

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146972.10 - phase: 0
(106 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
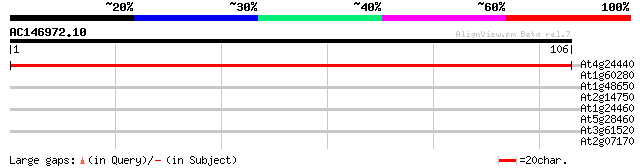
At4g24440 transcription factor IIA small subunit 198 4e-52
At1g60280 28 0.99
At1g48650 hypothetical protein 28 0.99
At2g14750 putative adenosine phosphosulfate kinase 25 4.9
At1g24460 unknown protein 25 4.9
At5g28460 putative protein 25 8.4
At3g61520 putative protein 25 8.4
At2g07170 hypothetical protein 25 8.4
>At4g24440 transcription factor IIA small subunit
Length = 106
Score = 198 bits (503), Expect = 4e-52
Identities = 96/106 (90%), Positives = 104/106 (97%)
Query: 1 MATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK 60
MATFELYRRSTIGMCLTETLDEMVQ+GTLSPE+AIQVLVQFDKSMTEALE+QVK+KVSIK
Sbjct: 1 MATFELYRRSTIGMCLTETLDEMVQSGTLSPELAIQVLVQFDKSMTEALESQVKTKVSIK 60
Query: 61 GHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLLSQ 106
GHLHTYRFCDNVWTFILQDA+FK++D QENV RVKIVACDSKLL+Q
Sbjct: 61 GHLHTYRFCDNVWTFILQDAMFKSDDRQENVSRVKIVACDSKLLTQ 106
>At1g60280
Length = 347
Score = 27.7 bits (60), Expect = 0.99
Identities = 25/104 (24%), Positives = 45/104 (43%), Gaps = 8/104 (7%)
Query: 2 ATFELYRRSTI----GMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKV 57
AT+ L + I + ++ L M+ NG P+ I+ + F+K+ E +Q V
Sbjct: 23 ATYWLIKEELIKAEDDVIISRYLKRMIVNGDSWPDHFIEDVDVFNKNPNEEFHSQSPRFV 82
Query: 58 SIKGHLHTYRFCD----NVWTFILQDALFKNEDNQENVGRVKIV 97
+K D W I +D L K+++ + +G KI+
Sbjct: 83 IVKPRTENCGRTDGCQSGCWRIIGRDKLIKSKETGKILGFKKIL 126
>At1g48650 hypothetical protein
Length = 1167
Score = 27.7 bits (60), Expect = 0.99
Identities = 15/45 (33%), Positives = 27/45 (59%), Gaps = 6/45 (13%)
Query: 5 ELYRRSTIG---MCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMT 46
E + + T+ + L LDE++QN ++P++ IQ+ +DK MT
Sbjct: 985 EFFMKPTLAYTYLSLKRELDELIQNKLVNPKLDIQL---YDKLMT 1026
>At2g14750 putative adenosine phosphosulfate kinase
Length = 276
Score = 25.4 bits (54), Expect = 4.9
Identities = 14/29 (48%), Positives = 17/29 (58%)
Query: 70 DNVWTFILQDALFKNEDNQENVGRVKIVA 98
DNV + +D FK ED EN+ RV VA
Sbjct: 138 DNVRHGLNRDLSFKAEDRAENIRRVGEVA 166
>At1g24460 unknown protein
Length = 1791
Score = 25.4 bits (54), Expect = 4.9
Identities = 11/40 (27%), Positives = 22/40 (54%)
Query: 20 LDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSI 59
++E + G+L E ++ LV+ MTE +E S +++
Sbjct: 587 MEETAERGSLEREEIVRRLVETSGLMTEGVEDHTSSDINL 626
>At5g28460 putative protein
Length = 766
Score = 24.6 bits (52), Expect = 8.4
Identities = 11/39 (28%), Positives = 21/39 (53%)
Query: 2 ATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQ 40
A F+ T G L + +DEMV+ +I +++L++
Sbjct: 692 ALFKCLNEKTQGETLLKLMDEMVEQSCEPNQITMEILME 730
>At3g61520 putative protein
Length = 766
Score = 24.6 bits (52), Expect = 8.4
Identities = 11/39 (28%), Positives = 21/39 (53%)
Query: 2 ATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQ 40
A F+ T G L + +DEMV+ +I +++L++
Sbjct: 692 ALFKCLNEKTQGETLLKLMDEMVEQSCEPNQITMEILME 730
>At2g07170 hypothetical protein
Length = 893
Score = 24.6 bits (52), Expect = 8.4
Identities = 12/40 (30%), Positives = 21/40 (52%), Gaps = 4/40 (10%)
Query: 5 ELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKS 44
+LY + G C+T+T E T +PE+ + + DK+
Sbjct: 395 DLYNEESEGSCITKTFAET----TNTPEVTYEYIPMKDKA 430
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.133 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,100,297
Number of Sequences: 26719
Number of extensions: 67658
Number of successful extensions: 169
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 164
Number of HSP's gapped (non-prelim): 8
length of query: 106
length of database: 11,318,596
effective HSP length: 82
effective length of query: 24
effective length of database: 9,127,638
effective search space: 219063312
effective search space used: 219063312
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146972.10


