

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147001.15 + phase: 0
(146 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
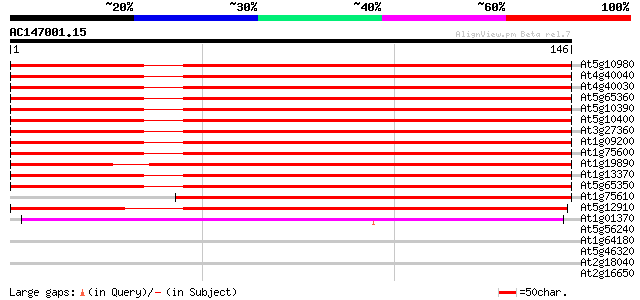
Score E
Sequences producing significant alignments: (bits) Value
At5g10980 histon H3 protein 235 5e-63
At4g40040 Histon H3 235 5e-63
At4g40030 histone H3.3 235 5e-63
At5g65360 histone H3 (sp|P05203) 231 1e-61
At5g10390 histone H3 - like protein 231 1e-61
At5g10400 histone H3 - like protein 231 1e-61
At3g27360 histone H3 like protein 231 1e-61
At1g09200 histone H3 like protein 231 1e-61
At1g75600 histone H3, putative 227 2e-60
At1g19890 hypothetical protein 226 3e-60
At1g13370 unknown protein 224 1e-59
At5g65350 histone H3 222 5e-59
At1g75610 histone H3, putative 195 8e-51
At5g12910 histone H3 -like protein 175 7e-45
At1g01370 centromeric histone H3 HTR12 111 2e-25
At5g56240 unknown protein 29 1.0
At1g64180 unknown protein 28 1.7
At5g46320 unknown protein 28 2.3
At2g18040 putative peptidyl-prolyl cis-trans isomerase 27 5.0
At2g16650 hypothetical protein 27 5.0
>At5g10980 histon H3 protein
Length = 136
Score = 235 bits (600), Expect = 5e-63
Identities = 126/146 (86%), Positives = 128/146 (87%), Gaps = 10/146 (6%)
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHR+R GTVALR
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGG----------VKKPHRYRPGTVALR 50
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL
Sbjct: 51 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 110
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 111 CAIHAKRVTIMPKDIQLARRIRGERA 136
>At4g40040 Histon H3
Length = 136
Score = 235 bits (600), Expect = 5e-63
Identities = 126/146 (86%), Positives = 128/146 (87%), Gaps = 10/146 (6%)
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHR+R GTVALR
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGG----------VKKPHRYRPGTVALR 50
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL
Sbjct: 51 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 110
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 111 CAIHAKRVTIMPKDIQLARRIRGERA 136
>At4g40030 histone H3.3
Length = 136
Score = 235 bits (600), Expect = 5e-63
Identities = 126/146 (86%), Positives = 128/146 (87%), Gaps = 10/146 (6%)
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHR+R GTVALR
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGG----------VKKPHRYRPGTVALR 50
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL
Sbjct: 51 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 110
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 111 CAIHAKRVTIMPKDIQLARRIRGERA 136
>At5g65360 histone H3 (sp|P05203)
Length = 136
Score = 231 bits (589), Expect = 1e-61
Identities = 125/146 (85%), Positives = 126/146 (85%), Gaps = 10/146 (6%)
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHRFR GTVALR
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGG----------VKKPHRFRPGTVALR 50
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQS AV ALQEAAEAYLVGLFEDTNL
Sbjct: 51 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNL 110
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 111 CAIHAKRVTIMPKDIQLARRIRGERA 136
>At5g10390 histone H3 - like protein
Length = 136
Score = 231 bits (589), Expect = 1e-61
Identities = 125/146 (85%), Positives = 126/146 (85%), Gaps = 10/146 (6%)
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHRFR GTVALR
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGG----------VKKPHRFRPGTVALR 50
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQS AV ALQEAAEAYLVGLFEDTNL
Sbjct: 51 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNL 110
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 111 CAIHAKRVTIMPKDIQLARRIRGERA 136
>At5g10400 histone H3 - like protein
Length = 136
Score = 231 bits (589), Expect = 1e-61
Identities = 125/146 (85%), Positives = 126/146 (85%), Gaps = 10/146 (6%)
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHRFR GTVALR
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGG----------VKKPHRFRPGTVALR 50
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQS AV ALQEAAEAYLVGLFEDTNL
Sbjct: 51 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNL 110
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 111 CAIHAKRVTIMPKDIQLARRIRGERA 136
>At3g27360 histone H3 like protein
Length = 136
Score = 231 bits (589), Expect = 1e-61
Identities = 125/146 (85%), Positives = 126/146 (85%), Gaps = 10/146 (6%)
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHRFR GTVALR
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGG----------VKKPHRFRPGTVALR 50
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQS AV ALQEAAEAYLVGLFEDTNL
Sbjct: 51 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNL 110
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 111 CAIHAKRVTIMPKDIQLARRIRGERA 136
>At1g09200 histone H3 like protein
Length = 136
Score = 231 bits (589), Expect = 1e-61
Identities = 125/146 (85%), Positives = 126/146 (85%), Gaps = 10/146 (6%)
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHRFR GTVALR
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGG----------VKKPHRFRPGTVALR 50
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQS AV ALQEAAEAYLVGLFEDTNL
Sbjct: 51 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNL 110
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 111 CAIHAKRVTIMPKDIQLARRIRGERA 136
>At1g75600 histone H3, putative
Length = 136
Score = 227 bits (578), Expect = 2e-60
Identities = 122/146 (83%), Positives = 125/146 (85%), Gaps = 10/146 (6%)
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKS GGKAPR LATK AR GG VKKPHR+R GTVALR
Sbjct: 1 MARTKQTARKSHGGKAPRTLLATKAARKSAPTTGG----------VKKPHRYRPGTVALR 50
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQD+KTDLRFQSHAVLALQEAAEAYLVGLFEDTNL
Sbjct: 51 EIRKYQKSTELLIRKLPFQRLVREIAQDYKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 110
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKD+QLARRIRGERA
Sbjct: 111 CAIHAKRVTIMPKDVQLARRIRGERA 136
>At1g19890 hypothetical protein
Length = 137
Score = 226 bits (577), Expect = 3e-60
Identities = 122/146 (83%), Positives = 125/146 (85%), Gaps = 9/146 (6%)
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGK PRKELATK AR R+ G VK+ HRFR GTVALR
Sbjct: 1 MARTKQTARKSTGGKGPRKELATKAAR---------KTRRPYRGGVKRAHRFRPGTVALR 51
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKST+LLIRKLPFQRLVREIAQDFK DLRFQSHAVLALQEAAEAYLVGLFEDTNL
Sbjct: 52 EIRKYQKSTDLLIRKLPFQRLVREIAQDFKVDLRFQSHAVLALQEAAEAYLVGLFEDTNL 111
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIM KDIQLARRIRGERA
Sbjct: 112 CAIHAKRVTIMSKDIQLARRIRGERA 137
>At1g13370 unknown protein
Length = 136
Score = 224 bits (572), Expect = 1e-59
Identities = 121/146 (82%), Positives = 124/146 (84%), Gaps = 10/146 (6%)
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQ+ARKS GGKAP K+LATK AR GG VKKPHRFR GTVALR
Sbjct: 1 MARTKQSARKSHGGKAPTKQLATKAARKSAPTTGG----------VKKPHRFRPGTVALR 50
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELL RKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL
Sbjct: 51 EIRKYQKSTELLNRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 110
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKD+QLARRIR ERA
Sbjct: 111 CAIHAKRVTIMPKDVQLARRIRAERA 136
>At5g65350 histone H3
Length = 139
Score = 222 bits (566), Expect = 5e-59
Identities = 120/146 (82%), Positives = 124/146 (84%), Gaps = 10/146 (6%)
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTAR STGGKAPRK+LA K AR GG VKKPHRFR GTVALR
Sbjct: 1 MARTKQTARISTGGKAPRKQLAPKAARQSAPATGG----------VKKPHRFRPGTVALR 50
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
+IRKYQKSTE+LIRKLPFQRLVREIAQDFKTDLRFQS AV ALQEAAEAYLVGLFEDTNL
Sbjct: 51 DIRKYQKSTEILIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNL 110
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPK+IQLARRIRGERA
Sbjct: 111 CAIHAKRVTIMPKEIQLARRIRGERA 136
>At1g75610 histone H3, putative
Length = 115
Score = 195 bits (495), Expect = 8e-51
Identities = 99/103 (96%), Positives = 101/103 (97%)
Query: 44 GHVKKPHRFRCGTVALREIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLAL 103
G VKKPHR+R GTVALREIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLAL
Sbjct: 13 GGVKKPHRYRPGTVALREIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLAL 72
Query: 104 QEAAEAYLVGLFEDTNLCAIHAKRVTIMPKDIQLARRIRGERA 146
QEAAEAYLVGLFEDTNLCAIHAKRVTIMPKD+QLARRIRGERA
Sbjct: 73 QEAAEAYLVGLFEDTNLCAIHAKRVTIMPKDVQLARRIRGERA 115
>At5g12910 histone H3 -like protein
Length = 131
Score = 175 bits (444), Expect = 7e-45
Identities = 92/145 (63%), Positives = 111/145 (76%), Gaps = 15/145 (10%)
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MAR+ QTARK+TGGKAP + P +KKP+R++ GTVALR
Sbjct: 1 MARSNQTARKATGGKAPHFAMRVWQHSTPP---------------LKKPYRYKPGTVALR 45
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQK+T+L+IRKLPFQRLV+EIAQ K DLRFQ+ AV ALQEAAEA++VG+FEDTNL
Sbjct: 46 EIRKYQKTTDLVIRKLPFQRLVKEIAQSLKADLRFQTGAVSALQEAAEAFMVGMFEDTNL 105
Query: 121 CAIHAKRVTIMPKDIQLARRIRGER 145
CA+HAKR TIMPKDIQLA+R+RG+R
Sbjct: 106 CAMHAKRSTIMPKDIQLAKRLRGDR 130
>At1g01370 centromeric histone H3 HTR12
Length = 178
Score = 111 bits (277), Expect = 2e-25
Identities = 66/143 (46%), Positives = 85/143 (59%), Gaps = 2/143 (1%)
Query: 4 TKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALREIR 63
T T R GG ++ T +G +S + M G KK +R+R GTVAL+EIR
Sbjct: 32 TTPTRRGGEGGDNTQQTNPTTSPATGTRRGAKRSRQAMPRGSQKKSYRYRPGTVALKEIR 91
Query: 64 KYQKSTELLIRKLPFQRLVREIAQDFKTDL--RFQSHAVLALQEAAEAYLVGLFEDTNLC 121
+QK T LLI F R VR I R+ + A++ALQEAAE YLVGLF D+ LC
Sbjct: 92 HFQKQTNLLIPAASFIREVRSITHMLAPPQINRWTAEALVALQEAAEDYLVGLFSDSMLC 151
Query: 122 AIHAKRVTIMPKDIQLARRIRGE 144
AIHA+RVT+M KD +LARR+ G+
Sbjct: 152 AIHARRVTLMRKDFELARRLGGK 174
>At5g56240 unknown protein
Length = 986
Score = 28.9 bits (63), Expect = 1.0
Identities = 13/22 (59%), Positives = 17/22 (77%), Gaps = 2/22 (9%)
Query: 31 AKGGGKSMRKMREGHVKKPHRF 52
+KG KS+RK+R G KKPH+F
Sbjct: 348 SKGTNKSLRKIRRG--KKPHKF 367
>At1g64180 unknown protein
Length = 593
Score = 28.1 bits (61), Expect = 1.7
Identities = 12/26 (46%), Positives = 16/26 (61%)
Query: 22 ATKVARAPRAKGGGKSMRKMREGHVK 47
ATK+ RAP G + R++R GH K
Sbjct: 89 ATKMHRAPLGSAGPSNSRRLRHGHGK 114
>At5g46320 unknown protein
Length = 118
Score = 27.7 bits (60), Expect = 2.3
Identities = 21/83 (25%), Positives = 41/83 (49%)
Query: 2 ARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALRE 61
AR +++ + T + +L + +A + K K +RK +E ++KK +R + L +
Sbjct: 31 ARRVRSSTRQTRILKNQAKLDSLMAELEQIKAYEKVLRKGQEQNIKKYNRKEISDLKLED 90
Query: 62 IRKYQKSTELLIRKLPFQRLVRE 84
+ +QK E L L +R+ E
Sbjct: 91 LVVFQKKLENLQDDLKKKRVELE 113
>At2g18040 putative peptidyl-prolyl cis-trans isomerase
Length = 119
Score = 26.6 bits (57), Expect = 5.0
Identities = 18/47 (38%), Positives = 26/47 (55%), Gaps = 3/47 (6%)
Query: 14 GKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
GKA +E+AT+V+ AK GG + G ++KP F T AL+
Sbjct: 55 GKANFEEVATRVSDCSSAKRGG-DLGSFGRGQMQKP--FEEATYALK 98
>At2g16650 hypothetical protein
Length = 369
Score = 26.6 bits (57), Expect = 5.0
Identities = 28/108 (25%), Positives = 47/108 (42%), Gaps = 14/108 (12%)
Query: 9 RKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALREIRKYQKS 68
R + G +P + T VAR AKG G K+ + V G V++ +R Y +
Sbjct: 99 RMVSSGISPNEASVTSVARLAAAKGNGDYAFKVVKEFVS------VGGVSIPRLRTYAPA 152
Query: 69 TELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFE 116
KL E + ++ + ++ A +AL+EA + L+ L E
Sbjct: 153 LLCFCEKL-------EAEKGYEVEEHMEA-AGIALEEAEISALLKLRE 192
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.135 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,834,582
Number of Sequences: 26719
Number of extensions: 99743
Number of successful extensions: 285
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 18
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 253
Number of HSP's gapped (non-prelim): 24
length of query: 146
length of database: 11,318,596
effective HSP length: 90
effective length of query: 56
effective length of database: 8,913,886
effective search space: 499177616
effective search space used: 499177616
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC147001.15


