

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147002.8 + phase: 0 /pseudo
(1664 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
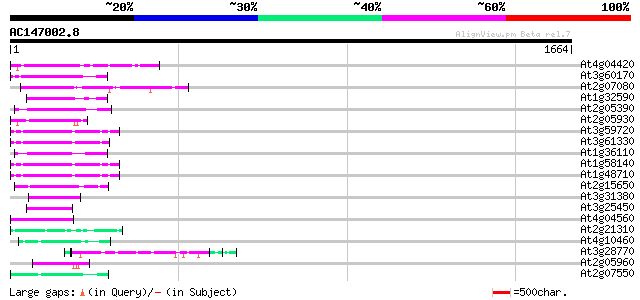
Score E
Sequences producing significant alignments: (bits) Value
At4g04420 putative transposon protein 117 7e-26
At3g60170 putative protein 105 3e-22
At2g07080 putative gag-protease polyprotein 99 2e-20
At1g32590 hypothetical protein, 5' partial 96 2e-19
At2g05390 putative retroelement pol polyprotein 87 8e-17
At2g05930 copia-like retroelement pol polyprotein 86 2e-16
At3g59720 copia-type reverse transcriptase-like protein 85 4e-16
At3g61330 copia-type polyprotein 84 7e-16
At1g36110 hypothetical protein 84 9e-16
At1g58140 hypothetical protein 83 1e-15
At1g48710 hypothetical protein 83 1e-15
At2g15650 putative retroelement pol polyprotein 75 2e-13
At3g31380 hypothetical protein 70 1e-11
At3g25450 hypothetical protein 69 2e-11
At4g04560 putative transposon protein 69 3e-11
At2g21310 putative retroelement pol polyprotein 67 9e-11
At4g10460 putative retrotransposon 61 5e-09
At3g28770 hypothetical protein 59 2e-08
At2g05960 putative retroelement pol polyprotein 59 2e-08
At2g07550 putative retroelement pol polyprotein 59 2e-08
>At4g04420 putative transposon protein
Length = 1008
Score = 117 bits (292), Expect = 7e-26
Identities = 115/462 (24%), Positives = 200/462 (42%), Gaps = 57/462 (12%)
Query: 1 WKTNMYSFIMGLDEELWDIL-----------EDGVDDLDLDEEGAAIDRKIHTPAQKKLY 49
WK M + I GL +E W EDG D L ++ A++
Sbjct: 24 WKVKMRALIRGLGKEAWIATSIGWKAPVIKGEDGEDVLKTKDQW--------NDAEEAKA 75
Query: 50 KKHHKIRGIIVASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELF 109
K + + +I + + ++ ++ + +AK + L +EG+ VK ++ ML Q+E
Sbjct: 76 KANSRALSLIFNFVNQNQFKRIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFENL 135
Query: 110 RMKDDESIEEMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLN 169
M++ E+IEE + + S L K Y V K+LR LPSR+ K TA+ + D +
Sbjct: 136 SMEETENIEEFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAMGTSLDTD 195
Query: 170 TLSVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSD 229
++ E++V L+ +E+ + + SK +AL + K ++ ++E D S
Sbjct: 196 SIDFEEVVGMLQAYELEITSGK-GGYSKGLALAASAKKNE----------IQELKDTMSM 244
Query: 230 EDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFK----KEDQKGCFNCKKPGHFIADCP 285
+ AM R +KK + ++ K D+ C C+ GH A+CP
Sbjct: 245 MAKDFSRAM--------RRVEKKGFGRNQGTDRYRDRSSKRDEIQCHECQGYGHIKAECP 296
Query: 286 DLQKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVAT 345
L+++ K S+ + +KF KS +SES S+ +++D K V V
Sbjct: 297 SLKRKDLK-CSECNGLGHTKFDCVGSKSKPDKSCSSESESDSNDGDSEDYIKGFVSFVGI 355
Query: 346 V-SSEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQ 404
+ + S++E+D EDE+ E D K++ + E L + ++ L K+
Sbjct: 356 IEEKDESSDSEADGEDEDN-----SADEDSDIEKDV-----NINEEFRKLYDSWLMLSKE 405
Query: 405 QKSTLLE-LKASE--EELKGFNLISATYEDRLKSLCQKLQEK 443
+ + L E LK E E+LKG + L C +EK
Sbjct: 406 KVAWLEEKLKVQELTEKLKGELTAANQKNSELIQKCSVAEEK 447
>At3g60170 putative protein
Length = 1339
Score = 105 bits (261), Expect = 3e-22
Identities = 75/289 (25%), Positives = 129/289 (43%), Gaps = 40/289 (13%)
Query: 1 WKTNMYSFIMGLDEELWDILEDGVDDLDLDEEGAAIDRKIHTPAQKKLYKKHHKIRGIIV 60
W M +F+ ELW ++E+G+ + + G + A ++ K K++ +
Sbjct: 22 WSMTMENFLRS--RELWRLVEEGIPAIVV---GTTPVSEAQRSAVEEAKLKDLKVKNFLF 76
Query: 61 ASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIEEM 120
+I R + DKST+KA++ S+ ++GS KVK A+ L ++EL MK+ E I+
Sbjct: 77 QAIDREILETILDKSTSKAIWESMKKKYQGSTKVKRAQLQALRKEFELLAMKEGEKIDTF 136
Query: 121 YSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSL 180
R T+V+ ++ + S VSKILRSL ++ V +IEE+ DL+TLS+++L SL
Sbjct: 137 LGRTLTVVNKMKTNGEVMEQSTIVSKILRSLTPKFNYVVCSIEESNDLSTLSIDELHGSL 196
Query: 181 KVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLS 240
VHE LN H +++ + + + ++ S
Sbjct: 197 LVHEQRLNGHVQEEQALKVTHEERPSQGRGRGVFRGSRG--------------------- 235
Query: 241 NKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQK 289
RG ++ C+ C GHF +CP+ +K
Sbjct: 236 --------------RGRGRGRSGTNRAIVECYKCHNLGHFQYECPEWEK 270
>At2g07080 putative gag-protease polyprotein
Length = 627
Score = 99.4 bits (246), Expect = 2e-20
Identities = 119/517 (23%), Positives = 227/517 (43%), Gaps = 51/517 (9%)
Query: 31 EEGAAIDR-KIHTPAQKKLYKKHH-KIRGIIVASIPRTEYMKMSDKSTAKAMFASLCANF 88
E+G I + K + A++KL K + + I + E+ + +AK + +L +
Sbjct: 36 EDGFKITKPKANWTAEEKLQSKFNARAMKAIFNGVDEDEFKLIQGCKSAKQAWDTLQKSH 95
Query: 89 EGSKKVKEAKALMLVHQYELFRMKDDESIEEMYSRFQTLVSGLQILKKSYVSSDHVSKIL 148
EG+ VK + + Q+E +M+ DE I + S+ L + +++ K+Y V K+L
Sbjct: 96 EGTSSVKRTRLDHIATQFEYLKMEPDEKIVKFSSKISALANEAEVMGKTYKDQKLVKKLL 155
Query: 149 RSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTS 208
R LP ++ + A + + +S DLV LK+ EM ++ + K SK+IA
Sbjct: 156 RCLPPKFAAHKAVMRVAGNTDKISFVDLVGMLKLEEMKADQDKV-KPSKNIAF------- 207
Query: 209 KSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQ 268
A + SE +E DG + ++ A+ +++ R + +F R + +K+ +
Sbjct: 208 ---NADQGSEQFQEIKDGMALLARNFGKALKRVEIDG-ERSRGRF--SRSENDDLRKKKE 261
Query: 269 KGCFNCKKPGHFIADCPDLQKEKFK-------GKSKKSSFNSSKFRKQIKKSLMATWEDL 321
C+ C GH +CP ++++ K G +K N SK + +KSL+
Sbjct: 262 IQCYECGGFGHIKPECPITKRKEMKCLKCKGVGHTKFECPNKSKLK---EKSLI------ 312
Query: 322 DSESGSDKEEADDDAKAAVGLVATVSSEAVSE--AESDSEDENEVYSKIPRQELVDSLKE 379
S SD E+DD+ + + VA ++S S+ +++DS+ + E+ K + L DS +
Sbjct: 313 ---SFSD-SESDDEGEELLNFVAFMASSDSSKFMSDTDSDCDEELNPKDKYRVLYDSWVQ 368
Query: 380 LLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKAS-------EEELKGFNLISATYEDR 432
L E L+ K ++ + K L + +++L DR
Sbjct: 369 LSKDKLKLVKEKLTLEAKLANVSTEDKQKLSGITVDGNSQDYYQKKLDCLQKECHRERDR 428
Query: 433 LKSLCQKLQEKCDK-GSGNKHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKS 491
K L ++L +K + NK +LD + G S+ + Y Y K K +E+
Sbjct: 429 TKLLERELNDKYKQIRMLNKGLESLDKILAMGRTDSQQRGLGYQGYTGKIK-----KEEG 483
Query: 492 KEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEA 528
+ + S + ++ +++ K+ V+ K E+
Sbjct: 484 RVINFVSGGSTSETVVRQSYIEPKKQVKSHVEIKRES 520
>At1g32590 hypothetical protein, 5' partial
Length = 1263
Score = 95.5 bits (236), Expect = 2e-19
Identities = 67/244 (27%), Positives = 110/244 (44%), Gaps = 40/244 (16%)
Query: 51 KHHKIRGIIVASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFR 110
K HK++ + ASI +T + K T+K ++ S+ ++G+ +V+ A+ L +E+
Sbjct: 24 KDHKVKNYLFASIDKTILKTILQKETSKDLWESMKRKYQGNDRVQSAQLQRLRRSFEVLE 83
Query: 111 MKDDESIEEMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNT 170
MK E+I +SR + + ++ L + S V KILR+L ++ V AIEE+ ++
Sbjct: 84 MKIGETITGYFSRVMEITNDMRNLGEDMPDSKVVEKILRTLVEKFTYVVCAIEESNNIKE 143
Query: 171 LSVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDE 230
L+V+ L SSL VHE +L+ H+ + + KA + PDG
Sbjct: 144 LTVDGLQSSLMVHEQNLSRHDVEE-----------------RVLKA--ETQWRPDGGRGR 184
Query: 231 DQSVKMAMLSNKLEYLARKQKKFLSKRGSY----KNFKKEDQKGCFNCKKPGHFIADCPD 286
S RG Y + + D CF C K GH+ A+CP
Sbjct: 185 GGSPSRG-----------------RGRGGYQGRGRGYVNRDTVECFKCHKMGHYKAECPS 227
Query: 287 LQKE 290
+KE
Sbjct: 228 WEKE 231
>At2g05390 putative retroelement pol polyprotein
Length = 1307
Score = 87.0 bits (214), Expect = 8e-17
Identities = 61/288 (21%), Positives = 127/288 (43%), Gaps = 54/288 (18%)
Query: 15 ELWDILEDGVDDLDLDEEGAAIDRKIHTPAQKKLYKKHHKIRGIIVASIPRTEYMKMSDK 74
++W+ ++ G DD++ K+ R ++ S+P + +++
Sbjct: 9 KVWETIDPGSDDME----------------------KNDMARALLFQSVPESTILQVGKH 46
Query: 75 STAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIEEMYSRFQTLVSGLQIL 134
T+KAM+ ++ G+++VKEAK L+ +++ MKD+E+I+E R + + + L
Sbjct: 47 KTSKAMWEAIKTRNLGAERVKEAKLQTLMAEFDRLNMKDNETIDEFVGRISEISTKSESL 106
Query: 135 KKSYVSSDHVSKILRSLP-SRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETS 193
+ S V K L+SLP ++ + A+E+ DLNT ED+V +K +E + + + S
Sbjct: 107 GEEIEESKIVKKFLKSLPRKKYIHIIAALEQILDLNTTGFEDIVGRMKTYEDRVCDEDDS 166
Query: 194 KKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKF 253
+ + + + ++S ++ + S G
Sbjct: 167 PEEQGKLMYANSESSYDTRGGRGRGRGRSSGRG--------------------------- 199
Query: 254 LSKRGSYKNFKKEDQKG-CFNCKKPGHFIADCPDLQKEKFKGKSKKSS 300
RG Y +++ K C+ C K GH+ ++C D + K + ++ +
Sbjct: 200 ---RGGYGYQQRDKSKVICYRCDKTGHYASECLDRLLKLIKAQEQQQN 244
>At2g05930 copia-like retroelement pol polyprotein
Length = 916
Score = 85.9 bits (211), Expect = 2e-16
Identities = 68/265 (25%), Positives = 115/265 (42%), Gaps = 47/265 (17%)
Query: 1 WKTNMYSFIMGLDEELWDIL-----------EDGVDDLDLDEEGAAIDRKIHTPAQKKLY 49
WK M + I GL +E W EDG D L +++ + T + L
Sbjct: 36 WKVKMRALIRGLGKEAWIATSIGWKAPVIKGEDGEDVLKTEDQWNDAEEAKATANSRAL- 94
Query: 50 KKHHKIRGIIVASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELF 109
+I S+ + ++ ++ + +AK + L +EG+ VK ++ ML Q+E
Sbjct: 95 -------SLIFNSVNQNQFKQIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFENL 147
Query: 110 RMKDDESIEEMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLN 169
M++ E+IEE + + S L K Y V K+LR LPSR+ K TA+ + D N
Sbjct: 148 TMEETENIEEFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAMGTSLDTN 207
Query: 170 TLSVEDLVSSLKVHEMSLNEHE----------TSKKSKSI--------------ALPSKG 205
++ E++V + +E+ + + S K K + + SK
Sbjct: 208 SIDFEEVVGMFQAYELEITSGKGGYGHIKAECPSLKRKDLKCSECKGLGHIKFDCVGSKS 267
Query: 206 KTSKSSKAYKASESVEESPDGDSDE 230
K +S +SES +S DGDS++
Sbjct: 268 KPDRSC----SSESESDSNDGDSED 288
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 84.7 bits (208), Expect = 4e-16
Identities = 72/327 (22%), Positives = 147/327 (44%), Gaps = 20/327 (6%)
Query: 1 WKTNMYSFIMGLDEELWDILEDGVDDLDLDEEGAAIDRKIHTPAQKKLYKKHHKIRGIIV 60
W M + + D +W+I+E G ++ + EG+ + + K+ K +I
Sbjct: 21 WSLRMKAILGAHD--VWEIVEKGF--IEPENEGSL--SQTQKDGLRDSRKRDKKALCLIY 74
Query: 61 ASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIEEM 120
+ + K+ + ++AK + L +++G+ +VK+ + L ++E +MK+ E + +
Sbjct: 75 QGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDY 134
Query: 121 YSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSL 180
+SR T+ + L+ + + K+LRSL ++ VT IEE KDL +++E L+ SL
Sbjct: 135 FSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSL 194
Query: 181 KVHEMSLNEHETSKKSKSI---ALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMA 237
+ +E E KK + I L + ++ ++Y+ + G
Sbjct: 195 QAYE------EKKKKKEDIVEQVLNMQITKEENGQSYQRRGGGQVRGRGRGGYGNGRGWR 248
Query: 238 MLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSK 297
+ + K + K K C+NC K GH+ ++C +KFK +
Sbjct: 249 PHEDNTNQRGENSSRGRGKGHPKSRYDKSSVK-CYNCGKFGHYASECKAPSNKKFK---E 304
Query: 298 KSSFNSSKFRKQIKKSLMATWEDLDSE 324
K+++ K +++ LMA+++ + E
Sbjct: 305 KANYVEEKIQEE-DMLLMASYKKDEQE 330
>At3g61330 copia-type polyprotein
Length = 1352
Score = 84.0 bits (206), Expect = 7e-16
Identities = 67/299 (22%), Positives = 133/299 (44%), Gaps = 16/299 (5%)
Query: 1 WKTNMYSFIMGLDEELWDILEDGVDDLDLDEEGAAIDRKIHTPAQKKLYKKHHKIRGIIV 60
W M + + D +W+I+E G ++ + EG+ + + K+ K +I
Sbjct: 21 WSLRMKAILGAHD--VWEIVEKGF--IEPENEGSL--SQTQKDGLRDSRKRDKKALCLIY 74
Query: 61 ASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIEEM 120
+ + K+ + ++AK + L +++G+ +VK+ + L ++E +MK+ E + +
Sbjct: 75 QGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDY 134
Query: 121 YSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSL 180
+SR T+ + L+ + + K+LRSL ++ VT IEE KDL +++E L+ SL
Sbjct: 135 FSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSL 194
Query: 181 KVHEMSLNEHETSKKSKSIA---LPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMA 237
+ +E E KK + IA L + ++ ++Y+ + G
Sbjct: 195 QAYE------EKKKKKEDIAEQVLNMQITKEENGQSYQRRGGGQVRGRGRGGYGNGRGWR 248
Query: 238 MLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKS 296
+ + K + K K C+NC K GH+ ++C +KF+ K+
Sbjct: 249 PHEDNTNQRGENSSRGRGKGHPKSRYDKSSVK-CYNCGKFGHYASECKAPSNKKFEEKA 306
>At1g36110 hypothetical protein
Length = 745
Score = 83.6 bits (205), Expect = 9e-16
Identities = 68/283 (24%), Positives = 120/283 (42%), Gaps = 66/283 (23%)
Query: 15 ELWDILEDGVDDLDLDEEGAAIDRKIHTPAQKKLYKKHHKIRGIIVASIPRTEYMKMSDK 74
++W+ +E G+DD K+ RG+I SIP + +++
Sbjct: 42 KMWETIEPGIDD----------------------GYKNTMARGLIFQSIPESLTLQVGTL 79
Query: 75 STAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIEEMYSRFQTLVSGLQIL 134
+TAK ++ S+ + G+ +VKEA+ L+ ++E +MK+ E I+ R L + L
Sbjct: 80 ATAKLVWDSIKTRYVGADRVKEARLQTLMAEFEKMKMKESEKIDVFAGRLAELATRSDAL 139
Query: 135 KKSYVSSDHVSKILRSLPSR-WRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETS 193
+ +S V K L +LP R + + ++E+ DLN S ED+V +KV+E
Sbjct: 140 GSNIETSKLVKKFLNALPLRKYIHIIASLEQVLDLNNTSFEDIVGRIKVYE--------- 190
Query: 194 KKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLE---YLARKQ 250
E DG+ ED K+ + + Y +R +
Sbjct: 191 ---------------------------ERVWDGEEQEDDQGKLMYANTDTQDSWYASRGR 223
Query: 251 KK---FLSK-RGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQK 289
+ F + RG + + + C+ C K GH+ ++CPD K
Sbjct: 224 GQGGGFNGRGRGRGRGSRDTSKVTCYRCDKLGHYASNCPDSVK 266
>At1g58140 hypothetical protein
Length = 1320
Score = 83.2 bits (204), Expect = 1e-15
Identities = 71/327 (21%), Positives = 147/327 (44%), Gaps = 20/327 (6%)
Query: 1 WKTNMYSFIMGLDEELWDILEDGVDDLDLDEEGAAIDRKIHTPAQKKLYKKHHKIRGIIV 60
W M + + D +W+I+E G ++ + EG+ + + K+ K +I
Sbjct: 21 WSLRMKAILGAHD--VWEIVEKGF--IEPENEGSL--SQTQKDGLRDSRKRDKKALCLIY 74
Query: 61 ASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIEEM 120
+ + K+ + ++AK + L +++G+ +VK+ + L ++E +MK+ E + +
Sbjct: 75 QGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDY 134
Query: 121 YSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSL 180
+SR T+ + L+ + + K+LRSL ++ VT IEE KDL +++E L+ SL
Sbjct: 135 FSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSL 194
Query: 181 KVHEMSLNEHETSKKSKSI---ALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMA 237
+ +E E KK + I L + ++ ++Y+ + G
Sbjct: 195 QAYE------EKKKKKEDIVEQVLNMQITKEENGQSYQRRGGGQVRGRGRGGYGNGRGWR 248
Query: 238 MLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSK 297
+ + K + K K C+NC K GH+ ++C +KF+ +
Sbjct: 249 PHEDNTNQRGENSSRGRGKGHPKSRYDKSSVK-CYNCGKFGHYASECKAPSNKKFE---E 304
Query: 298 KSSFNSSKFRKQIKKSLMATWEDLDSE 324
K+++ K +++ LMA+++ + E
Sbjct: 305 KANYVEEKIQEE-DMLLMASYKKDEQE 330
>At1g48710 hypothetical protein
Length = 1352
Score = 83.2 bits (204), Expect = 1e-15
Identities = 71/327 (21%), Positives = 147/327 (44%), Gaps = 20/327 (6%)
Query: 1 WKTNMYSFIMGLDEELWDILEDGVDDLDLDEEGAAIDRKIHTPAQKKLYKKHHKIRGIIV 60
W M + + D +W+I+E G ++ + EG+ + + K+ K +I
Sbjct: 21 WSLRMKAILGAHD--VWEIVEKGF--IEPENEGSL--SQTQKDGLRDSRKRDKKALCLIY 74
Query: 61 ASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIEEM 120
+ + K+ + ++AK + L +++G+ +VK+ + L ++E +MK+ E + +
Sbjct: 75 QGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDY 134
Query: 121 YSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSL 180
+SR T+ + L+ + + K+LRSL ++ VT IEE KDL +++E L+ SL
Sbjct: 135 FSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSL 194
Query: 181 KVHEMSLNEHETSKKSKSI---ALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMA 237
+ +E E KK + I L + ++ ++Y+ + G
Sbjct: 195 QAYE------EKKKKKEDIIEQVLNMQITKEENGQSYQRRGGGQVRGRGRGGYGNGRGWR 248
Query: 238 MLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSK 297
+ + K + K K C+NC K GH+ ++C +KF+ +
Sbjct: 249 PHEDNTNQRGENSSRGRGKGHPKSRYDKSSVK-CYNCGKFGHYASECKAPSNKKFE---E 304
Query: 298 KSSFNSSKFRKQIKKSLMATWEDLDSE 324
K+++ K +++ LMA+++ + E
Sbjct: 305 KANYVEEKIQEE-DMLLMASYKKDEQE 330
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 75.5 bits (184), Expect = 2e-13
Identities = 59/277 (21%), Positives = 125/277 (44%), Gaps = 14/277 (5%)
Query: 15 ELWDILEDGVDDLDLDEEGAAIDRKIHTPAQKKLYKKHHKIRGIIVASIPRTEYMKMSDK 74
+LW ++E+GV + E + T ++ + ++ I+ ++ + +++
Sbjct: 32 KLWSVVEEGVPVEPVQAEETPETARAKTLREEAVTNDTMALQ-ILQTAVTDQIFSRIAAA 90
Query: 75 STAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIEEMYSRFQTLVSGLQIL 134
S++K + L ++GS +V+ K L +YE +M D+++I+ + L L
Sbjct: 91 SSSKEAWDVLKDEYQGSPQVRLVKLQSLRREYENLKMYDNDNIKTFTDKLIVLEIQLTYH 150
Query: 135 KKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETSK 194
+ ++ + KIL SLP+++ V+ +E+ +DL+ L++ +L+ LK E + E S
Sbjct: 151 GEKKTNTQLIQKILISLPAKFDSIVSVLEQTRDLDALTMSELLGILKAQEARVTAREEST 210
Query: 195 KSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFL 254
K + + SKG+ ES + + ++ +Q K ++ + ++
Sbjct: 211 KEGAFYVRSKGR-----------ESGFKQDNTNNRVNQDKKWCGFHKSSKHTEEECREKP 259
Query: 255 SKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEK 291
KN K C+ C K GH+ +C KE+
Sbjct: 260 KNDDHGKN--KRSNIKCYKCGKIGHYANECRSKNKER 294
>At3g31380 hypothetical protein
Length = 262
Score = 69.7 bits (169), Expect = 1e-11
Identities = 45/156 (28%), Positives = 78/156 (49%), Gaps = 1/156 (0%)
Query: 56 RGIIVASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDE 115
RG++V SIP +++ + TAK ++ S+ G+++VKEA+ L+ +E +MK+ E
Sbjct: 3 RGLLVQSIPEAFTLQVGNLQTAKEVWDSIKTRHVGAERVKEARVQTLMADFEKMKMKEAE 62
Query: 116 SIEEMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLP-SRWRPKVTAIEEAKDLNTLSVE 174
I++ R L + L + V K L SLP ++ + ++E+ DLN + E
Sbjct: 63 KIDDFAGRLSELSTKSAPLGVNIEVPKLVKKFLNSLPRKKYIHIIASLEQVLDLNNTTFE 122
Query: 175 DLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKS 210
D+V +KV+E + + E L SKS
Sbjct: 123 DIVGCMKVNEERVYDPEEETNEDQNKLMYTSSDSKS 158
>At3g25450 hypothetical protein
Length = 1343
Score = 69.3 bits (168), Expect = 2e-11
Identities = 37/136 (27%), Positives = 76/136 (55%), Gaps = 1/136 (0%)
Query: 50 KKHHKIRGIIVASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELF 109
+K+ R ++ SIP + +++ + T+ A++ ++ + G+++VKEA+ L+ +++
Sbjct: 58 EKNDMARALLFQSIPESLILQVGKQKTSSAVWEAIKSRNLGAERVKEARLQTLMAEFDKL 117
Query: 110 RMKDDESIEEMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLP-SRWRPKVTAIEEAKDL 168
+MKD E+I++ R + + L + S V K L+SLP ++ V A+E+ DL
Sbjct: 118 KMKDSETIDDYVGRISEITTKAAALGEDIEESKIVKKFLKSLPRKKYIHIVAALEQVLDL 177
Query: 169 NTLSVEDLVSSLKVHE 184
T + ED+ +K +E
Sbjct: 178 KTTTFEDIAGRIKTYE 193
>At4g04560 putative transposon protein
Length = 590
Score = 68.6 bits (166), Expect = 3e-11
Identities = 45/192 (23%), Positives = 91/192 (46%), Gaps = 3/192 (1%)
Query: 1 WKTNMYSFIMGLDEELWDILEDGVDDLDLDEEGAAIDRKIHTP--AQKKLYKKHH-KIRG 57
WK + I +D + W +EDG + I K T A +K H+ +
Sbjct: 356 WKVLIKRSIQSIDMDAWFAVEDGWMPPTTKDAKRDIVSKSRTEWIADEKTAANHNSQALS 415
Query: 58 IIVASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESI 117
+I S+ R ++ ++ +AK ++ L +FE + VK + ML ++E M+ +ES+
Sbjct: 416 VIFGSLLRNKFTQVQGCLSAKEVWEILQVSFECTNNVKRTRLDMLASEFENLTMEAEESV 475
Query: 118 EEMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLV 177
++ + ++ +L K+Y V K LRSLP +++ +AI+ + + + L + +V
Sbjct: 476 DDFNGKLSSITQEAVVLGKTYKDKKMVKKFLRSLPDKFQSHKSAIDVSLNSDQLKFDQVV 535
Query: 178 SSLKVHEMSLNE 189
++ ++ E
Sbjct: 536 GMMQAYDTDKEE 547
Score = 36.6 bits (83), Expect = 0.13
Identities = 36/174 (20%), Positives = 74/174 (41%), Gaps = 29/174 (16%)
Query: 1 WKTNMYSFIMGLDEELWDILEDGVDD--LDLDEEGAAIDRKIHTPAQKKLYKKHHKIRGI 58
W+ M + ++LW+I+++GV + D A+ +K A
Sbjct: 18 WRIKMKTIFQ--TKKLWEIVDEGVPKPPAEGDHSPEAVQQKTRCEA-------------- 61
Query: 59 IVASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIE 118
AS+ +++ + + ++F + + AL +YE +MK+ ++I
Sbjct: 62 --ASLKDLTALQILQTAVSDSIFPRIAP---------ASSALGKPWEYENLKMKESDNIN 110
Query: 119 EMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLS 172
++ + + L++ + V KIL SLP R+ V +++ KDL +LS
Sbjct: 111 TFMTKLIEMGNQLRVHGEEKSDYQIVQKILISLPKRFDIIVAMMKQTKDLTSLS 164
Score = 31.2 bits (69), Expect = 5.3
Identities = 35/162 (21%), Positives = 73/162 (44%), Gaps = 18/162 (11%)
Query: 128 VSGLQILKKSYVSS--DHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVE--DLVSSLKVH 183
++ LQIL+ + S ++ +L W + ++E+ ++NT + ++ + L+VH
Sbjct: 67 LTALQILQTAVSDSIFPRIAPASSALGKPWEYENLKMKESDNINTFMTKLIEMGNQLRVH 126
Query: 184 EMSLNEHETSKKSKSIALPSKGKTSKSS-KAYKASESVEESPDGDSDEDQSVKMAMLSNK 242
++++ +K I+LP + + K K S+ D E ++ N+
Sbjct: 127 GEEKSDYQIVQKIL-ISLPKRFDIIVAMMKQTKDLTSLSAGKWCDVCERKN------HNE 179
Query: 243 LEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADC 284
+ +K K LS++ +++ CF C KPGH +C
Sbjct: 180 SDCWMKKNKGVLSQQVG------NNERRCFVCNKPGHLAKNC 215
>At2g21310 putative retroelement pol polyprotein
Length = 838
Score = 67.0 bits (162), Expect = 9e-11
Identities = 76/337 (22%), Positives = 136/337 (39%), Gaps = 54/337 (16%)
Query: 1 WKTNMYSFIMGLDEELWDILEDGVDDLDLDEEG--AAIDRKIHTPAQKKLYKKHHKIRGI 58
WK + + I EL +LE +D ++EE A D + K L +K K R
Sbjct: 20 WKEKLLAHI-----ELLGLLEGLEEDEAIEEEESTAETDSLLTKTEDKVLKEKRGKARST 74
Query: 59 IVASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIE 118
++ S+ K+ + TA M L F ++ Y +M D +IE
Sbjct: 75 VILSLGNHVLRKVIKEKTAAGMIRVLDKLFMAKSLPNRIYLKQRLYGY---KMSDSMTIE 131
Query: 119 EMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVS 178
E + F L+S L+ +K S D +L SLP ++ ++ K TL+++++
Sbjct: 132 ENVNDFFKLISDLENVKVSVPDEDQAIVLLMSLPKQFDQLKDTLKYGK--TTLALDEITG 189
Query: 179 SLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAM 238
+++ SK + L + GK K+S S+++ G S+
Sbjct: 190 AIR--------------SKVLELGASGKMLKNS-----SDALFVQDRGRSE--------- 221
Query: 239 LSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKK 298
K+ K + S K ++K C+ C K GHF C +++ KG + +
Sbjct: 222 ----------KRDKSSERNKSQSRSKSREKKVCWVCGKEGHFKKQCYVWKEKNKKGNNSE 271
Query: 299 SSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDD 335
+S+ + + +A E ES +D +E D++
Sbjct: 272 KGESSNVIGQAADAAALAVRE----ESNADNQEVDNE 304
>At4g10460 putative retrotransposon
Length = 1230
Score = 61.2 bits (147), Expect = 5e-09
Identities = 60/279 (21%), Positives = 112/279 (39%), Gaps = 51/279 (18%)
Query: 25 DDLDLDEEGAAIDRKIHTPAQKKLYKKHHKIRGIIVASIPRTEYMKMSDKSTAKAMFASL 84
D L+ +EEG ++ + + +K K R IV S+ K + TA +M +L
Sbjct: 45 DPLESEEEGKESEKG---DKEALMEEKRQKARSTIVLSVSDQVLRKSKKEKTAPSMLEAL 101
Query: 85 CANFEGSKKVKEAKAL----MLVHQYELFRMKDDESIEEMYSRFQTLVSGLQILKKSYVS 140
K+ +KAL L + ++M+++ S+E F L++ L+
Sbjct: 102 -------DKLYMSKALPNRIYLKQKLYSYKMQENLSVEGNIDEFLRLIADLENTNVLVSD 154
Query: 141 SDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKSKSIA 200
D +L SLP ++ ++ TLSV+++V+++ E+ L ++ S + ++
Sbjct: 155 EDQAILLLMSLPKQFDQLKDTLKYGSGRTTLSVDEVVAAIYSKELELGSNKKSIRGQAEG 214
Query: 201 LPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSY 260
L K K + E+ G+ +S
Sbjct: 215 LYVKDKPETRGMS-------EQKEKGNKGRSRS--------------------------- 240
Query: 261 KNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKS 299
+ + KGC+ C + GHF CP+ K++ KGK + S
Sbjct: 241 ---RSKGWKGCWICGEEGHFKTSCPNKGKQQNKGKDQAS 276
>At3g28770 hypothetical protein
Length = 2081
Score = 59.3 bits (142), Expect = 2e-08
Identities = 95/416 (22%), Positives = 183/416 (43%), Gaps = 37/416 (8%)
Query: 184 EMSLNEHETSK-KSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNK 242
E +E SK + K K KT + +K K ++ + DS+E +S K S
Sbjct: 998 EKKESEDSASKNREKKEYEEKKSKTKEEAKKEKKKSQDKKREEKDSEERKSKKEKEESRD 1057
Query: 243 LEYLARKQKKFLSKRGS--YKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKSS 300
L+ +K+++ K+ S +K+ KKED+K D ++KE+ K + KK
Sbjct: 1058 LK-AKKKEEETKEKKESENHKSKKKEDKKE----------HEDNKSMKKEEDKKEKKKHE 1106
Query: 301 FNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAA--VGLVATVSSEAVSEAESDS 358
+ S+ +++ KK + E L+ ++ + K+E ++ K + V LV S + + +
Sbjct: 1107 ESKSRKKEEDKKDM----EKLEDQNSNKKKEDKNEKKKSQHVKLVKKESDKKEKKENEEK 1162
Query: 359 EDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEE 418
+ E+ S ++ VD KE S + + + ++KE +K+ + + + EE
Sbjct: 1163 SETKEIESSKSQKNEVDK-KEKKSSKDQQKKKEKEMKESEEKKLKKNEEDRKKQTSVEEN 1221
Query: 419 LKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDRSKVASMIYSTYK 478
K T +++ K K + K SG K E + A + A+ + +
Sbjct: 1222 KKQ----KETKKEKNKPKDDK--KNTTKQSGGKKESMESESKEAENQQKSQATTQADSDE 1275
Query: 479 NKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEASGSQAKITSK 538
+K + + ++ ++ +S S D D K+ + + + T ++ E + K TS
Sbjct: 1276 SKNEILMQADSQADSHS-DSQAD--SDESKNEILMQADSQATTQRNNEE---DRKKQTSV 1329
Query: 539 PENLKIKVM--TKSDPKSQKIKILKRSEPVHQNLIKPESKIPKQKDQKNKAATASE 592
EN K K K+ PK K K+S +++ + ESK + QK++A T ++
Sbjct: 1330 AENKKQKETKEEKNKPKDDKKNTTKQSGGKKESM-ESESK-EAENQQKSQATTQAD 1383
Score = 47.0 bits (110), Expect = 9e-05
Identities = 98/480 (20%), Positives = 190/480 (39%), Gaps = 59/480 (12%)
Query: 181 KVHEMSLNEHETSKKSKSIALPSKGKT-------SKSSKAYKASESVEESPDGDSDEDQS 233
K E E +K + + + KGK SK+S K E +E + + + +
Sbjct: 917 KKDEKKEGNKEENKDTINTSSKQKGKDKKKKKKESKNSNMKKKEEDKKEYVNNELKKQED 976
Query: 234 VKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFK 293
K ++ L + K K+ S + K +K + KK + +KEK K
Sbjct: 977 NKKETTKSENSKLKEENKDNKEKKESEDSASKNREKKEYEEKKS----KTKEEAKKEKKK 1032
Query: 294 GKSKKSSFNSS---KFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEA 350
+ KK S K +K+ ++S + + E+ KE + +K +++
Sbjct: 1033 SQDKKREEKDSEERKSKKEKEESRDLKAKKKEEETKEKKESENHKSKKKEDKKEHEDNKS 1092
Query: 351 VSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTD-----LKEKYVDLMKQQ 405
+ + E E + SK ++E + K++ L + +N+ + K ++V L+K++
Sbjct: 1093 MKKEEDKKEKKKHEESKSRKKE--EDKKDMEKLEDQNSNKKKEDKNEKKKSQHVKLVKKE 1150
Query: 406 KSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGI- 464
S E K +EE+ + + S+ + K+ K ++K K K E + + +
Sbjct: 1151 -SDKKEKKENEEKSETKEIESSKSQ---KNEVDKKEKKSSKDQQKKKEKEMKESEEKKLK 1206
Query: 465 ----DRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKT 520
DR K S+ EE K+ K + KD K+T G ++
Sbjct: 1207 KNEEDRKKQTSV---------------EENKKQKETKKEKNKPKDDKKNTTKQSGGKKES 1251
Query: 521 AVQSKPEASG---SQAKITSKPENLKIKVMTKSDPK--SQKIKILKRSEPVHQNLIKPES 575
EA SQA + + K +++ ++D + S E ++ L++ +S
Sbjct: 1252 MESESKEAENQQKSQATTQADSDESKNEILMQADSQADSHSDSQADSDESKNEILMQADS 1311
Query: 576 KIPKQK----DQKNKAATASEK--TIPKGVKPKVLNDQKPLSIHPKVQGRKSKTSKANPK 629
+ Q+ D+K + + A K K K K +D+K + K G K ++ ++ K
Sbjct: 1312 QATTQRNNEEDRKKQTSVAENKKQKETKEEKNKPKDDKKNTT---KQSGGKKESMESESK 1368
Score = 44.7 bits (104), Expect = 5e-04
Identities = 108/562 (19%), Positives = 210/562 (37%), Gaps = 94/562 (16%)
Query: 164 EAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEES 223
E K+L+ + ++ + K E++ N+ + +K+ + + G++ K+ E+ E+
Sbjct: 553 EDKNLDNIGADEQKKNDKSVEVTTNDGDHTKEKREETQGNNGESVKNENL----ENKEDK 608
Query: 224 PDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIAD 283
+ DE K +N L K+++ + N K D KG
Sbjct: 609 KELKDDESVGAK----TNNETSLEEKREQTQKGHDNSINSKIVDNKG------------G 652
Query: 284 CPDLQKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLV 343
D KEK ++ N+ + ++ K + D SE G + +E + D+ L
Sbjct: 653 NADSNKEKEVHVGDSTNDNNMESKEDTKSEVEVKKNDGSSEKGEEGKENNKDSMEDKKLE 712
Query: 344 ATVSSEAVSEAES--DSEDENEVYSKIPRQ----ELVDSLKELLSLFEHRTNELTDLKEK 397
S + +S D ++E ++Y + E KE + +TNE ++ K
Sbjct: 713 NKESQTDSKDDKSVDDKQEEAQIYGGESKDDKSVEAKGKKKESKENKKTKTNE-NRVRNK 771
Query: 398 YVDLMKQQKSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALD 457
++ +K + K ++E K + +L S + + K G NK +
Sbjct: 772 EENVQGNKKESEKVEKGEKKESKDAKSVETKDNKKLSSTENRDEAKERSGEDNKED---- 827
Query: 458 DFIMAGIDRSKVASMIYSTYKNKGKG----IGYSEEK-------------SKEYSLKSYC 500
+ SK + + KN+ G +G E+ +KE S+K
Sbjct: 828 ------KEESKDYQSVEAKEKNENGGVDTNVGNKEDSKDLKDDRSVEVKANKEESMKKKR 881
Query: 501 DCIKDGLKSTFVP---------------EGTNAKTAVQSKPEASGSQAK---ITSKPENL 542
+ ++ KS+ G + K K E + + K TS +
Sbjct: 882 EEVQRNDKSSTKEVRDFANNMDIDVQKGSGESVKYKKDEKKEGNKEENKDTINTSSKQKG 941
Query: 543 KIKVMTKSDPKSQKIK------------ILKRSEPVHQNLIKPESKIPKQKDQKNKAATA 590
K K K + K+ +K LK+ E + K E+ K++++ NK
Sbjct: 942 KDKKKKKKESKNSNMKKKEEDKKEYVNNELKKQEDNKKETTKSENSKLKEENKDNKEKKE 1001
Query: 591 SEKTIPKGVKPKVLNDQKPLSIHPKVQGRKSKTSKANPKGPMKIWVPKSELAKNAGVLKG 650
SE + K + K ++K + K + +K K + K K SE K+ K
Sbjct: 1002 SEDSASKNREKKEYEEKKSKT---KEEAKKEKKKSQDKKREEK----DSEERKSK---KE 1051
Query: 651 KRETKVMVPRQRMFKAHDWRES 672
K E++ + +++ + + +ES
Sbjct: 1052 KEESRDLKAKKKEEETKEKKES 1073
>At2g05960 putative retroelement pol polyprotein
Length = 1200
Score = 59.3 bits (142), Expect = 2e-08
Identities = 49/199 (24%), Positives = 92/199 (45%), Gaps = 31/199 (15%)
Query: 69 MKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIEEMYSRFQTLV 128
+++ T+KA++ + + G+++VKEAK L+ +++ +MKD+E+I+E R +
Sbjct: 35 LQVGKLKTSKAVWDKIQSRNLGAERVKEAKLKTLMAEFDKLKMKDNETIDECAGRLSEIS 94
Query: 129 SGLQILKKSYVSSDHVSKILRSLPS-RWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSL 187
+ L + + V K L+SLP+ ++ V A+E+ DL + +D+V +K +E +
Sbjct: 95 TKSTSLGEDIEETKVVKKFLKSLPTKKYIHIVAALEQVLDLKNTTFKDIVGRIKTYEDKI 154
Query: 188 --------NEHETSKK-------------SKSIALP---------SKGKTSKSSKAYKAS 217
E E +K S+ I L + + K K
Sbjct: 155 WVLITCLKKEAEEEEKSVVGVEAEEELVISREITLRLLVIAVINYASNCPDRLLKLIKLQ 214
Query: 218 ESVEESPDGDSDEDQSVKM 236
E +E+ D D DE +S+ M
Sbjct: 215 ERQQEAEDDDDDEVESLMM 233
>At2g07550 putative retroelement pol polyprotein
Length = 1356
Score = 58.9 bits (141), Expect = 2e-08
Identities = 63/299 (21%), Positives = 118/299 (39%), Gaps = 54/299 (18%)
Query: 1 WKTNMYSF--IMGLDEELWDILEDGVDDLDLDEEGAAIDRKIHTPAQKKLYKKHHKIRGI 58
WK + + I+GL+ L + G LDE + K+ + L +K K R
Sbjct: 20 WKEKLLAHMDILGLNTALKESESTGEKKSVLDESDEDYEEKLEK--FEALEEKKKKARSA 77
Query: 59 IVASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKAL--MLVHQYEL--FRMKDD 114
IV S+ K+ +STA AM +L K+ +KAL + + +L F+M ++
Sbjct: 78 IVLSVTDRVLRKIKKESTAAAMLLAL-------DKLYMSKALPNRIYPKQKLYSFKMSEN 130
Query: 115 ESIEEMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVE 174
S+E F +++ L+ + D +L +LP + ++ + + L+++
Sbjct: 131 LSVEGNIDEFLQIITDLENMNVIISDEDQAILLLTALPKAFDQLKDTLKYSSGKSILTLD 190
Query: 175 DLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSV 234
++ +++ E+ L + S K ++ L K K K +
Sbjct: 191 EVAAAIYSKELELGSVKKSIKVQAEGLYVKDKNENKGKGEQKG----------------- 233
Query: 235 KMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFK 293
+G K K + + GC+ C + GHF + CP+ K +FK
Sbjct: 234 ----------------------KGKGKKGKSKKKPGCWTCGEEGHFRSSCPNQNKPQFK 270
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.357 0.156 0.559
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,716,085
Number of Sequences: 26719
Number of extensions: 1185156
Number of successful extensions: 12141
Number of sequences better than 10.0: 275
Number of HSP's better than 10.0 without gapping: 55
Number of HSP's successfully gapped in prelim test: 225
Number of HSP's that attempted gapping in prelim test: 11270
Number of HSP's gapped (non-prelim): 863
length of query: 1664
length of database: 11,318,596
effective HSP length: 113
effective length of query: 1551
effective length of database: 8,299,349
effective search space: 12872290299
effective search space used: 12872290299
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC147002.8


