

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147001.13 + phase: 0
(136 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
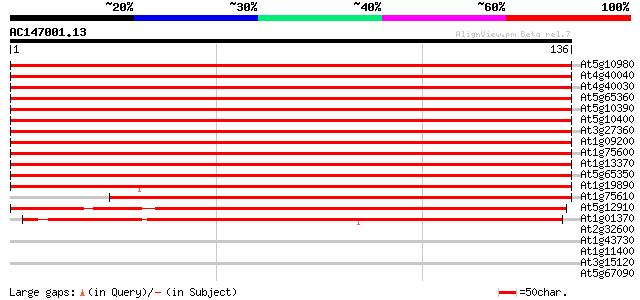
Score E
Sequences producing significant alignments: (bits) Value
At5g10980 histon H3 protein 249 2e-67
At4g40040 Histon H3 249 2e-67
At4g40030 histone H3.3 249 2e-67
At5g65360 histone H3 (sp|P05203) 241 1e-64
At5g10390 histone H3 - like protein 241 1e-64
At5g10400 histone H3 - like protein 241 1e-64
At3g27360 histone H3 like protein 241 1e-64
At1g09200 histone H3 like protein 241 1e-64
At1g75600 histone H3, putative 240 1e-64
At1g13370 unknown protein 236 3e-63
At5g65350 histone H3 231 7e-62
At1g19890 hypothetical protein 226 3e-60
At1g75610 histone H3, putative 209 5e-55
At5g12910 histone H3 -like protein 176 3e-45
At1g01370 centromeric histone H3 HTR12 108 8e-25
At2g32600 putative spliceosome associated protein 28 1.9
At1g43730 hypothetical protein 27 2.5
At1g11400 unknown protein 27 2.5
At3g15120 chaperone-like ATPase 26 5.5
At5g67090 subtilisin-type protease-like 26 7.2
>At5g10980 histon H3 protein
Length = 136
Score = 249 bits (637), Expect = 2e-67
Identities = 126/136 (92%), Positives = 131/136 (95%)
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS PTTGG+KKPHRYRPGTVALREIRKYQK TE
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 60
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQSHAVLALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAKRVTI 120
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLARRIRGERA
Sbjct: 121 MPKDIQLARRIRGERA 136
>At4g40040 Histon H3
Length = 136
Score = 249 bits (637), Expect = 2e-67
Identities = 126/136 (92%), Positives = 131/136 (95%)
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS PTTGG+KKPHRYRPGTVALREIRKYQK TE
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 60
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQSHAVLALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAKRVTI 120
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLARRIRGERA
Sbjct: 121 MPKDIQLARRIRGERA 136
>At4g40030 histone H3.3
Length = 136
Score = 249 bits (637), Expect = 2e-67
Identities = 126/136 (92%), Positives = 131/136 (95%)
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS PTTGG+KKPHRYRPGTVALREIRKYQK TE
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 60
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQSHAVLALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAKRVTI 120
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLARRIRGERA
Sbjct: 121 MPKDIQLARRIRGERA 136
>At5g65360 histone H3 (sp|P05203)
Length = 136
Score = 241 bits (614), Expect = 1e-64
Identities = 122/136 (89%), Positives = 128/136 (93%)
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS P TGG+KKPHR+RPGTVALREIRKYQK TE
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 60
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQS AV ALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNLCAIHAKRVTI 120
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLARRIRGERA
Sbjct: 121 MPKDIQLARRIRGERA 136
>At5g10390 histone H3 - like protein
Length = 136
Score = 241 bits (614), Expect = 1e-64
Identities = 122/136 (89%), Positives = 128/136 (93%)
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS P TGG+KKPHR+RPGTVALREIRKYQK TE
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 60
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQS AV ALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNLCAIHAKRVTI 120
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLARRIRGERA
Sbjct: 121 MPKDIQLARRIRGERA 136
>At5g10400 histone H3 - like protein
Length = 136
Score = 241 bits (614), Expect = 1e-64
Identities = 122/136 (89%), Positives = 128/136 (93%)
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS P TGG+KKPHR+RPGTVALREIRKYQK TE
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 60
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQS AV ALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNLCAIHAKRVTI 120
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLARRIRGERA
Sbjct: 121 MPKDIQLARRIRGERA 136
>At3g27360 histone H3 like protein
Length = 136
Score = 241 bits (614), Expect = 1e-64
Identities = 122/136 (89%), Positives = 128/136 (93%)
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS P TGG+KKPHR+RPGTVALREIRKYQK TE
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 60
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQS AV ALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNLCAIHAKRVTI 120
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLARRIRGERA
Sbjct: 121 MPKDIQLARRIRGERA 136
>At1g09200 histone H3 like protein
Length = 136
Score = 241 bits (614), Expect = 1e-64
Identities = 122/136 (89%), Positives = 128/136 (93%)
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS P TGG+KKPHR+RPGTVALREIRKYQK TE
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 60
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQS AV ALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNLCAIHAKRVTI 120
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLARRIRGERA
Sbjct: 121 MPKDIQLARRIRGERA 136
>At1g75600 histone H3, putative
Length = 136
Score = 240 bits (613), Expect = 1e-64
Identities = 121/136 (88%), Positives = 128/136 (93%)
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+S GKAPR LATKAARKS PTTGG+KKPHRYRPGTVALREIRKYQK TE
Sbjct: 1 MARTKQTARKSHGGKAPRTLLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 60
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ++KTDLRFQSHAVLALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 61 LLIRKLPFQRLVREIAQDYKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAKRVTI 120
Query: 121 MVKDIQLARRIRGERA 136
M KD+QLARRIRGERA
Sbjct: 121 MPKDVQLARRIRGERA 136
>At1g13370 unknown protein
Length = 136
Score = 236 bits (602), Expect = 3e-63
Identities = 119/136 (87%), Positives = 127/136 (92%)
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQ+AR+S GKAP KQLATKAARKS PTTGG+KKPHR+RPGTVALREIRKYQK TE
Sbjct: 1 MARTKQSARKSHGGKAPTKQLATKAARKSAPTTGGVKKPHRFRPGTVALREIRKYQKSTE 60
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LL RKLPFQRLVREIAQ+FKTDLRFQSHAVLALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 61 LLNRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAKRVTI 120
Query: 121 MVKDIQLARRIRGERA 136
M KD+QLARRIR ERA
Sbjct: 121 MPKDVQLARRIRAERA 136
>At5g65350 histone H3
Length = 139
Score = 231 bits (590), Expect = 7e-62
Identities = 117/136 (86%), Positives = 126/136 (92%)
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR ST GKAPRKQLA KAAR+S P TGG+KKPHR+RPGTVALR+IRKYQK TE
Sbjct: 1 MARTKQTARISTGGKAPRKQLAPKAARQSAPATGGVKKPHRFRPGTVALRDIRKYQKSTE 60
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
+LIRKLPFQRLVREIAQ+FKTDLRFQS AV ALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 61 ILIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNLCAIHAKRVTI 120
Query: 121 MVKDIQLARRIRGERA 136
M K+IQLARRIRGERA
Sbjct: 121 MPKEIQLARRIRGERA 136
>At1g19890 hypothetical protein
Length = 137
Score = 226 bits (576), Expect = 3e-60
Identities = 116/137 (84%), Positives = 126/137 (91%), Gaps = 1/137 (0%)
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSV-PTTGGIKKPHRYRPGTVALREIRKYQKGT 59
MARTKQTAR+ST GK PRK+LATKAARK+ P GG+K+ HR+RPGTVALREIRKYQK T
Sbjct: 1 MARTKQTARKSTGGKGPRKELATKAARKTRRPYRGGVKRAHRFRPGTVALREIRKYQKST 60
Query: 60 ELLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRIT 119
+LLIRKLPFQRLVREIAQ+FK DLRFQSHAVLALQEA EAYLVGLFEDTNLCAIHAKR+T
Sbjct: 61 DLLIRKLPFQRLVREIAQDFKVDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAKRVT 120
Query: 120 VMVKDIQLARRIRGERA 136
+M KDIQLARRIRGERA
Sbjct: 121 IMSKDIQLARRIRGERA 137
>At1g75610 histone H3, putative
Length = 115
Score = 209 bits (531), Expect = 5e-55
Identities = 103/112 (91%), Positives = 108/112 (95%)
Query: 25 AARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTELLIRKLPFQRLVREIAQNFKTDLR 84
AARKS PTTGG+KKPHRYRPGTVALREIRKYQK TELLIRKLPFQRLVREIAQ+FKTDLR
Sbjct: 4 AARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLR 63
Query: 85 FQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITVMVKDIQLARRIRGERA 136
FQSHAVLALQEA EAYLVGLFEDTNLCAIHAKR+T+M KD+QLARRIRGERA
Sbjct: 64 FQSHAVLALQEAAEAYLVGLFEDTNLCAIHAKRVTIMPKDVQLARRIRGERA 115
>At5g12910 histone H3 -like protein
Length = 131
Score = 176 bits (447), Expect = 3e-45
Identities = 91/135 (67%), Positives = 112/135 (82%), Gaps = 5/135 (3%)
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MAR+ QTAR++T GKAP A + + S P +KKP+RY+PGTVALREIRKYQK T+
Sbjct: 1 MARSNQTARKATGGKAPH--FAMRVWQHSTPP---LKKPYRYKPGTVALREIRKYQKTTD 55
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
L+IRKLPFQRLV+EIAQ+ K DLRFQ+ AV ALQEA EA++VG+FEDTNLCA+HAKR T+
Sbjct: 56 LVIRKLPFQRLVKEIAQSLKADLRFQTGAVSALQEAAEAFMVGMFEDTNLCAMHAKRSTI 115
Query: 121 MVKDIQLARRIRGER 135
M KDIQLA+R+RG+R
Sbjct: 116 MPKDIQLAKRLRGDR 130
>At1g01370 centromeric histone H3 HTR12
Length = 178
Score = 108 bits (270), Expect = 8e-25
Identities = 65/133 (48%), Positives = 85/133 (63%), Gaps = 5/133 (3%)
Query: 4 TKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTELLI 63
T+QT T A + K +R+++P G KK +RYRPGTVAL+EIR +QK T LLI
Sbjct: 45 TQQT--NPTTSPATGTRRGAKRSRQAMPR-GSQKKSYRYRPGTVALKEIRHFQKQTNLLI 101
Query: 64 RKLPFQRLVREIAQNFKTDL--RFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITVM 121
F R VR I R+ + A++ALQEA E YLVGLF D+ LCAIHA+R+T+M
Sbjct: 102 PAASFIREVRSITHMLAPPQINRWTAEALVALQEAAEDYLVGLFSDSMLCAIHARRVTLM 161
Query: 122 VKDIQLARRIRGE 134
KD +LARR+ G+
Sbjct: 162 RKDFELARRLGGK 174
>At2g32600 putative spliceosome associated protein
Length = 277
Score = 27.7 bits (60), Expect = 1.9
Identities = 30/101 (29%), Positives = 39/101 (37%), Gaps = 9/101 (8%)
Query: 12 TLGKAPRKQLATKAAR-------KSVPTTGGIKKPHRYRPGTVALREIRKYQKGTELLIR 64
T GK + LA +AAR K P + + G R ++Y EL R
Sbjct: 71 TQGKRHQTNLAKRAAREAKDAPTKPQPLKRNVSVRRTVKIGRPGYRVTKQYDP--ELQQR 128
Query: 65 KLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLF 105
L FQ EI N K RF S +Q ++Y LF
Sbjct: 129 SLLFQIEYPEIEDNIKPRHRFMSSYEQKVQPYDKSYQYLLF 169
>At1g43730 hypothetical protein
Length = 320
Score = 27.3 bits (59), Expect = 2.5
Identities = 16/48 (33%), Positives = 28/48 (58%), Gaps = 4/48 (8%)
Query: 88 HAVLALQEAVEAYLVGLFEDTNLCAIH---AKRITVMVKDIQLARRIR 132
+ L ++ A +AY+ ++ + NLC +H A+ ++KDIQL R R
Sbjct: 247 NTTLIIRLAFQAYVYAIWRERNLC-LHTGVARVFDSVLKDIQLTIRAR 293
>At1g11400 unknown protein
Length = 204
Score = 27.3 bits (59), Expect = 2.5
Identities = 13/36 (36%), Positives = 20/36 (55%)
Query: 34 GGIKKPHRYRPGTVALREIRKYQKGTELLIRKLPFQ 69
G ++KP R RPG E+ KYQ L+ +++ Q
Sbjct: 34 GTLRKPIRIRPGYTPEDEVVKYQSKGSLMKKEMASQ 69
>At3g15120 chaperone-like ATPase
Length = 1954
Score = 26.2 bits (56), Expect = 5.5
Identities = 19/75 (25%), Positives = 35/75 (46%), Gaps = 2/75 (2%)
Query: 35 GIKKPHRYRPGTVALREIRKYQKGTELLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQ 94
G++ H + + +R K L +P++ L +I Q FKTDL + ++
Sbjct: 1248 GVRVSHAWNTFFEQVETLRVSTKMMILATSGMPYKLLPPKIQQFFKTDLSKECQPTMS-- 1305
Query: 95 EAVEAYLVGLFEDTN 109
EAV + V + E ++
Sbjct: 1306 EAVPQFNVQVVESSD 1320
>At5g67090 subtilisin-type protease-like
Length = 736
Score = 25.8 bits (55), Expect = 7.2
Identities = 14/48 (29%), Positives = 23/48 (47%)
Query: 2 ARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVAL 49
A K R++ +G P ++ T ++R + I KP PGT+ L
Sbjct: 454 ATAKLEFRKTVIGTKPAPEVGTYSSRGPFTSFPQILKPDILAPGTLIL 501
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.135 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,492,486
Number of Sequences: 26719
Number of extensions: 85106
Number of successful extensions: 201
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 181
Number of HSP's gapped (non-prelim): 22
length of query: 136
length of database: 11,318,596
effective HSP length: 89
effective length of query: 47
effective length of database: 8,940,605
effective search space: 420208435
effective search space used: 420208435
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 54 (25.4 bits)
Medicago: description of AC147001.13


