

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147011.10 - phase: 0
(165 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
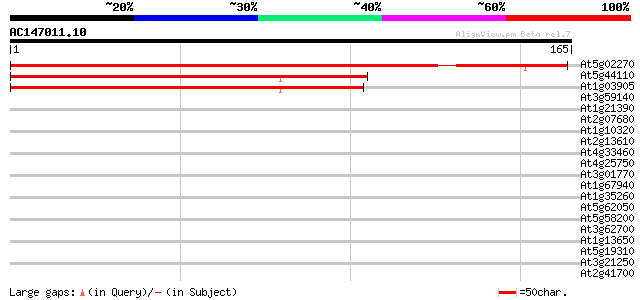
Score E
Sequences producing significant alignments: (bits) Value
At5g02270 ABC transporter -like protein 259 4e-70
At5g44110 NBD-like protein (gb|AAD20643.1) 122 9e-29
At1g03905 unknown protein 116 5e-27
At3g59140 ABC transporter-like protein 31 0.26
At1g21390 hypothetical protein 31 0.26
At2g07680 putative ABC transporter 31 0.35
At1g10320 unknown protein 29 1.0
At2g13610 putative ABC transporter 29 1.3
At4g33460 unknown protein 28 1.7
At4g25750 putative membrane transporter 28 1.7
At3g01770 unknown protein 28 1.7
At1g67940 ABC transporter like protein 28 1.7
At1g35260 unknown protein 28 1.7
At5g62050 AtOXA1 28 2.2
At5g58200 phosphoesterase like protein 28 2.2
At3g62700 ABC transporter-like protein 28 2.2
At1g13650 hypothetical protein 28 2.2
At5g19310 homeotic gene regulator - like protein 28 2.9
At3g21250 unknown protein 28 2.9
At2g41700 ATP-binding cassette transporter ABCA1 27 3.8
>At5g02270 ABC transporter -like protein
Length = 328
Score = 259 bits (663), Expect = 4e-70
Identities = 131/165 (79%), Positives = 149/165 (89%), Gaps = 6/165 (3%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLLKPF+VLLLDEITVDLDVLARADLLKFLRKEC+ERGATIIYATHIFDGLEDWPT+IV
Sbjct: 169 MGLLKPFKVLLLDEITVDLDVLARADLLKFLRKECEERGATIIYATHIFDGLEDWPTHIV 228
Query: 61 YVAHGRLELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPEFDKQVDG 120
YVA+G+L+LA+P+EKVKETSK SLMRTVE WLRKERDE+R++RKERKA GLPEF+ + +
Sbjct: 229 YVANGKLQLALPMEKVKETSKKSLMRTVESWLRKERDEERKRRKERKANGLPEFETRTEE 288
Query: 121 SRVVGGDPARAPVRVTNNGWAAGRLHSTIA-GEENFLLSSNRVLR 164
SRV G P R+ NNGWAAGRLHST+A GE+NF+LSSNRVLR
Sbjct: 289 SRVTGD-----PARMLNNGWAAGRLHSTVAGGEDNFVLSSNRVLR 328
>At5g44110 NBD-like protein (gb|AAD20643.1)
Length = 282
Score = 122 bits (306), Expect = 9e-29
Identities = 55/106 (51%), Positives = 83/106 (77%), Gaps = 1/106 (0%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLL PF+VLLLDE+TVDLDV+AR DLL+F ++EC++RGATI+YATHIFDGLE W +++
Sbjct: 160 MGLLHPFKVLLLDEVTVDLDVVARMDLLEFFKEECEQRGATIVYATHIFDGLETWASHLA 219
Query: 61 YVAHGRLELAMPIEKVKE-TSKLSLMRTVEVWLRKERDEDRRQRKE 105
Y+ G L+L+ ++++K+ + +L+ VE WLR E +++ +K+
Sbjct: 220 YINGGELKLSAKLDEIKDLKTSPNLLSVVEAWLRSETKVEKKTKKK 265
>At1g03905 unknown protein
Length = 290
Score = 116 bits (291), Expect = 5e-27
Identities = 56/105 (53%), Positives = 77/105 (73%), Gaps = 1/105 (0%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLL PF+VLLLDE+TVDLDV+AR DLL+F ++ECD+RGATI+YATHIFDGLE W T++
Sbjct: 161 MGLLHPFKVLLLDEVTVDLDVVARMDLLEFFKEECDQRGATIVYATHIFDGLETWATHLA 220
Query: 61 YVAHGRLELAMPIEKVKE-TSKLSLMRTVEVWLRKERDEDRRQRK 104
Y+ G L + ++E + +L+ VE WLR E ++++K
Sbjct: 221 YIQDGELNRLSKMTDIEELKTSPNLLSVVESWLRSEIKLVKKKKK 265
>At3g59140 ABC transporter-like protein
Length = 1389
Score = 31.2 bits (69), Expect = 0.26
Identities = 25/90 (27%), Positives = 45/90 (49%), Gaps = 4/90 (4%)
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYV 62
+L+ +VL+LDE T +D L K +R+E + T+I H + D T ++ +
Sbjct: 1294 VLRRSRVLVLDEATASIDNATDLILQKTIRREFAD--CTVITVAHRIPTVMDC-TMVLSI 1350
Query: 63 AHGRL-ELAMPIEKVKETSKLSLMRTVEVW 91
+ GR+ E P++ +K+ + L E W
Sbjct: 1351 SDGRIVEYDEPMKLMKDENSLFGKLVKEYW 1380
>At1g21390 hypothetical protein
Length = 260
Score = 31.2 bits (69), Expect = 0.26
Identities = 22/73 (30%), Positives = 38/73 (51%), Gaps = 7/73 (9%)
Query: 62 VAHGRLELAMPIEKVKETS-KLSLMRTVEVWLRKERDED------RRQRKERKAAGLPEF 114
+A G+ EL + K+ E+ +LSL VEV + +E + +R ++ K +
Sbjct: 64 IARGQRELMEMVSKMPESCYELSLKDLVEVKVNQENERKVFDELPKRANRQSKVVRKTKS 123
Query: 115 DKQVDGSRVVGGD 127
DK+VD +R GG+
Sbjct: 124 DKRVDPNRSGGGN 136
>At2g07680 putative ABC transporter
Length = 1146
Score = 30.8 bits (68), Expect = 0.35
Identities = 17/45 (37%), Positives = 25/45 (54%), Gaps = 2/45 (4%)
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATH 47
LLK ++L LDE T ++DV + L + EC +G T+I H
Sbjct: 1060 LLKSSKILCLDECTANIDVHTASLLHNTISSEC--KGVTVITIAH 1102
>At1g10320 unknown protein
Length = 757
Score = 29.3 bits (64), Expect = 1.0
Identities = 14/46 (30%), Positives = 25/46 (53%)
Query: 77 KETSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPEFDKQVDGSR 122
KE+ K + T V + ++D DR ++++R + PE D+ G R
Sbjct: 664 KESDKKRSVETSPVGYQSDKDRDRSKQRQRYKSDDPESDQSRKGKR 709
>At2g13610 putative ABC transporter
Length = 649
Score = 28.9 bits (63), Expect = 1.3
Identities = 15/40 (37%), Positives = 22/40 (54%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATH 47
+VL+LDE T LD + ++ L+ + RG TII H
Sbjct: 202 KVLILDEPTSGLDSTSALLIIDMLKHMAETRGRTIILTIH 241
>At4g33460 unknown protein
Length = 271
Score = 28.5 bits (62), Expect = 1.7
Identities = 19/67 (28%), Positives = 36/67 (53%), Gaps = 3/67 (4%)
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRK--ECDERGATIIYATHIFDGLEDWPTNIV 60
L + +VLLLDE+T LD + ++K ++ + T ++ TH + L+ + V
Sbjct: 180 LAEACKVLLLDELTTFLDESDQMGVIKAVKDLINAKKGDVTALWVTHRLEELK-YADGAV 238
Query: 61 YVAHGRL 67
Y+ +GR+
Sbjct: 239 YMENGRV 245
>At4g25750 putative membrane transporter
Length = 577
Score = 28.5 bits (62), Expect = 1.7
Identities = 16/47 (34%), Positives = 25/47 (53%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATH 47
+ LL +VLLLDE T LD + D+++ L+ R +I + H
Sbjct: 159 LSLLHDPEVLLLDEPTSGLDSKSAFDVVQILKSIATSRERIVILSIH 205
>At3g01770 unknown protein
Length = 620
Score = 28.5 bits (62), Expect = 1.7
Identities = 18/68 (26%), Positives = 32/68 (46%), Gaps = 1/68 (1%)
Query: 51 GLEDWPTNIVYVAHGRLELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKERKAAG 110
G + P+N+ +AH + + P+ K ++ + + +E R DED R + R
Sbjct: 236 GTKSEPSNLATLAHKDIAIPEPVAKKRKMNAVKRNSLLEPAKRVMTDED-RVKLGRDLGS 294
Query: 111 LPEFDKQV 118
L EF Q+
Sbjct: 295 LTEFPVQI 302
>At1g67940 ABC transporter like protein
Length = 263
Score = 28.5 bits (62), Expect = 1.7
Identities = 19/68 (27%), Positives = 33/68 (47%), Gaps = 1/68 (1%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
+VLLLDE T LD ++ ++ + K +RG T + +H ++ + V G +
Sbjct: 179 EVLLLDEPTSALDPISTENIEDVIVKLKKQRGITTVIVSHSIKQIQKVADIVCLVVDGEI 238
Query: 68 -ELAMPIE 74
E+ P E
Sbjct: 239 VEVLKPSE 246
>At1g35260 unknown protein
Length = 152
Score = 28.5 bits (62), Expect = 1.7
Identities = 28/98 (28%), Positives = 45/98 (45%), Gaps = 18/98 (18%)
Query: 10 LLLDEITVDLDVLARADLL-KFLRKECDERGATIIYATHIFDGLE----DWPT------- 57
++ +EI VD+D+ RAD KF+R R + ATH G + +W
Sbjct: 1 MIEEEIEVDVDIKTRADKFHKFIR-----RSQHVPKATHYIKGCDLLEGEWGKVGSILLW 55
Query: 58 NIVYVAHGRLELAMPIEKVKETSKLSLMRTVEVWLRKE 95
+V+ R+ M IE + E + +R +E L+KE
Sbjct: 56 KLVFDGEPRVSKDM-IEVIDEEKNVIQLRVLEGPLKKE 92
>At5g62050 AtOXA1
Length = 429
Score = 28.1 bits (61), Expect = 2.2
Identities = 21/73 (28%), Positives = 36/73 (48%), Gaps = 6/73 (8%)
Query: 47 HIFDGLEDWPTNIVYVAHGRLE-LAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKE 105
H F G E W + +V R + + I+++K+T+KL+LMR R E + Q K
Sbjct: 145 HTFTGFEWWASIVVATILIRSSTVPLLIKQMKDTTKLALMRP-----RLESIREEMQNKG 199
Query: 106 RKAAGLPEFDKQV 118
+ + E K++
Sbjct: 200 MDSVTMAEGQKKM 212
>At5g58200 phosphoesterase like protein
Length = 222
Score = 28.1 bits (61), Expect = 2.2
Identities = 15/41 (36%), Positives = 22/41 (53%)
Query: 64 HGRLELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRK 104
HG +L I ++KET+KLS+ V + KE + RK
Sbjct: 103 HGDPDLEQAIRQLKETTKLSIPLVVFGHMHKELQRGKGNRK 143
>At3g62700 ABC transporter-like protein
Length = 1539
Score = 28.1 bits (61), Expect = 2.2
Identities = 21/78 (26%), Positives = 35/78 (43%), Gaps = 2/78 (2%)
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYV 62
+LK ++L LDE T +D A + K +R++ + TII H + D +V
Sbjct: 1448 MLKRSRILFLDEATASVDSQTDAMIQKIIREDFSD--CTIISIAHRIPTVMDCDRVLVID 1505
Query: 63 AHGRLELAMPIEKVKETS 80
A E P+ ++ S
Sbjct: 1506 AGKAKEYDSPVRLLERQS 1523
>At1g13650 hypothetical protein
Length = 281
Score = 28.1 bits (61), Expect = 2.2
Identities = 16/34 (47%), Positives = 23/34 (67%), Gaps = 1/34 (2%)
Query: 74 EKVKETSKLSLMRTVEVWLRKERDEDRRQRKERK 107
EK +ET +L ++ E LRKE+DE R+ RK +K
Sbjct: 236 EKAEET-QLRMVIAEERQLRKEKDEKRQLRKGKK 268
>At5g19310 homeotic gene regulator - like protein
Length = 1041
Score = 27.7 bits (60), Expect = 2.9
Identities = 12/52 (23%), Positives = 27/52 (51%)
Query: 62 VAHGRLELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPE 113
++ G L + +E ++L+ E W+ ++ DE+RR+++ K + E
Sbjct: 843 MSKGTSSLGEDVPSEREINRLAARTEEEFWMFEQMDEERRKKENYKTRLMEE 894
>At3g21250 unknown protein
Length = 1305
Score = 27.7 bits (60), Expect = 2.9
Identities = 23/89 (25%), Positives = 39/89 (42%), Gaps = 3/89 (3%)
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYV 62
LLK ++L+LDE T +D A + + +R+E + T+I H + D ++ +
Sbjct: 1207 LLKRNKILVLDEATASIDSATDAIIQRIIREEFAD--CTVITVAHRVPTVID-SDMVMVL 1263
Query: 63 AHGRLELAMPIEKVKETSKLSLMRTVEVW 91
+ G L K+ ET E W
Sbjct: 1264 SFGDLVEYNEPSKLMETDSYFSKLVAEYW 1292
>At2g41700 ATP-binding cassette transporter ABCA1
Length = 1884
Score = 27.3 bits (59), Expect = 3.8
Identities = 18/68 (26%), Positives = 35/68 (51%), Gaps = 2/68 (2%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
+ L+ +V++LDE T +D + + ++K ++G I+ TH D E+ I
Sbjct: 660 IALIGNSKVIILDEPTSGMDPYSMRLTWQLIKKI--KKGRIILLTTHSMDEAEELGDRIG 717
Query: 61 YVAHGRLE 68
+A+G L+
Sbjct: 718 IMANGSLK 725
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.139 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,543,020
Number of Sequences: 26719
Number of extensions: 144308
Number of successful extensions: 585
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 18
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 561
Number of HSP's gapped (non-prelim): 48
length of query: 165
length of database: 11,318,596
effective HSP length: 92
effective length of query: 73
effective length of database: 8,860,448
effective search space: 646812704
effective search space used: 646812704
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC147011.10


