

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147000.4 + phase: 1 /pseudo
(187 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
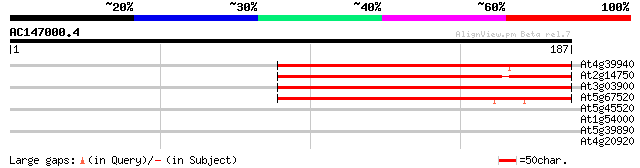
Score E
Sequences producing significant alignments: (bits) Value
At4g39940 adenosine-5'-phosphosulfate-kinase 156 7e-39
At2g14750 putative adenosine phosphosulfate kinase 152 1e-37
At3g03900 putative adenylylsulfate kinase 142 1e-34
At5g67520 adenylylsulfate kinase-like protein 138 2e-33
At5g45520 putative protein 28 3.8
At1g54000 unknown protein 27 4.9
At5g39890 unknown protein 27 6.4
At4g20920 putative protein 27 6.4
>At4g39940 adenosine-5'-phosphosulfate-kinase
Length = 293
Score = 156 bits (394), Expect = 7e-39
Identities = 71/99 (71%), Positives = 86/99 (86%), Gaps = 1/99 (1%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY++DRDACR+LLP+GDF+EVF+DVPLHVCE+RDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 194 LISPYRRDRDACRSLLPDGDFVEVFMDVPLHVCESRDPKGLYKLARAGKIKGFTGIDDPY 253
Query: 150 EPPCCCEIILQQKGSD-CKSPKDMAETVISYLEKSGHLQ 187
E P CE++L+ G D SP+ MAE +ISYL+ G+L+
Sbjct: 254 EAPVNCEVVLKHTGDDESCSPRQMAENIISYLQNKGYLE 292
>At2g14750 putative adenosine phosphosulfate kinase
Length = 276
Score = 152 bits (383), Expect = 1e-37
Identities = 73/98 (74%), Positives = 83/98 (84%), Gaps = 2/98 (2%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+ DRDACR+LLPEGDF+EVF+DVPL VCEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 180 LISPYRTDRDACRSLLPEGDFVEVFMDVPLSVCEARDPKGLYKLARAGKIKGFTGIDDPY 239
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP CEI L ++G SP +MAE V+ YL+ G+LQ
Sbjct: 240 EPPLNCEISLGREGG--TSPIEMAEKVVGYLDNKGYLQ 275
>At3g03900 putative adenylylsulfate kinase
Length = 208
Score = 142 bits (357), Expect = 1e-34
Identities = 68/98 (69%), Positives = 78/98 (79%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+KDRDACR ++ FIEVF+++ L +CEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 109 LISPYRKDRDACREMIQNSSFIEVFMNMSLQLCEARDPKGLYKLARAGKIKGFTGIDDPY 168
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
E P CEI L++K +C SP MAE VISYLE G LQ
Sbjct: 169 ESPLNCEIELKEKEGECPSPVAMAEEVISYLEDKGFLQ 206
>At5g67520 adenylylsulfate kinase-like protein
Length = 310
Score = 138 bits (348), Expect = 2e-33
Identities = 69/113 (61%), Positives = 84/113 (74%), Gaps = 15/113 (13%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+ +R ACRALLP+GDFIEVF+DVPLHVCEARDPKGLYK ARAGKIKGFTG+DDPY
Sbjct: 188 LISPYRIERAACRALLPQGDFIEVFMDVPLHVCEARDPKGLYKRARAGKIKGFTGVDDPY 247
Query: 150 EPPCCCEIILQ--------QKGSDCKSPK-------DMAETVISYLEKSGHLQ 187
E P CEI++Q S SP +MA+ V+SYL+++G+L+
Sbjct: 248 EAPLDCEIVIQNSRDKGLSSSSSSSSSPSSSSSSLCEMADIVVSYLDQNGYLK 300
>At5g45520 putative protein
Length = 1167
Score = 27.7 bits (60), Expect = 3.8
Identities = 14/41 (34%), Positives = 22/41 (53%)
Query: 113 VFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYEPPC 153
+++DV R PKG+ KL+R +KGF + +E C
Sbjct: 465 IYLDVSECYMLDRMPKGIAKLSRLQVLKGFVISESDHENNC 505
>At1g54000 unknown protein
Length = 391
Score = 27.3 bits (59), Expect = 4.9
Identities = 17/62 (27%), Positives = 26/62 (41%), Gaps = 4/62 (6%)
Query: 93 PYQKDRDACRALLPEG----DFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDP 148
PY K RD +G DF+ F+ +PL + A P + ++G T + P
Sbjct: 64 PYGKSRDDPNGKFSDGLITPDFLAKFMKIPLAIAPALQPNVNVSRGASFAVEGATLLGAP 123
Query: 149 YE 150
E
Sbjct: 124 VE 125
>At5g39890 unknown protein
Length = 276
Score = 26.9 bits (58), Expect = 6.4
Identities = 11/31 (35%), Positives = 18/31 (57%)
Query: 78 SRRSFRKHTKDCLISPYQKDRDACRALLPEG 108
SR+ ++ +K LI P QK D C+ + +G
Sbjct: 31 SRKKIQRRSKKTLICPVQKLFDTCKKVFADG 61
>At4g20920 putative protein
Length = 870
Score = 26.9 bits (58), Expect = 6.4
Identities = 11/28 (39%), Positives = 15/28 (53%)
Query: 108 GDFIEVFIDVPLHVCEARDPKGLYKLAR 135
GD I + P +C P G+YKL+R
Sbjct: 316 GDAIVASVGYPWRICCGISPNGIYKLSR 343
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.340 0.150 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,191,899
Number of Sequences: 26719
Number of extensions: 164704
Number of successful extensions: 357
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 347
Number of HSP's gapped (non-prelim): 8
length of query: 187
length of database: 11,318,596
effective HSP length: 93
effective length of query: 94
effective length of database: 8,833,729
effective search space: 830370526
effective search space used: 830370526
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC147000.4


