

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146972.11 + phase: 0
(362 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
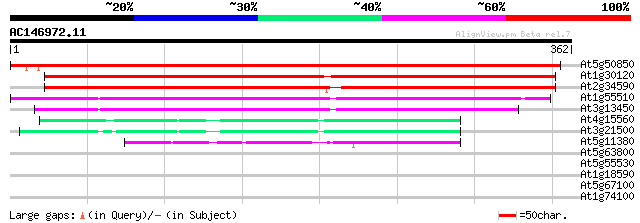
Score E
Sequences producing significant alignments: (bits) Value
At5g50850 pyruvate dehydrogenase E1 component beta subunit, mito... 608 e-174
At1g30120 unknown protein 266 1e-71
At2g34590 putative pyruvate dehydrogenase E1 beta subunit 265 2e-71
At1g55510 branched-chain alpha-keto acid decarboxylase E1 beta s... 220 8e-58
At3g13450 branched chain alpha-keto acid dehydrogenase E1 beta s... 217 7e-57
At4g15560 1-D-deoxyxylulose 5-phosphate synthase, putative 73 3e-13
At3g21500 1-D-deoxyxylulose 5-phosphate synthase, putative 68 9e-12
At5g11380 1-D-deoxyxylulose 5-phosphate synthase - like protein 50 2e-06
At5g63800 beta-galactosidase like protein 31 1.2
At5g55530 unknown protein 30 2.1
At1g18590 unknown protein 29 4.6
At5g67100 DNA polymerase alpha 1 28 6.0
At1g74100 putative flavonol sulfotransferase 28 7.8
>At5g50850 pyruvate dehydrogenase E1 component beta subunit,
mitochondrial precursor (PDHE1-B) (sp|Q38799)
Length = 363
Score = 608 bits (1569), Expect = e-174
Identities = 311/362 (85%), Positives = 332/362 (90%), Gaps = 7/362 (1%)
Query: 1 MLGVIRNKV-----NLLRPSFS--AFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMG 53
MLG++R + L R F+ + R ++ AK+MTVRDALNSA+DEEMSADPKVF+MG
Sbjct: 1 MLGILRQRAIDGASTLRRTRFALVSARSYAAGAKEMTVRDALNSAIDEEMSADPKVFVMG 60
Query: 54 EEVGEYQGAYKISKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSM 113
EEVG+YQGAYKI+KGLLEKYGPERV DTPITEAGFTGIGVGAAY GLKPVVEFMTFNFSM
Sbjct: 61 EEVGQYQGAYKITKGLLEKYGPERVYDTPITEAGFTGIGVGAAYAGLKPVVEFMTFNFSM 120
Query: 114 QAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAP 173
QAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHS CYA+WY S PGLKVLAP
Sbjct: 121 QAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSQCYAAWYASVPGLKVLAP 180
Query: 174 YSSEDARGLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDV 233
YS+EDARGLLKAAIRDPDPVVFLENELLYGESFP+S E LDSSFCLPIGKAKIEREGKDV
Sbjct: 181 YSAEDARGLLKAAIRDPDPVVFLENELLYGESFPISEEALDSSFCLPIGKAKIEREGKDV 240
Query: 234 TITAFSKMVGFALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFP 293
TI FSKMVGFALKAAE L +EGISAEVINLRSIRPLDRATINASVRKT+RLVTVEEGFP
Sbjct: 241 TIVTFSKMVGFALKAAEKLAEEGISAEVINLRSIRPLDRATINASVRKTSRLVTVEEGFP 300
Query: 294 QHGVGAEICASVIEESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRACH 353
QHGV AEICASV+EESF YLDAPVERIAGADVPMPYAANLERLA+PQIEDIVRA+KRAC+
Sbjct: 301 QHGVCAEICASVVEESFSYLDAPVERIAGADVPMPYAANLERLALPQIEDIVRASKRACY 360
Query: 354 RS 355
RS
Sbjct: 361 RS 362
>At1g30120 unknown protein
Length = 406
Score = 266 bits (681), Expect = 1e-71
Identities = 137/330 (41%), Positives = 208/330 (62%), Gaps = 4/330 (1%)
Query: 23 SSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTP 82
+S+ ++ + +AL L+EEM DP V +MGE+VG Y G+YK++KGL +K+G RVLDTP
Sbjct: 80 ASTGHELLLFEALQEGLEEEMDRDPHVCVMGEDVGHYGGSYKVTKGLADKFGDLRVLDTP 139
Query: 83 ITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRG 142
I E FTG+G+GAA GL+PV+E M F + A + I N+ +Y S GQ +P+V RG
Sbjct: 140 ICENAFTGMGIGAAMTGLRPVIEGMNMGFLLLAFNQISNNCGMLHYTSGGQFTIPVVIRG 199
Query: 143 PNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLY 202
P G +GA+HS S++ S PG++++A + +A+GL+KAAIR +PV+ E+ LLY
Sbjct: 200 PGGVGRQLGAEHSQRLESYFQSIPGIQMVACSTPYNAKGLMKAAIRSENPVILFEHVLLY 259
Query: 203 GESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVI 262
+ ++ D + + +A++ R G+ +TI +S+M ++AA+TL +G EVI
Sbjct: 260 N----LKEKIPDEDYVCNLEEAEMVRPGEHITILTYSRMRYHVMQAAKTLVNKGYDPEVI 315
Query: 263 NLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAG 322
++RS++P D TI SV+KT+R++ VEE G+GA + A++ E YLDAPV ++
Sbjct: 316 DIRSLKPFDLHTIGNSVKKTHRVLIVEECMRTGGIGASLTAAINENFHDYLDAPVMCLSS 375
Query: 323 ADVPMPYAANLERLAVPQIEDIVRAAKRAC 352
DVP PYA LE V Q IV A ++ C
Sbjct: 376 QDVPTPYAGTLEEWTVVQPAQIVTAVEQLC 405
>At2g34590 putative pyruvate dehydrogenase E1 beta subunit
Length = 406
Score = 265 bits (678), Expect = 2e-71
Identities = 140/332 (42%), Positives = 206/332 (61%), Gaps = 8/332 (2%)
Query: 23 SSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTP 82
S ++ + +AL L+EEM DP V +MGE+VG Y G+YK++KGL +K+G RVLDTP
Sbjct: 80 SKPGHELLLFEALQEGLEEEMDRDPHVCVMGEDVGHYGGSYKVTKGLADKFGDLRVLDTP 139
Query: 83 ITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRG 142
I E FTG+G+GAA GL+PV+E M F + A + I N+ +Y S GQ +P+V RG
Sbjct: 140 ICENAFTGMGIGAAMTGLRPVIEGMNMGFLLLAFNQISNNCGMLHYTSGGQFTIPVVIRG 199
Query: 143 PNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLY 202
P G +GA+HS S++ S PG++++A + +A+GL+KAAIR +PV+ E+ LLY
Sbjct: 200 PGGVGRQLGAEHSQRLESYFQSIPGIQMVACSTPYNAKGLMKAAIRSENPVILFEHVLLY 259
Query: 203 G--ESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAE 260
ES P D + + +A++ R G+ +TI +S+M ++AA+TL +G E
Sbjct: 260 NLKESIP------DEEYICNLEEAEMVRPGEHITILTYSRMRYHVMQAAKTLVNKGYDPE 313
Query: 261 VINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERI 320
VI++RS++P D TI SV+KT+R++ VEE G+GA + A++ E YLDAPV +
Sbjct: 314 VIDIRSLKPFDLYTIGNSVKKTHRVLIVEECMRTGGIGASLTAAINENFHDYLDAPVMCL 373
Query: 321 AGADVPMPYAANLERLAVPQIEDIVRAAKRAC 352
+ DVP PYA LE V Q IV A ++ C
Sbjct: 374 SSQDVPTPYAGTLEEWTVVQPAQIVTAVEQLC 405
>At1g55510 branched-chain alpha-keto acid decarboxylase E1 beta
subunit
Length = 352
Score = 220 bits (561), Expect = 8e-58
Identities = 126/350 (36%), Positives = 194/350 (55%), Gaps = 7/350 (2%)
Query: 1 MLGVIRNKVNLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQ 60
+LG K++ S A R + + K + + A+N AL + DP+ ++ GE+VG +
Sbjct: 4 LLGRSCRKLSFPSLSHGARRVSTETGKPLNLYSAINQALHIALDTDPRSYVFGEDVG-FG 62
Query: 61 GAYKISKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHII 120
G ++ + GL E++G RV +TP+ E G G G+G A G + +VE ++ A D I+
Sbjct: 63 GVFRCTTGLAERFGKNRVFNTPLCEQGIVGFGIGLAAMGNRAIVEIQFADYIYPAFDQIV 122
Query: 121 NSAAKSNYMSAGQINVP-IVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDA 179
N AAK Y S Q N + R P GA G HS +++ PG+KV+ P S +A
Sbjct: 123 NEAAKFRYRSGNQFNCGGLTIRAPYGAVGHGGHYHSQSPEAFFCHVPGIKVVIPRSPREA 182
Query: 180 RGLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFS 239
+GLL + IRDP+PVVF E + LY ++ EV + + +P+ +A++ REG D+T+ +
Sbjct: 183 KGLLLSCIRDPNPVVFFEPKWLYRQAVE---EVPEHDYMIPLSEAEVIREGNDITLVGWG 239
Query: 240 KMVGFALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGA 299
+ +A EKEGIS E+I+L+++ P D+ T+ ASV+KT RL+ E G GA
Sbjct: 240 AQLTVMEQACLDAEKEGISCELIDLKTLLPWDKETVEASVKKTGRLLISHEAPVTGGFGA 299
Query: 300 EICASVIEESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAK 349
EI A+++E F L+APV R+ G D P P E +P I+ A K
Sbjct: 300 EISATILERCFLKLEAPVSRVCGLDTPFPLV--FEPFYMPTKNKILDAIK 347
>At3g13450 branched chain alpha-keto acid dehydrogenase E1 beta
subunit
Length = 358
Score = 217 bits (553), Expect = 7e-57
Identities = 115/313 (36%), Positives = 178/313 (56%), Gaps = 5/313 (1%)
Query: 17 SAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPE 76
S ++S S K M + A+N AL + DP+ ++ GE+VG + G ++ + GL E++G
Sbjct: 26 STVENVSESGKSMNLYSAINQALHIALETDPRSYVFGEDVG-FGGVFRCTTGLAERFGKS 84
Query: 77 RVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINV 136
RV +TP+ E G G G+G A G + + E ++ A D I+N AAK Y S Q N
Sbjct: 85 RVFNTPLCEQGIVGFGIGLAAMGNRVIAEIQFADYIFPAFDQIVNEAAKFRYRSGNQFNC 144
Query: 137 P-IVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVF 195
+ R P GA G HS +++ PG+KV+ P S +A+GLL ++IRDP+PVVF
Sbjct: 145 GGLTIRAPYGAVGHGGHYHSQSPEAFFCHVPGIKVVIPRSPREAKGLLLSSIRDPNPVVF 204
Query: 196 LENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKE 255
E + LY ++ +V + + +P+ +A++ REG D+T+ + + +A E E
Sbjct: 205 FEPKWLYRQAVE---DVPEDDYMIPLSEAEVMREGSDITLVGWGAQLTIMEQACLDAENE 261
Query: 256 GISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDA 315
GIS E+I+L+++ P D+ + SVRKT RL+ E G GAEI A+++E F L+A
Sbjct: 262 GISCELIDLKTLIPWDKEIVETSVRKTGRLLISHEAPVTGGFGAEIAATIVERCFLRLEA 321
Query: 316 PVERIAGADVPMP 328
PV R+ G D P P
Sbjct: 322 PVSRVCGLDTPFP 334
>At4g15560 1-D-deoxyxylulose 5-phosphate synthase, putative
Length = 717
Score = 72.8 bits (177), Expect = 3e-13
Identities = 69/275 (25%), Positives = 108/275 (39%), Gaps = 20/275 (7%)
Query: 20 RHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVL 79
R ++ K + AL E D V + +G G L ++ P R
Sbjct: 390 RQFKTTNKTQSYTTYFAEALVAEAEVDKDVVAIHAAMGGGTGL-----NLFQRRFPTRCF 444
Query: 80 DTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIV 139
D I E G A GLKP + +F +A D +++ +P+
Sbjct: 445 DVGIAEQHAVTFAAGLACEGLKPFCAIYS-SFMQRAYDQVVHDVDLQK--------LPVR 495
Query: 140 FRGPNGAAAGV-GAQHSHCYASWYGSC-PGLKVLAPYSSEDARGLLKAAIR-DPDPVVFL 196
F G G H + + +C P + V+AP D ++ A+ D P F
Sbjct: 496 FAMDRAGLVGADGPTHCGAFDVTFMACLPNMIVMAPSDEADLFNMVATAVAIDDRPSCFR 555
Query: 197 ENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEG 256
G V+ + + IGK +I +EG+ V + + V L AA LE+ G
Sbjct: 556 YPR---GNGIGVALPPGNKGVPIEIGKGRILKEGERVALLGYGSAVQSCLGAAVMLEERG 612
Query: 257 ISAEVINLRSIRPLDRATINASVRKTNRLVTVEEG 291
++ V + R +PLDRA I + + L+TVEEG
Sbjct: 613 LNVTVADARFCKPLDRALIRSLAKSHEVLITVEEG 647
>At3g21500 1-D-deoxyxylulose 5-phosphate synthase, putative
Length = 628
Score = 67.8 bits (164), Expect = 9e-12
Identities = 70/288 (24%), Positives = 115/288 (39%), Gaps = 20/288 (6%)
Query: 7 NKVNLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKIS 66
+K ++L+ + + +K + AL E AD + + +G G ++
Sbjct: 322 DKYHVLKFDPETGKQFKNISKTQSYTSCFVEALIAEAEADKDIVAIHAAMG---GGTMLN 378
Query: 67 KGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKS 126
L E P R D I E G A GLKP + +F +A D +++
Sbjct: 379 --LFESRFPTRCFDVGIAEQHAVTFAAGLACEGLKPFCTIYS-SFMQRAYDQVVHDVDLQ 435
Query: 127 NYMSAGQINVPIVFRGPNGAAAGV-GAQHSHCYASWYGSC-PGLKVLAPYSSEDARGLLK 184
+P+ F G G H + + +C P + V+AP + ++
Sbjct: 436 K--------LPVRFAIDRAGLMGADGPTHCGAFDVTFMACLPNMIVMAPSDEAELFNMVA 487
Query: 185 -AAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVG 243
AA D P F + G VS + L IG+ +I R+G+ V + + V
Sbjct: 488 TAAAIDDRPSCFRYHR---GNGIGVSLPPGNKGVPLQIGRGRILRDGERVALLGYGSAVQ 544
Query: 244 FALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEG 291
L+AA L + G+ V + R +PLD A I + + L+TVEEG
Sbjct: 545 RCLEAASMLSERGLKITVADARFCKPLDVALIRSLAKSHEVLITVEEG 592
>At5g11380 1-D-deoxyxylulose 5-phosphate synthase - like protein
Length = 700
Score = 50.1 bits (118), Expect = 2e-06
Identities = 52/224 (23%), Positives = 91/224 (40%), Gaps = 24/224 (10%)
Query: 75 PERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQI 134
P+R + + E G + GLKP + F +A D +++ + +
Sbjct: 421 PDRFFNVGMAEQHAVTFSAGLSSGGLKPFC-IIPSAFLQRAYDQVVHDVDRQRKA----V 475
Query: 135 NVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPD-PV 193
I G G+ V Q ++ S P + +AP ++ ++ A D PV
Sbjct: 476 RFVITSAGLVGSDGPV--QCGAFDIAFMSSLPNMIAMAPADEDELVNMVATAAYVTDRPV 533
Query: 194 VFLENELLYGESFPVSAEVLDSSFCLP------IGKAKIEREGKDVTITAFSKMVGFALK 247
F FP +++ ++ +P IG+ ++ EG+DV + + MV L
Sbjct: 534 CF---------RFP-RGSIVNMNYLVPTGLPIEIGRGRVLVEGQDVALLGYGAMVQNCLH 583
Query: 248 AAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEG 291
A L K G++ V + R +PLD + + L+TVEEG
Sbjct: 584 AHSLLSKLGLNVTVADARFCKPLDIKLVRDLCQNHKFLITVEEG 627
>At5g63800 beta-galactosidase like protein
Length = 657
Score = 30.8 bits (68), Expect = 1.2
Identities = 34/105 (32%), Positives = 41/105 (38%), Gaps = 16/105 (15%)
Query: 109 FNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSH--CYASWYGSCP 166
F F MQ I KS + A Q I+ + N A GA H Y W G
Sbjct: 91 FKFHMQKFTAKIVDLMKSEGLYASQGGPIILSQIENEYANVEGAFHEKGASYIKWAGQMA 150
Query: 167 -GLKVLAPY---SSEDARGLLKAAIRDPDPVVFLENELLYGESFP 207
GLK P+ S DA PDPV+ N + GE+FP
Sbjct: 151 VGLKTGVPWIMCKSPDA----------PDPVINTCNGMKCGETFP 185
>At5g55530 unknown protein
Length = 405
Score = 30.0 bits (66), Expect = 2.1
Identities = 23/70 (32%), Positives = 29/70 (40%), Gaps = 15/70 (21%)
Query: 160 SWYGSCPGLKVLA--------------PYSSEDARGLLKAAIRDPDPVVFLENELLYGES 205
S+YGS P + + P SE G L I PDP V ENE + E
Sbjct: 177 SYYGSYPDVMAIPSMPSSVSIDETTKDPEGSESVPGELDK-IEFPDPNVANENEKMVSEY 235
Query: 206 FPVSAEVLDS 215
F +S +DS
Sbjct: 236 FGISCSTIDS 245
>At1g18590 unknown protein
Length = 346
Score = 28.9 bits (63), Expect = 4.6
Identities = 20/69 (28%), Positives = 30/69 (42%), Gaps = 4/69 (5%)
Query: 165 CPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKA 224
C GL PY + G KA +PD ++FL+ E + + P + + + G
Sbjct: 207 CQGLSAYGPYL-DHVLGYWKAYQANPDQILFLKYETMRADPLPYVKRLAE---FMGYGFT 262
Query: 225 KIEREGKDV 233
K E EG V
Sbjct: 263 KEEEEGNVV 271
>At5g67100 DNA polymerase alpha 1
Length = 1492
Score = 28.5 bits (62), Expect = 6.0
Identities = 20/72 (27%), Positives = 36/72 (49%), Gaps = 3/72 (4%)
Query: 222 GKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVI---NLRSIRPLDRATINAS 278
G+ K +++GK+ T K V ALKAA T+ EG + + + + ++ D+A
Sbjct: 117 GRLKKKKKGKEQTQQPQVKKVNPALKAAATITGEGRLSSMFTSSSFKKVKETDKAQYEGI 176
Query: 279 VRKTNRLVTVEE 290
+ + VT +E
Sbjct: 177 LDEIIAQVTPDE 188
>At1g74100 putative flavonol sulfotransferase
Length = 338
Score = 28.1 bits (61), Expect = 7.8
Identities = 15/43 (34%), Positives = 20/43 (45%), Gaps = 1/43 (2%)
Query: 165 CPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLYGESFP 207
C GL V PY + G KA +PD ++FL E + P
Sbjct: 199 CKGLSVYGPYL-DHVLGYWKAYQENPDRILFLRYETMRANPLP 240
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.134 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,455,257
Number of Sequences: 26719
Number of extensions: 309758
Number of successful extensions: 678
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 658
Number of HSP's gapped (non-prelim): 13
length of query: 362
length of database: 11,318,596
effective HSP length: 101
effective length of query: 261
effective length of database: 8,619,977
effective search space: 2249813997
effective search space used: 2249813997
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146972.11


