

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147010.4 - phase: 0 /pseudo
(618 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
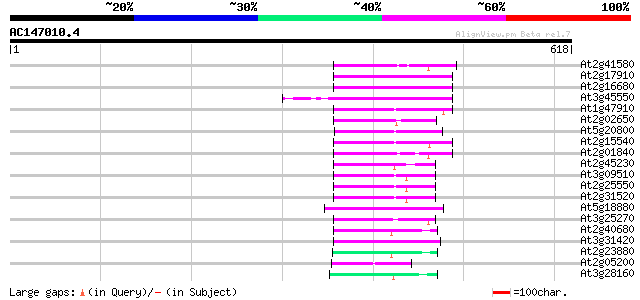
Score E
Sequences producing significant alignments: (bits) Value
At2g41580 putative non-LTR retroelement reverse transcriptase 76 5e-14
At2g17910 putative non-LTR retroelement reverse transcriptase 73 4e-13
At2g16680 putative non-LTR retroelement reverse transcriptase 72 9e-13
At3g45550 putative protein 70 3e-12
At1g47910 reverse transcriptase, putative 68 1e-11
At2g02650 putative reverse transcriptase 62 7e-10
At5g20800 putative protein 62 9e-10
At2g15540 putative non-LTR retroelement reverse transcriptase 60 3e-09
At2g01840 putative non-LTR retroelement reverse transcriptase 60 4e-09
At2g45230 putative non-LTR retroelement reverse transcriptase 60 5e-09
At3g09510 putative non-LTR reverse transcriptase 59 6e-09
At2g25550 putative non-LTR retroelement reverse transcriptase 59 6e-09
At2g31520 putative non-LTR retroelement reverse transcriptase 58 2e-08
At5g18880 SAE1-S9-protein - like 56 7e-08
At3g25270 hypothetical protein 56 7e-08
At2g40680 putative non-LTR retroelement reverse transcriptase 53 4e-07
At3g31420 hypothetical protein 53 6e-07
At2g23880 putative non-LTR retroelement reverse transcriptase 52 1e-06
At2g05200 putative non-LTR retroelement reverse transcriptase 52 1e-06
At3g28160 non-LTR reverse transcriptase, putative 52 1e-06
>At2g41580 putative non-LTR retroelement reverse transcriptase
Length = 1094
Score = 76.3 bits (186), Expect = 5e-14
Identities = 44/140 (31%), Positives = 68/140 (48%), Gaps = 7/140 (5%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IW L PK K +W + +G RL +G+ C CP +E ++HI F+CP A
Sbjct: 775 IWNLNSAPKIKVFLWKVLKGAVAVEDRLRTRGVLIEDGCSMCPEKNETLNHILFQCPLAR 834
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTS----LFWSLWKHRNLRA 472
VW +T + H + NV ++ + EL+ L + W LWK+RN R
Sbjct: 835 QVWALTPMQSP-NHGFGDSIFT-NVNHVIGNCHNTELSPHLRYVSPWIIWILWKNRNKRL 892
Query: 473 WEDVTEVSVVVVERA*LAEC 492
+E + VS+ +V +A L +C
Sbjct: 893 FEGIGSVSLSIVGKA-LEDC 911
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 73.2 bits (178), Expect = 4e-13
Identities = 39/133 (29%), Positives = 64/133 (47%), Gaps = 2/133 (1%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IW L PK + +W G P RL +GI+ C+ C + +E ++HI FECP A
Sbjct: 1024 IWNLHTAPKIRIFLWKALHGAIPVEDRLRTRGIRSDDGCLMCDTENETINHILFECPLAR 1083
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTS--LFWSLWKHRNLRAWE 474
VW +T L +S + ++ + L + + R S + W LWK+RN +E
Sbjct: 1084 QVWAITHLSSAGSEFSNSVYTNMSRLIDLTQQNDLPHHLRFVSPWILWFLWKNRNALLFE 1143
Query: 475 DVTEVSVVVVERA 487
++ +V++A
Sbjct: 1144 GKGSITTTLVDKA 1156
>At2g16680 putative non-LTR retroelement reverse transcriptase
Length = 1319
Score = 72.0 bits (175), Expect = 9e-13
Identities = 38/133 (28%), Positives = 62/133 (46%), Gaps = 2/133 (1%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
+W L+ PK K +W +G RL +GI+ C+ C E ++H+ F+CPFA
Sbjct: 998 VWSLETVPKIKVFMWKALKGALAVEDRLRSRGIRTADGCLFCKEEIETINHLLFQCPFAR 1057
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTS--LFWSLWKHRNLRAWE 474
VW ++ + +S S IN + + F + R S L W +WK+RN ++
Sbjct: 1058 QVWALSLIQAPATGFGTSIFSNINHVIQNSQNFGIPRHMRTVSPWLLWEIWKNRNKTLFQ 1117
Query: 475 DVTEVSVVVVERA 487
S +V +A
Sbjct: 1118 GTGLTSSEIVAKA 1130
>At3g45550 putative protein
Length = 851
Score = 70.1 bits (170), Expect = 3e-12
Identities = 52/190 (27%), Positives = 84/190 (43%), Gaps = 21/190 (11%)
Query: 301 NSTGVQC*HC*ENSSNAASWSSTRGPSHLVLCS*CLLVMCHGTR*LFSPYVGFWSGIWKL 360
NSTG + + + W ST PS+ + + HG+ V + IW L
Sbjct: 629 NSTG-------DYTVRSGYWLSTHDPSNTIPT----MAKPHGS-------VDLKTKIWNL 670
Query: 361 KVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAIHVW* 420
+ PK K+ +W + PT RL +G++ C C +E ++H F CPFA W
Sbjct: 671 PIMPKLKHFLWRILSKALPTTDRLTTRGMRIDPGCPRCRRENESINHALFTCPFATMAWR 730
Query: 421 MTSLWGTIQHALSST-TSAINVIFMLLETFSAELNQRLTS--LFWSLWKHRNLRAWEDVT 477
++ LS+ I+ I +LL+ + +Q+L L W +WK RN + ++
Sbjct: 731 LSDTPLYRSSILSNNIEDNISNILLLLQNTTITDSQKLIPFWLLWRIWKARNNVVFNNLR 790
Query: 478 EVSVVVVERA 487
E + V RA
Sbjct: 791 ESPSITVVRA 800
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 68.2 bits (165), Expect = 1e-11
Identities = 37/135 (27%), Positives = 66/135 (48%), Gaps = 5/135 (3%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IWK++ PPK ++ +W + G P L +GI C CVSC ++ E ++H F+C A
Sbjct: 828 IWKVQCPPKLRHFLWQILSGCVPVSENLRKRGILCDKGCVSCGASEESINHTLFQCHPAR 887
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSLWKHRNLRAWEDV 476
+W ++ + T S + N+ + S + + W +WK RN + +E+V
Sbjct: 888 QIWALSQI-PTAPGIFPSNSIFTNLDHLFWRIPSGVDSAPYPWIIWYIWKARNEKVFENV 946
Query: 477 ----TEVSVVVVERA 487
E+ ++ V+ A
Sbjct: 947 DKDPMEILLLAVKEA 961
>At2g02650 putative reverse transcriptase
Length = 365
Score = 62.4 bits (150), Expect = 7e-10
Identities = 38/121 (31%), Positives = 51/121 (41%), Gaps = 11/121 (9%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IWKL V PK K+ +W G T RL + I C C E + HI F CP+
Sbjct: 37 IWKLHVAPKIKHFLWRCVTGALATNTRLRSRNIDADPICQRCCIEEETIHHIMFNCPYTQ 96
Query: 417 HVW*MTSL-----WGTIQHALSSTTSAINVIFMLLETFSAELNQRLTS--LFWSLWKHRN 469
VW ++ WG SS +N + L +T + R + W LWK RN
Sbjct: 97 SVWRSANIIIGNQWG----PPSSFEDNLNRLIQLSKTQTTNSLDRFLPFWIMWRLWKSRN 152
Query: 470 L 470
+
Sbjct: 153 V 153
>At5g20800 putative protein
Length = 359
Score = 62.0 bits (149), Expect = 9e-10
Identities = 30/119 (25%), Positives = 57/119 (47%), Gaps = 1/119 (0%)
Query: 358 WKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAIH 417
WK++ PPK ++ +W + P R L +GI C C C + + ++H F+C A
Sbjct: 128 WKVQCPPKLRHFLWQILASCVPVRENLRKRGINCDIGCARCGATEKTINHTLFQCHPARQ 187
Query: 418 VW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSLWKHRNLRAWEDV 476
+W ++ + T+ S + N+ + SA + + W +WK RN + +++V
Sbjct: 188 IWALSQI-PTVIGIFPSDSIYENLDHLFWRVQSAVDSFSYPWIIWYIWKARNAKLFDNV 245
>At2g15540 putative non-LTR retroelement reverse transcriptase
Length = 1225
Score = 60.5 bits (145), Expect = 3e-09
Identities = 34/135 (25%), Positives = 63/135 (46%), Gaps = 5/135 (3%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
+WK+K KFK+ W G T RL + I C C + E ++H+ F CP +
Sbjct: 926 VWKIKTTRKFKHFEWQCLSGCLATNQRLFSRHIGTEKVCPRCGAEEESINHLLFLCPPSR 985
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLF----WSLWKHRNLRA 472
+W ++ + + ++ + N F+L ++ + + +F W +WK RN
Sbjct: 986 QIWALSPI-PSSEYIFPRNSLFYNFDFLLSRGKEFDIAEDIMEIFPWILWYIWKSRNRFI 1044
Query: 473 WEDVTEVSVVVVERA 487
+E+V E V+++ A
Sbjct: 1045 FENVIESPQVILDFA 1059
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 60.1 bits (144), Expect = 4e-09
Identities = 39/139 (28%), Positives = 63/139 (45%), Gaps = 13/139 (9%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IW+LK+ PK K+ +W G T +L ++ I C C + E ++HI F C +A
Sbjct: 1388 IWRLKITPKIKHFIWRCLSGALSTTTQLRNRNIPADPTCQRCCNADETINHIIFTCSYAQ 1447
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSL--------FWSLWKHR 468
VW + G+ + L T + I ++L+ + NQ L L W LWK R
Sbjct: 1448 VVWRSANFSGS--NRLCFTDNLEENIRLILQ---GKKNQNLPILNGLMPFWIMWRLWKSR 1502
Query: 469 NLRAWEDVTEVSVVVVERA 487
N ++ + V ++A
Sbjct: 1503 NEYLFQQLDRFPWKVAQKA 1521
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 59.7 bits (143), Expect = 5e-09
Identities = 35/125 (28%), Positives = 56/125 (44%), Gaps = 21/125 (16%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IWKL VPPK + +W L + + +CV CPS+ E ++H+ F+CPFA
Sbjct: 1054 IWKLDVPPKIHHFLWRCVNNCLSVASNLAYRHLAREKSCVRCPSHGETVNHLLFKCPFAR 1113
Query: 417 HVW*MT------------SLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSL 464
W ++ SL+ + H LS S + ++ + + + W L
Sbjct: 1114 LTWAISPLPAPPGGEWAESLFRNMHHVLSVHKS---------QPEESDHHALIPWILWRL 1164
Query: 465 WKHRN 469
WK+RN
Sbjct: 1165 WKNRN 1169
>At3g09510 putative non-LTR reverse transcriptase
Length = 484
Score = 59.3 bits (142), Expect = 6e-09
Identities = 33/117 (28%), Positives = 57/117 (48%), Gaps = 5/117 (4%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IW L + PK K+ +W T RL +G++ +C C +E ++H F CPFA
Sbjct: 162 IWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPSCPRCHRENESINHALFTCPFAT 221
Query: 417 HVW*MTSLWGTIQHALSST---TSAINVIFMLLETFSAELNQRL-TSLFWSLWKHRN 469
W ++ I++ L S + N++ + +T ++ ++ L L W +WK RN
Sbjct: 222 MAWRLSDS-SLIRNQLMSNDFEENISNILNFVQDTTMSDFHKLLPVWLIWRIWKARN 277
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 59.3 bits (142), Expect = 6e-09
Identities = 33/117 (28%), Positives = 57/117 (48%), Gaps = 5/117 (4%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IW L + PK K+ +W T RL +G++ +C C +E ++H F CPFA
Sbjct: 1428 IWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPSCPRCHRENESINHALFTCPFAT 1487
Query: 417 HVW*MTSLWGTIQHALSST---TSAINVIFMLLETFSAELNQRL-TSLFWSLWKHRN 469
W ++ I++ L S + N++ + +T ++ ++ L L W +WK RN
Sbjct: 1488 MAWRLSDS-SLIRNQLMSNDFEENISNILNFVQDTTMSDFHKLLPVWLIWRIWKARN 1543
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 57.8 bits (138), Expect = 2e-08
Identities = 33/117 (28%), Positives = 56/117 (47%), Gaps = 5/117 (4%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IW L + PK K+ +W T RL +G++ C C +E ++H F CPFA
Sbjct: 1202 IWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPICPRCHRENESINHALFTCPFAT 1261
Query: 417 HVW*MTSLWGTIQHALSST---TSAINVIFMLLETFSAELNQRL-TSLFWSLWKHRN 469
W ++ I++ L S + N++ + +T ++ ++ L L W +WK RN
Sbjct: 1262 MAWWLSDS-SLIRNQLMSNDFEENISNILNFVQDTTMSDFHKLLPVWLIWRIWKARN 1317
>At5g18880 SAE1-S9-protein - like
Length = 295
Score = 55.8 bits (133), Expect = 7e-08
Identities = 34/131 (25%), Positives = 60/131 (44%), Gaps = 1/131 (0%)
Query: 348 SPYVGFWSGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSH 407
SP V + +W + P+F + W PTR RL G+ P++ V C + E +H
Sbjct: 122 SPTVPWAKVVWFKEYIPRFSLITWMSFLERLPTRDRLRGWGMNIPSSWVLCSNGDETHAH 181
Query: 408 IFFECPFAIHVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAE-LNQRLTSLFWSLWK 466
+FFEC F++ +W + ++ ++ + L + S L L S + +WK
Sbjct: 182 LFFECSFSLAIWEFFASKFRPSPPFGLPAASSWILQLPLRSHSTTILKLLLQSAVYHVWK 241
Query: 467 HRNLRAWEDVT 477
RN R + ++
Sbjct: 242 ERNARIFTSIS 252
>At3g25270 hypothetical protein
Length = 343
Score = 55.8 bits (133), Expect = 7e-08
Identities = 38/119 (31%), Positives = 52/119 (42%), Gaps = 10/119 (8%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IWKLK PK K+ +W + G T L + I+ C C E H+FF+C +A
Sbjct: 18 IWKLKTAPKIKHFLWKLLSGALATGDNLKRRHIRNHPQCHRCCQEDETSQHLFFDCFYAQ 77
Query: 417 HVW*MTSLWGTIQHALSSTTSAI--NVIFMLLETFSAELNQRLTSL----FWSLWKHRN 469
VW + I H TT + +LL + A +L +L W LWK RN
Sbjct: 78 QVWRASG----IPHQELRTTGITMETKMELLLSSCLANRQPQLFNLAIWILWRLWKSRN 132
>At2g40680 putative non-LTR retroelement reverse transcriptase
Length = 296
Score = 53.1 bits (126), Expect = 4e-07
Identities = 33/125 (26%), Positives = 52/125 (41%), Gaps = 18/125 (14%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IW + PKF + W T R+ Q T C C E +H+FF+CP++
Sbjct: 132 IWFSEATPKFFFITWLAIHDRLTTGARMRSWNTQVDTTCKFCAEPVETRNHLFFQCPYST 191
Query: 417 HVW----------*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSLWK 466
VW TS W I L+S++ + +L TF +++ ++W+
Sbjct: 192 QVWEKLMKGLLQGHYTSSWDRIVTLLTSSSLGRKHLLLLRYTFQIKVH--------TIWR 243
Query: 467 HRNLR 471
RN R
Sbjct: 244 ERNNR 248
>At3g31420 hypothetical protein
Length = 1491
Score = 52.8 bits (125), Expect = 6e-07
Identities = 31/119 (26%), Positives = 50/119 (41%), Gaps = 1/119 (0%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
+WK K PK K+ W + G T L + I + C C + E H+FFEC +A
Sbjct: 1168 LWKTKTAPKIKHFCWKILSGAIATGEMLRYRHINKQSICKRCCRDEETSQHLFFECDYAK 1227
Query: 417 HVW*MTSLWGTI-QHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSLWKHRNLRAWE 474
W L I Q ++ + +F + + Q + W LW+ RN+ ++
Sbjct: 1228 ATWRGAGLPNLIFQDSIVTLEEKFRAMFTFNPSTTNYWRQLPFWILWRLWRSRNILTFQ 1286
>At2g23880 putative non-LTR retroelement reverse transcriptase
Length = 1216
Score = 52.0 bits (123), Expect = 1e-06
Identities = 31/126 (24%), Positives = 50/126 (39%), Gaps = 18/126 (14%)
Query: 356 GIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFA 415
G+W PKF W R T R++ PT CV C S E H+FF+C ++
Sbjct: 781 GVWFAHATPKFSFCAWLAIRNRLSTGDRMMTWNNGTPTTCVFCSSPMETRDHLFFQCCYS 840
Query: 416 IHVW----------*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSLW 465
+W ++ W + + +S + F+ TF ++ S+W
Sbjct: 841 SEIWTSIAKNVYKDRFSTKWSAVVNYISDSQPDRIQSFLSRYTFQVSIH--------SIW 892
Query: 466 KHRNLR 471
+ RN R
Sbjct: 893 RERNSR 898
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 52.0 bits (123), Expect = 1e-06
Identities = 28/88 (31%), Positives = 44/88 (49%), Gaps = 1/88 (1%)
Query: 355 SGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPF 414
S +WKL+ PK K+L+W P ++L+ + I C C + E +H+FF C F
Sbjct: 932 SNLWKLQTLPKIKHLMWKAAMEALPVGIQLVRRHISPSAACHRCGA-PESTTHLFFHCEF 990
Query: 415 AIHVW*MTSLWGTIQHALSSTTSAINVI 442
A VW + L T SS A++++
Sbjct: 991 AAQVWELAPLQETTVPPGSSMLDALSLL 1018
>At3g28160 non-LTR reverse transcriptase, putative
Length = 630
Score = 51.6 bits (122), Expect = 1e-06
Identities = 34/129 (26%), Positives = 48/129 (36%), Gaps = 18/129 (13%)
Query: 353 FWSGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFEC 412
++ G+W PK+ + W T RL + CV C E H+FF C
Sbjct: 279 WYRGVWFPSSTPKYSFVTWLAFHNRLATGDRLYKWNSEARATCVFCDEELETRDHLFFSC 338
Query: 413 PFAIHVW*M----------TSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFW 462
P++ +W S W I L ++ +F L TF A L
Sbjct: 339 PYSSQIWIALAKGLLNGRNVSSWSLITPHLLDSSQPYLHVFTLRYTFQA--------LIH 390
Query: 463 SLWKHRNLR 471
SLW+ RN R
Sbjct: 391 SLWRERNGR 399
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.364 0.161 0.683
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,176,049
Number of Sequences: 26719
Number of extensions: 450880
Number of successful extensions: 1994
Number of sequences better than 10.0: 69
Number of HSP's better than 10.0 without gapping: 61
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 1873
Number of HSP's gapped (non-prelim): 83
length of query: 618
length of database: 11,318,596
effective HSP length: 105
effective length of query: 513
effective length of database: 8,513,101
effective search space: 4367220813
effective search space used: 4367220813
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 63 (28.9 bits)
Medicago: description of AC147010.4


