKAHPWHKIFHK--RMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRD 364
Q+ P K +P A+DLV ++L + P R + L HP+ D
Sbjct: 297 RQLPTLPRQSFTEKFPNVPPLAIDLVEKMLTFDPKQRISVKEALAHPYLSSFHD 350
>At3g45640 mitogen-activated protein kinase 3
Length = 370
Score = 178 bits (452), Expect = 4e-45
Identities = 100/295 (33%), Positives = 163/295 (54%), Gaps = 16/295 (5%)
Query: 81 VGHGSFGVVFQAKCLETGETVAIKKVLQ------DKRYKNRELQTMRLLDHPNVVSLKHC 134
+G G++G+V ET E VA+KK+ D + RE++ +R LDH N+++++
Sbjct: 44 IGRGAYGIVCSVLDTETNELVAMKKIANAFDNHMDAKRTLREIKLLRHLDHENIIAIRDV 103
Query: 135 FFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRCI 194
+ + + E + +H++I+ NQ + + + + YQ+ R L YIH
Sbjct: 104 VPPPLRRQFSDVYISTELMDTDLHQIIRS----NQSLSEEHCQYFLYQLLRGLKYIHSA- 158
Query: 195 GVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYT 254
+ HRD+KP NLL+N + +K+CDFG A+ + + Y+ +R+YRAPEL+ +++YT
Sbjct: 159 NIIHRDLKPSNLLLNANC-DLKICDFGLARPTSENDFMTEYVVTRWYRAPELLLNSSDYT 217
Query: 255 TAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIK-CMNPNYTEF--K 311
AIDVWSVGC+ EL+ +PLFPG+ V Q+ + ++LGTPT ++ N + + +
Sbjct: 218 AAIDVWSVGCIFMELMNRKPLFPGKDHVHQMRLLTELLGTPTESDLGFTHNEDAKRYIRQ 277
Query: 312 FPQIKAHPWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRDPN 366
P P K+F P A+DLV R+L + PN R L H + +L DPN
Sbjct: 278 LPNFPRQPLAKLFSHVNPM-AIDLVDRMLTFDPNRRITVEQALNHQYLAKLHDPN 331
>At2g42880 MAP kinase 4 homolog
Length = 606
Score = 176 bits (446), Expect = 2e-44
Identities = 109/311 (35%), Positives = 172/311 (55%), Gaps = 23/311 (7%)
Query: 75 YMAERVVGHGSFGVVFQAKCLETGETVAIKKV------LQDKRYKNRELQTMRLLDHPNV 128
+ + V+G GS+GVV A TGE VAIKK+ + D RE++ +RLL HP++
Sbjct: 25 FKVQEVIGKGSYGVVCSAIDTLTGEKVAIKKIHDIFEHISDAARILREIKLLRLLRHPDI 84
Query: 129 VSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALS 188
V +KH + ++ + +V E + +H+VIK L + + + + YQ+ RAL
Sbjct: 85 VEIKHIMLPPSRREFKDIYVVFELMESDLHQVIKANDDLTRE----HYQFFLYQLLRALK 140
Query: 189 YIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNI----SYICSRYYRAP 244
YIH V HRD+KP+N+L N + ++K+CDFG A+V P Y+ +R+YRAP
Sbjct: 141 YIHTA-NVYHRDLKPKNILANANC-KLKICDFGLARVAFNDTPTTIFWTDYVATRWYRAP 198
Query: 245 ELIFGA-TEYTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCM 303
EL ++YT AID+WS+GC+ AE+L+G+PLFPG++ V QL + +LGTP+ + I +
Sbjct: 199 ELCGSFYSKYTPAIDIWSIGCIFAEVLMGKPLFPGKNVVHQLDLMTDLLGTPSLDTISRV 258
Query: 304 NPNYTEFKFPQIKAHPWHKIFHKRMPAE--AVDLVSRLLQYSPNLRCQALDCLTHPFFDE 361
++ P K A+ ++ L+ RLL + P R A + L P+F
Sbjct: 259 RNEKARRYLTSMRKKPPIPFAQKFPNADPLSLKLLERLLAFDPKDRPTAEEALADPYFKG 318
Query: 362 L----RDPNAR 368
L R+P+ +
Sbjct: 319 LAKVEREPSCQ 329
>At5g19010 MAP kinase -like protein
Length = 567
Score = 175 bits (444), Expect = 3e-44
Identities = 114/313 (36%), Positives = 170/313 (53%), Gaps = 27/313 (8%)
Query: 75 YMAERVVGHGSFGVVFQAKCLETGETVAIKKV------LQDKRYKNRELQTMRLLDHPNV 128
Y E V+G GS+GVV A TGE VAIKK+ + D RE++ +RLL HP++
Sbjct: 25 YRIEEVIGKGSYGVVCSAYDTHTGEKVAIKKINDIFEHVSDATRILREIKLLRLLRHPDI 84
Query: 129 VSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALS 188
V +KH + ++ + +V E + +H+VIK N + + + + YQ+ R L
Sbjct: 85 VEIKHILLPPSRREFRDIYVVFELMESDLHQVIK----ANDDLTPEHYQFFLYQLLRGLK 140
Query: 189 YIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNI----SYICSRYYRAP 244
YIH V HRD+KP+N+L N ++K+CDFG A+V P Y+ +R+YRAP
Sbjct: 141 YIHTA-NVFHRDLKPKNILANADC-KLKICDFGLARVAFNDTPTAIFWTDYVATRWYRAP 198
Query: 245 ELIFGA-TEYTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEI-KC 302
EL ++YT AID+WS+GC+ AELL G+PLFPG++ V QL + +LGTP+ E I +
Sbjct: 199 ELCGSFFSKYTPAIDIWSIGCIFAELLTGKPLFPGKNVVHQLDLMTDMLGTPSAEAIGRV 258
Query: 303 MNPNYTEFKFPQIKAHP---WHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFF 359
N + K P HK H A+ L+ ++L + P R A + L +F
Sbjct: 259 RNEKARRYLSSMRKKKPIPFSHKFPH--TDPLALRLLEKMLSFEPKDRPTAEEALADVYF 316
Query: 360 DEL----RDPNAR 368
L R+P+A+
Sbjct: 317 KGLAKVEREPSAQ 329
>At4g01370 MAP kinase 4
Length = 376
Score = 175 bits (443), Expect = 4e-44
Identities = 102/297 (34%), Positives = 166/297 (55%), Gaps = 16/297 (5%)
Query: 79 RVVGHGSFGVVFQAKCLETGETVAIKKV------LQDKRYKNRELQTMRLLDHPNVVSLK 132
R +G G++G+V A ETGE VAIKK+ + D + RE++ ++ +DH NV+++K
Sbjct: 47 RPIGRGAYGIVCAATNSETGEEVAIKKIGNAFDNIIDAKRTLREIKLLKHMDHENVIAVK 106
Query: 133 HCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHR 192
++ + +V E + +H++I+ NQ + + + + YQ+ R L Y+H
Sbjct: 107 DIIKPPQRENFNDVYIVYELMDTDLHQIIRS----NQPLTDDHCRFFLYQLLRGLKYVHS 162
Query: 193 CIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATE 252
V HRD+KP NLL+N + +KL DFG A+ + + Y+ +R+YRAPEL+ +E
Sbjct: 163 A-NVLHRDLKPSNLLLNANC-DLKLGDFGLARTKSETDFMTEYVVTRWYRAPELLLNCSE 220
Query: 253 YTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKF 312
YT AID+WSVGC+L E + +PLFPG+ V QL I +++G+P + + +
Sbjct: 221 YTAAIDIWSVGCILGETMTREPLFPGKDYVHQLRLITELIGSPDDSSLGFLRSDNARRYV 280
Query: 313 PQIKAHPWHKIFHKRMP---AEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRDPN 366
Q+ +P + F R P A AVDL+ ++L + P+ R + L HP+ L D N
Sbjct: 281 RQLPQYP-RQNFAARFPNMSAGAVDLLEKMLVFDPSRRITVDEALCHPYLAPLHDIN 336
>At3g18040 MAP kinase like protein
Length = 510
Score = 175 bits (443), Expect = 4e-44
Identities = 113/322 (35%), Positives = 176/322 (54%), Gaps = 21/322 (6%)
Query: 75 YMAERVVGHGSFGVVFQAKCLETGETVAIKKV------LQDKRYKNRELQTMRLLDHPNV 128
Y + V+G GS+GVV A +GE VAIKK+ + D RE++ +RLL HP++
Sbjct: 23 YQIQEVIGKGSYGVVASAIDTHSGEKVAIKKINDVFEHVSDATRILREIKLLRLLRHPDI 82
Query: 129 VSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALS 188
V +KH + ++ + +V E + +H+VIK N + + + + YQ+ R L
Sbjct: 83 VEIKHVMLPPSRREFRDIYVVFELMESDLHQVIK----ANDDLTPEHYQFFLYQLLRGLK 138
Query: 189 YIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNI----SYICSRYYRAP 244
+IH V HRD+KP+N+L N ++K+CDFG A+V P+ Y+ +R+YRAP
Sbjct: 139 FIHTA-NVFHRDLKPKNILANSDC-KLKICDFGLARVSFNDAPSAIFWTDYVATRWYRAP 196
Query: 245 ELIFGA-TEYTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCM 303
EL ++YT AID+WS+GC+ AE+L G+PLFPG++ V QL + +LGTP E I +
Sbjct: 197 ELCGSFFSKYTPAIDIWSIGCIFAEMLTGKPLFPGKNVVHQLDIMTDLLGTPPPEAIARI 256
Query: 304 NPNYTEFKFPQIKAHPWHKIFHK--RMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDE 361
++ P HK + A+ L+ RLL + P R A + L P+F
Sbjct: 257 RNEKARRYLGNMRRKPPVPFTHKFPHVDPLALRLLHRLLAFDPKDRPSAEEALADPYFYG 316
Query: 362 LRDPNARLPTGRFLPPL-FNFK 382
L + + R P+ + +P L F F+
Sbjct: 317 LANVD-REPSTQPIPKLEFEFE 337
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.140 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,106,906
Number of Sequences: 26719
Number of extensions: 406450
Number of successful extensions: 3143
Number of sequences better than 10.0: 920
Number of HSP's better than 10.0 without gapping: 278
Number of HSP's successfully gapped in prelim test: 642
Number of HSP's that attempted gapping in prelim test: 1474
Number of HSP's gapped (non-prelim): 1061
length of query: 384
length of database: 11,318,596
effective HSP length: 101
effective length of query: 283
effective length of database: 8,619,977
effective search space: 2439453491
effective search space used: 2439453491
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147011.7


