

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146941.4 + phase: 0
(328 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
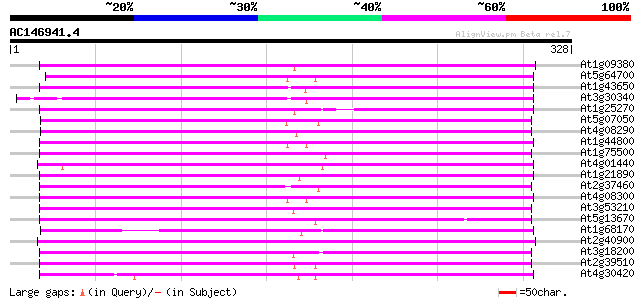
Score E
Sequences producing significant alignments: (bits) Value
At1g09380 nodulin N21 like protein 180 8e-46
At5g64700 nodulin-like protein 169 1e-42
At1g43650 nodulin-like protein 169 2e-42
At3g30340 unknown protein 159 2e-39
At1g25270 151 4e-37
At5g07050 MtN21 nodulin protein-like 150 9e-37
At4g08290 nodulin-like protein 149 2e-36
At1g44800 nodulin like protein 147 6e-36
At1g75500 nodulin-like protein 145 2e-35
At4g01440 predicted protein of unknown function 142 2e-34
At1g21890 nodulin-like protein 142 2e-34
At2g37460 nodulin-like protein 140 9e-34
At4g08300 nodulin-like protein 138 5e-33
At3g53210 unknown protein 138 5e-33
At5g13670 unknown protein 136 1e-32
At1g68170 MtN21-like protein 134 5e-32
At2g40900 putative integral membrane protein nodulin 134 9e-32
At3g18200 integral membrane protein, putative 131 6e-31
At2g39510 nodulin-like protein 131 6e-31
At4g30420 nodulin-like protein 130 1e-30
>At1g09380 nodulin N21 like protein
Length = 374
Score = 180 bits (457), Expect = 8e-46
Identities = 98/299 (32%), Positives = 153/299 (50%), Gaps = 9/299 (3%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+ VQ+ G I S++ + G L +YR + A + P+A + ER + +
Sbjct: 11 AMVLVQIGYAGMNITSKMAMEAGMKPLILVAYRQIFATIATFPVAFFLERKTRPKITLRI 70
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
L ++F G + VLY+ G++++S T A NL+P TFL A +FR E + I +
Sbjct: 71 LVQVFFCSITGATGNQVLYFVGLQNSSPTIACALTNLLPAVTFLLAAIFRQETVGIKKAS 130
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYI---------AQYHSFHSVAAHKTQMLRGTLFL 188
G+AK +G ++CV G + Y G I A+ + H ++ + G +
Sbjct: 131 GQAKVIGTLVCVIGAMVLSFYHGHTIGIGESKIHWAYAENITKHGSSSGHSNFFLGPFLI 190
Query: 189 IGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQ 248
+ A S++AWF +Q K+ E F Y +L C+M +IQ I D + W L L+
Sbjct: 191 MAAAVSWAAWFIIQTKMSETFAAPYTSTLLMCLMGSIQCGAIALISDHTISDWSLSSPLR 250
Query: 249 LITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGTY 307
I+ LY+G +++A FCL SWAM KGP Y S+F+PL LV VA +L E L GT+
Sbjct: 251 FISALYAGVVASALAFCLMSWAMQRKGPLYVSVFSPLLLVVVAIFSWALLEEKLYTGTF 309
>At5g64700 nodulin-like protein
Length = 359
Score = 169 bits (429), Expect = 1e-42
Identities = 93/297 (31%), Positives = 155/297 (51%), Gaps = 12/297 (4%)
Query: 22 VQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVLTKI 81
+Q+I T ++S+ + G F YR A + +APLA +FER S KI
Sbjct: 15 IQVIYTIMFLISKAVFNGGMNTFVFVFYRQAFATIFLAPLAFFFERKSAPPLSFVTFIKI 74
Query: 82 FLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNGRAK 141
F+ G+++ + L + TSAT A +P TF A+LF ME L + + G AK
Sbjct: 75 FMLSLFGVTLSLDLNGIALSYTSATLAAATTASLPAITFFLALLFGMERLKVKSIQGTAK 134
Query: 142 CVGAILCVAGTLAARLYKGK--------EFYIAQYHSFHSVAAH----KTQMLRGTLFLI 189
VG +C+ G + +YKG FY Q H + H T L+G + +I
Sbjct: 135 LVGITVCMGGVIILAIYKGPLLKLPLCPHFYHGQEHPHRNNPGHVSGGSTSWLKGCVLMI 194
Query: 190 GACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQL 249
+ + W +Q ++++V+P + + L C++++IQS VI ++ AW+L WNL+L
Sbjct: 195 TSNILWGLWLVLQGRVLKVYPSKLYFTTLHCLLSSIQSFVIAIALERDISAWKLGWNLRL 254
Query: 250 ITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
+ ++Y G + T + LQSW + +GP + SMF PL+L+F + A++L E +++G+
Sbjct: 255 VAVIYCGFIVTGVAYYLQSWVIEKRGPVFLSMFTPLSLLFTLLSSAILLCEIISLGS 311
>At1g43650 nodulin-like protein
Length = 341
Score = 169 bits (427), Expect = 2e-42
Identities = 88/292 (30%), Positives = 157/292 (53%), Gaps = 4/292 (1%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+FVQ++ G +LS++ + +GT F YR AA+ ++P A + E + S +
Sbjct: 8 AMVFVQIVYAGMPLLSKVAISQGTNPFVFVFYRQAFAALALSPFAFFLESSKSSPLSFIL 67
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
L KIF G+++ + LYY I +T+AT+A N IP TF+ A+LFR+E + + +
Sbjct: 68 LLKIFFISLCGLTLSLNLYYVAIENTTATFAAATTNAIPSITFVLALLFRLETVTLKKSH 127
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSF---HSVAAHKTQMLRGTLFLIGACFS 194
G AK G+++ + G L KG I Y+S + ++G++ ++ A
Sbjct: 128 GVAKVTGSMVGMLGALVFAFVKGPSL-INHYNSSTIPNGTVPSTKNSVKGSITMLAANTC 186
Query: 195 YSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITILY 254
+ W MQ K+++ +P + + LQC+ + IQSAV V+ + W++E+ L L+++ Y
Sbjct: 187 WCLWIIMQSKVMKEYPAKLRLVALQCLFSCIQSAVWAVAVNRNPSVWKIEFGLPLLSMAY 246
Query: 255 SGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
G + T + LQ WA+ KGP + +++ PLAL+ + + E +G+
Sbjct: 247 CGIMVTGLTYWLQVWAIEKKGPVFTALYTPLALILTCIVSSFLFKETFYLGS 298
>At3g30340 unknown protein
Length = 364
Score = 159 bits (402), Expect = 2e-39
Identities = 92/310 (29%), Positives = 157/310 (49%), Gaps = 14/310 (4%)
Query: 5 DVKEWFVLSSILWSMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIY 64
D K W + ++ SMI + L V ++ + ++ EG T+YR+ V + + P AI+
Sbjct: 5 DTKLWKAV--LMMSMINIGLSVVN--VMFKKMIDEGLNRMVATTYRLAVGTLFLIPFAIF 60
Query: 65 FERGQPKNFSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAI 124
ER + +L +F + +G S+V + G+ TS+T++L F N++P TF A+
Sbjct: 61 LERHNRPKLTGRILCSLFFSALLGTSLVQYFFLIGLEYTSSTFSLAFSNMVPSVTFALAL 120
Query: 125 LFRMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFH--------SVAA 176
+FR E LNI + GRAK +G ++C+ G L LYKG +++ HS H S A
Sbjct: 121 VFRQETLNIKSNVGRAKLLGTMICICGALVLTLYKGTA--LSREHSTHMETHTRTDSTGA 178
Query: 177 HKTQMLRGTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDS 236
+ G++ L+ + +S+WF +Q K+ V+P +Y + IQSA++ +
Sbjct: 179 MTQKWAMGSIMLVISIIIWSSWFIVQAKISRVYPCQYTSTTILSFFGVIQSALLSLISER 238
Query: 237 SKEAWRLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAM 296
S W ++ Q++ +LYSG + + + SW + +G + S F PL VF A
Sbjct: 239 STSMWVVKDKFQVLALLYSGIVGSGLCYVGMSWCLRQRGAVFTSSFIPLIQVFAAIFSFS 298
Query: 297 ILGEPLTVGT 306
L E + G+
Sbjct: 299 FLHEQIYCGS 308
>At1g25270
Length = 356
Score = 151 bits (382), Expect = 4e-37
Identities = 91/297 (30%), Positives = 152/297 (50%), Gaps = 19/297 (6%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+ VQ I G IL +I + +GT + L +YR+ A + + PLA+ F+R + F+ +
Sbjct: 6 AMVAVQFIFAGMFILFKITVDDGTNLKVLVAYRLSFATIFMLPLALIFQRKKRPEFTWRL 65
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
L F++G +G ++ +LY G+ TSAT++ + P+ T + ++FRME L + +
Sbjct: 66 LLLAFVSGLLGAAIPNILYLPGMARTSATFSAASSIISPLITLVLGLVFRMETLRLGSNE 125
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYI--------AQYHSFHSVAAHKTQMLRGTLFLI 189
GRAK VG +L G L YKG E +I H+ + H +L G L ++
Sbjct: 126 GRAKLVGTLLGACGALVFVFYKGIEIHIWSTHVDLLKGSHTGRATTNHHVSIL-GVLMVL 184
Query: 190 GACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQL 249
G+ K+ + YW L + ++ +I C D E W+L W++ L
Sbjct: 185 GS----------NAKIGKELGGLYWNTSLMNGVGSLVCVIIALCSDHDWEQWQLGWDINL 234
Query: 250 ITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
+ LYSG + + V L +W + KGP + ++F+P+ LV VA + L EPL +G+
Sbjct: 235 LATLYSGIVVSGMVVPLVAWCIATKGPLFVTVFSPIRLVIVALIGSFALEEPLHLGS 291
>At5g07050 MtN21 nodulin protein-like
Length = 381
Score = 150 bits (379), Expect = 9e-37
Identities = 84/300 (28%), Positives = 157/300 (52%), Gaps = 13/300 (4%)
Query: 19 MIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVL 78
MI +Q G I+++I L G + L YR +A +AP A +FER + +
Sbjct: 1 MISLQFGYAGMNIITKISLNTGMSHYVLVVYRHAIATAVIAPFAFFFERKAQPKITFSIF 60
Query: 79 TKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNG 138
++F+ G +G + YY G++ TS T++ N++P TF+ A+LFRME L++
Sbjct: 61 MQLFILGLLGPVIDQNFYYMGLKYTSPTFSCAMSNMLPAMTFILAVLFRMEMLDLKKLWC 120
Query: 139 RAKCVGAILCVAGTLAARLYKG---KEFYIAQYHSFHSVAAHKT---------QMLRGTL 186
+AK G ++ VAG + +YKG + F+ H S A+ T + L+G++
Sbjct: 121 QAKIAGTVVTVAGAMLMTIYKGPIVELFWTKYMHIQDSSHANTTSSKNSSSDKEFLKGSI 180
Query: 187 FLIGACFSYSAWFFMQVKLVEVFPLRYWGI-MLQCVMAAIQSAVIGACVDSSKEAWRLEW 245
LI A ++++ F +Q K+++ + + L C + +Q+ + ++ + AWR+ W
Sbjct: 181 LLIFATLAWASLFVLQAKILKTYAKHQLSLTTLICFIGTLQAVAVTFVMEHNPSAWRIGW 240
Query: 246 NLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVG 305
++ L+ YSG ++++ + +Q M +GP + + F+PL +V VA + +L E + +G
Sbjct: 241 DMNLLAAAYSGIVASSISYYVQGIVMKKRGPVFATAFSPLMMVIVAVMGSFVLAEKIFLG 300
>At4g08290 nodulin-like protein
Length = 384
Score = 149 bits (376), Expect = 2e-36
Identities = 88/294 (29%), Positives = 145/294 (48%), Gaps = 7/294 (2%)
Query: 19 MIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVL 78
MIF+Q GT I+ L +G + + YR LVAA+ +AP A+ FER + VL
Sbjct: 17 MIFLQFGAAGTYIVIMATLNQGQNRYVVIVYRNLVAALVLAPFALIFERKVRPKMTLSVL 76
Query: 79 TKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNG 138
KI GF+ + Y G+ TSATY +N++P TF+ A + RME +NI
Sbjct: 77 WKIMALGFLEPVLDQGFGYLGMNMTSATYTSAIMNILPSVTFIIAWILRMEKVNIAEVRS 136
Query: 139 RAKCVGAILCVAGTLAARLYKGKEFYIA-------QYHSFHSVAAHKTQMLRGTLFLIGA 191
+AK +G ++ + G L LYKG + Q + + + + GTL ++
Sbjct: 137 KAKIIGTLVGLGGALVMTLYKGPLIPLPWSNPNMDQQNGHTNNSQDHNNWVVGTLLILLG 196
Query: 192 CFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLIT 251
C ++S ++ +Q ++ +P L C+ A+QS + V+ W + W+ +L
Sbjct: 197 CVAWSGFYVLQSITIKTYPADLSLSALICLAGAVQSFAVALVVERHPSGWAVGWDARLFA 256
Query: 252 ILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVG 305
LY+G +S+ + +Q M +GP + + FNPL ++ VA + IL E + G
Sbjct: 257 PLYTGIVSSGITYYVQGMVMKTRGPVFVTAFNPLCMILVALIASFILHEQIHFG 310
>At1g44800 nodulin like protein
Length = 370
Score = 147 bits (372), Expect = 6e-36
Identities = 82/294 (27%), Positives = 154/294 (51%), Gaps = 5/294 (1%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
++I +Q G I++ + G + L +YR +VA V +AP A+ FER + +
Sbjct: 14 AIISLQFGYAGMYIITMVSFKHGMDHWVLATYRHVVATVVMAPFALMFERKIRPKMTLAI 73
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
++ G + M LYY G+++TSA+Y F N +P TF+ A++FR+E +N +
Sbjct: 74 FWRLLALGILEPLMDQNLYYIGLKNTSASYTSAFTNALPAVTFILALIFRLETVNFRKVH 133
Query: 138 GRAKCVGAILCVAGTLAARLYKGK--EFYIAQYHSFH---SVAAHKTQMLRGTLFLIGAC 192
AK VG ++ V G + LYKG E A ++SFH S + GT+ ++G+
Sbjct: 134 SVAKVVGTVITVGGAMIMTLYKGPAIEIVKAAHNSFHGGSSSTPTGQHWVLGTIAIMGSI 193
Query: 193 FSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITI 252
+++A+F +Q ++V+P + L C + I +A+ + AW++ + +
Sbjct: 194 STWAAFFILQSYTLKVYPAELSLVTLICGIGTILNAIASLIMVRDPSAWKIGMDSGTLAA 253
Query: 253 LYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
+YSG + + + +QS + +GP + + F+P+ ++ AF A++L E + +G+
Sbjct: 254 VYSGVVCSGIAYYIQSIVIKQRGPVFTTSFSPMCMIITAFLGALVLAEKIHLGS 307
>At1g75500 nodulin-like protein
Length = 389
Score = 145 bits (367), Expect = 2e-35
Identities = 83/300 (27%), Positives = 140/300 (46%), Gaps = 12/300 (4%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+ +Q G ++SR L G YR ++A + + P A + E+ + +
Sbjct: 23 AMLTLQFGYAGFHVVSRAALNMGISKLVFPVYRNIIALLLLLPFAYFLEKKERPAITLNF 82
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
L + F +G++ Y G+ +TS T+A + N +P TFL A L R+E + I+ +
Sbjct: 83 LIQFFFLALIGITANQGFYLLGLDNTSPTFASSMQNSVPAITFLMAALLRIEKVRINRRD 142
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAHKTQMLR------------GT 185
G +K +G LCVAG LYKG Y H + + +L G
Sbjct: 143 GISKILGTALCVAGASVITLYKGPTIYTPASHLHAHLLTTNSAVLAPLGNAAPKNWTLGC 202
Query: 186 LFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEW 245
++LIG C S+S W Q +++ +P R C IQ +I A + +AW
Sbjct: 203 IYLIGHCLSWSGWLVFQAPVLKSYPARLSVTSYTCFFGIIQFLIIAAFCERDSQAWVFHS 262
Query: 246 NLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVG 305
+L TILY+G +++ F +Q W + GP + +++ P+ + VA ++ LGE +G
Sbjct: 263 GWELFTILYAGIVASGIAFAVQIWCIDRGGPVFVAVYQPVQTLVVAIMASIALGEEFYLG 322
>At4g01440 predicted protein of unknown function
Length = 365
Score = 142 bits (359), Expect = 2e-34
Identities = 75/299 (25%), Positives = 147/299 (49%), Gaps = 9/299 (3%)
Query: 17 WSMIFVQLIVTGT----QILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKN 72
W+ + + +++ L + +L G + +YR+ ++ + +AP+A ++ER
Sbjct: 8 WTPVIIMVMINSALGLANALVKKVLDGGVNHMVIATYRLAISTLFLAPIAFFWERKTRPT 67
Query: 73 FSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLN 132
+ +L ++F + VG S+ + G+ TSAT A F+++ P TF+ A++FR+E LN
Sbjct: 68 LTLNILVQLFFSALVGASLTQYFFLLGLSYTSATLACAFISMTPAITFVMALIFRVEKLN 127
Query: 133 IHTWNGRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAHKTQM-----LRGTLF 187
+ + G +GA++C+ G L +YKG + H + + M + G +
Sbjct: 128 MKSKAGMGMVMGALICIGGALLLTMYKGVPLTKLRKLETHQLINNNHAMKPENWIIGCVL 187
Query: 188 LIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNL 247
L + +W +Q K+ E +P +Y ++ IQ A++ AW L L
Sbjct: 188 LFAGSSCFGSWMLIQAKVNEKYPCQYSSTVVLSFFGTIQCALLSLIKSRDITAWILTDKL 247
Query: 248 QLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
++TI+Y+GA++ SW + +GP + S+F P+ L+F + +IL + +G+
Sbjct: 248 DIVTIVYAGAVAQGICTVGTSWCIRKRGPIFTSIFTPVGLIFATLFDFLILHRQIFLGS 306
>At1g21890 nodulin-like protein
Length = 389
Score = 142 bits (359), Expect = 2e-34
Identities = 84/303 (27%), Positives = 143/303 (46%), Gaps = 14/303 (4%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+MI +Q G I++ + L G + L YR +A +AP A++ ER + +
Sbjct: 14 AMISMQFGYAGMYIITMVSLKHGMNHYVLAVYRHAIATAVIAPFALFHERKIRPKMTFRI 73
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
+I L GF+ + LYY G+ TSAT+A N++P TF+ AI+FR+E++N
Sbjct: 74 FLQIALLGFIEPVLDQNLYYVGMTYTSATFASATANVLPAITFVLAIIFRLESVNFKKVR 133
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYIAQY--------------HSFHSVAAHKTQMLR 183
AK VG ++ V+G L LYKG ++ H AA +
Sbjct: 134 SIAKVVGTVITVSGALLMTLYKGPIVDFIRFGGGGGGGSDGAGGSHGGAGAAAMDKHWIP 193
Query: 184 GTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRL 243
GTL L+G F ++ +F +Q ++ +P L C+M ++ + AW++
Sbjct: 194 GTLMLLGRTFGWAGFFILQSFTLKQYPAELSLTTLICLMGTLEGTAVSLVTVRDLSAWKI 253
Query: 244 EWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLT 303
++ L YSG + + + +Q M +GP + + FNPL +V A ++L E +
Sbjct: 254 GFDSNLFAAAYSGVICSGVAYYVQGVVMRERGPVFVATFNPLCVVITAALGVVVLSESIH 313
Query: 304 VGT 306
+G+
Sbjct: 314 LGS 316
>At2g37460 nodulin-like protein
Length = 380
Score = 140 bits (353), Expect = 9e-34
Identities = 80/295 (27%), Positives = 143/295 (48%), Gaps = 10/295 (3%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
SM+ +Q+ + G ILS+ +L +G + L YR VA + +AP A YF++ + +
Sbjct: 18 SMVVLQVGLAGMDILSKAVLNKGMSNYVLVVYRHAVATIVMAPFAFYFDKKVRPKMTLMI 77
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
KI L G + + LYY G++ T+AT+A N++P TF+ A +F +E + +
Sbjct: 78 FFKISLLGLLEPVIDQNLYYLGMKYTTATFATAMYNVLPAITFVLAYIFGLERVKLRCIR 137
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAHKT------QMLRGTLFLIGA 191
K VG + V G + L KG + V+AH T ++G + +
Sbjct: 138 STGKVVGTLATVGGAMIMTLVKGP---VLDLFWTKGVSAHNTAGTDIHSAIKGAVLVTIG 194
Query: 192 CFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVD-SSKEAWRLEWNLQLI 250
CFSY+ + +Q + +P C+M I+ + ++ + AW + W+ +L+
Sbjct: 195 CFSYACFMILQAITLRTYPAELSLTAWICLMGTIEGTAVALVMEKGNPSAWAIGWDTKLL 254
Query: 251 TILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVG 305
T YSG + +A + + M +GP + + F+PL ++ VA +I E + +G
Sbjct: 255 TATYSGIVCSALAYYVGGVVMKTRGPVFVTAFSPLCMIIVAIMSTIIFAEQMYLG 309
>At4g08300 nodulin-like protein
Length = 373
Score = 138 bits (347), Expect = 5e-33
Identities = 76/297 (25%), Positives = 148/297 (49%), Gaps = 8/297 (2%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
++I +Q G I++ + G + L +YR +VA + +AP A+ ER + +
Sbjct: 14 AIISLQFGYAGMYIITMVSFKHGMNHWILATYRHVVATIVIAPFALILERKIRPKMTWPL 73
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
+I GF+ + LYY G++ TSATY+ F+N +P TF+ A++FR+E +N+
Sbjct: 74 FLRILALGFLEPLLDQNLYYIGMKATSATYSSAFVNALPAITFIMAVIFRIETVNLKKTR 133
Query: 138 GRAKCVGAILCVAGTLAARLYKGK--EFYIAQYHSFH------SVAAHKTQMLRGTLFLI 189
AK +G + V G + LYKG E + + S H S + GTL ++
Sbjct: 134 SLAKVIGTAITVGGAMVMTLYKGPAIELFKTAHSSLHGGSSGTSSETTDQNWVTGTLAVM 193
Query: 190 GACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQL 249
G+ +++ +F +Q ++ +P +M C M + + + + AW++ +
Sbjct: 194 GSITTWAGFFILQSFTLKKYPAELSLVMWICAMGTVLNTIASLIMVRDVSAWKVGMDSGT 253
Query: 250 ITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
+ +YSG + + + +QS + +GP + + F+P+ ++ AF ++L E + +G+
Sbjct: 254 LAAVYSGVVCSGMAYYIQSIVIRERGPVFTTSFSPMCMIITAFLGVLVLAEKIHLGS 310
>At3g53210 unknown protein
Length = 369
Score = 138 bits (347), Expect = 5e-33
Identities = 86/294 (29%), Positives = 137/294 (46%), Gaps = 6/294 (2%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+ Q G ++ R L G YR +VA +AP A + E+ +
Sbjct: 13 AMVVFQTGYAGNHVIMRYALNLGVSKLVFPLYRTIVAFSVLAPSAYFLEKKERPAMKISF 72
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
L + FL G VG+++ Y +G+ +TS T+A N++P +FL A L +E + +
Sbjct: 73 LIQFFLLGLVGITLNQGFYIFGLDNTSPTFASATENVVPAVSFLMAALLGIEKVEWKRKD 132
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFY------IAQYHSFHSVAAHKTQMLRGTLFLIGA 191
G AK VG I+ VAG+L LYKG Y + Q G L L+G
Sbjct: 133 GIAKVVGTIVSVAGSLVITLYKGPTIYQPSLNIVNQTIKPEEAEEENKNWTLGCLCLMGH 192
Query: 192 CFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLIT 251
C +S+W +Q L++ +P R+ + C A IQ I A + E W++ +L
Sbjct: 193 CLCWSSWIVLQSPLLKKYPARFSFVSYSCFFAVIQFFGISAYFERDLERWKIISGGELYA 252
Query: 252 ILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVG 305
+LY+G + +A VF +Q + + GP + S + PL + A + LGE +G
Sbjct: 253 LLYTGLVGSAMVFAIQIYVVERGGPLFVSAYLPLQTLIAAVLATLALGEHFYLG 306
>At5g13670 unknown protein
Length = 377
Score = 136 bits (343), Expect = 1e-32
Identities = 74/297 (24%), Positives = 148/297 (48%), Gaps = 10/297 (3%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+++F+Q + I++++ L +G L +YR+ VA+ + P A+ ER + ++
Sbjct: 11 AIVFIQCLYALMSIVAKLALNKGMSPHVLVAYRMAVASALITPFALILERNTRPKLTFKI 70
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
L +I + + LYY G++ T+AT+ N +P TF+ A +F++E + I +
Sbjct: 71 LLQIAILSLFEPVVEQNLYYSGMKLTTATFTSALCNALPAMTFIMACVFKLEKVTIERRH 130
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAH--------KTQMLRGTLFLI 189
+AK VG ++ + G + KG + + + H + + RG++ L+
Sbjct: 131 SQAKLVGTMVAIGGAMLMTFVKGNVIELPWTSNSRGLNGHTHAMRIPKQADIARGSIMLV 190
Query: 190 GACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVD-SSKEAWRLEWNLQ 248
+CFS+S + +Q K++ + L C+M +++ V+G + + W++ ++
Sbjct: 191 ASCFSWSCYIILQAKILAQYKAELSLTALMCIMGMLEATVMGLIWERKNMSVWKINPDVT 250
Query: 249 LITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVG 305
L+ +Y G +S A + + WA +GP + S FNPL++V VA + E + VG
Sbjct: 251 LLASIYGGLVSGLAYYVI-GWASKERGPVFVSAFNPLSMVLVAILSTFVFLEKVYVG 306
>At1g68170 MtN21-like protein
Length = 329
Score = 134 bits (338), Expect = 5e-32
Identities = 83/297 (27%), Positives = 138/297 (45%), Gaps = 31/297 (10%)
Query: 19 MIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVL 78
M+ VQ+ G I ++ + +G L +YR+L A + + P+ F+
Sbjct: 1 MVVVQIATAGLNIFFKLAMEDGMNPSVLVAYRLLFATLFMIPICFIFQ------------ 48
Query: 79 TKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNG 138
+ + +L G+ TSAT+ L P+ TF+ A L RME++ + + G
Sbjct: 49 ---------SVVIPSILTITGLALTSATFTSAAGVLTPLVTFIFAALLRMESVRLGSSVG 99
Query: 139 RAKCVGAILCVAGTLAARLYKGKEFYIAQYH---------SFHSVAAHKTQMLRGTLFLI 189
AK G + V G L Y+G E + H S H +L G L +
Sbjct: 100 LAKVFGTLFGVGGALVFIFYRGIEIRLWSTHVNLVNQPRDSSRDATTHHISIL-GALLVF 158
Query: 190 GACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQL 249
G S S WF +QVK+ + F YW L +M + + ++ C + + WRL WN++L
Sbjct: 159 GGNISISLWFLLQVKISKQFGGPYWNATLMNMMGGVVAMLVALCWEHDLDEWRLGWNIRL 218
Query: 250 ITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
+TI Y+ L + V + +W + +GP + S+F+P+ LV VA + +L E L +G+
Sbjct: 219 LTIAYAAILISGMVVAVNAWCIESRGPLFVSVFSPVGLVIVALVGSFLLDETLHLGS 275
>At2g40900 putative integral membrane protein nodulin
Length = 400
Score = 134 bits (336), Expect = 9e-32
Identities = 73/291 (25%), Positives = 144/291 (49%)
Query: 17 WSMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCE 76
++M+ +Q G ++++ +L G + L +YR A +AP A+ ER +
Sbjct: 13 FAMVCLQFGYAGMNLVTKTVLDRGMSHYVLVAYRNAFATAAIAPFALLSERKVRSKMTFP 72
Query: 77 VLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTW 136
+ +IFL +G + LYY G++ TS T++ N++P T + A LFRME + +
Sbjct: 73 IFMRIFLLALLGPVIDQNLYYIGLKLTSPTFSSAVSNIVPAITIILATLFRMEKVEMRKV 132
Query: 137 NGRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAHKTQMLRGTLFLIGACFSYS 196
K +G ++ V G++ YKG + H + + L+ +FL+ A S++
Sbjct: 133 RCLVKVMGTLVTVVGSILMIFYKGPFINFFRSHLTAASSPPTADYLKAAVFLLLASLSWA 192
Query: 197 AWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITILYSG 256
++F +Q ++ + + C M +QS + ++ + A + +++ L+ Y+G
Sbjct: 193 SFFVLQAATLKKYSAHLSMSTMVCFMGTLQSLALAFVMEHNPSALNIGFDMNLLASAYAG 252
Query: 257 ALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGTY 307
+S++ + +Q M KGP + + FNPL +V V+ +LG+ + +G Y
Sbjct: 253 IMSSSIAYYVQGLMMQRKGPVFVTAFNPLIVVIVSIMSFFVLGQGIYLGGY 303
>At3g18200 integral membrane protein, putative
Length = 360
Score = 131 bits (329), Expect = 6e-31
Identities = 78/293 (26%), Positives = 142/293 (47%), Gaps = 7/293 (2%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
++I +Q G I+SR+ L G YR L+A + + P A +FE+ + + +
Sbjct: 15 ALITLQFCFAGFHIVSRVALNIGVSKVVYPVYRNLLALLLIGPFAYFFEKKERPPLTISL 74
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
L + F +G++ Y G+ + T+A N +P TF+ A R+E++++ +
Sbjct: 75 LAQFFFLALIGITANQGFYLLGLYYATPTFASAMQNSVPAITFIMACALRLEHIDLVRKH 134
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFY-----IAQYHSFHSVAAHKTQMLRGTLFLIGAC 192
G AK +G ++ + G LY+G + + + S +H + G L+L+G C
Sbjct: 135 GVAKVLGTLVSIGGATVITLYRGFPIFDQGLNMQKEEVVGSDNSHSLTL--GWLYLMGHC 192
Query: 193 FSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITI 252
S++ W +Q +++ +P + C IQ VI V++ W + +L TI
Sbjct: 193 LSWAGWMVLQAPVLKQYPAKLTLTSFTCFFGLIQFLVIALFVETDLNNWIIVSWEELFTI 252
Query: 253 LYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVG 305
LY+G +++ V LQ+W + GP + ++F PL + VA +ILG+ L G
Sbjct: 253 LYAGIIASGLVVYLQTWCIYKSGPVFVAVFQPLQTLLVAAMAFLILGDQLYSG 305
>At2g39510 nodulin-like protein
Length = 374
Score = 131 bits (329), Expect = 6e-31
Identities = 74/294 (25%), Positives = 149/294 (50%), Gaps = 6/294 (2%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+++ +Q G I+++ L +G L SYR +VA + +AP A + +R + +
Sbjct: 11 TVVSLQFGYAGLSIIAKFALNQGMSPHVLASYRHIVATIFIAPFAYFLDRKIRPKMTLSI 70
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
KI L G + ++ LYY G++ TSAT+ N++P F+ A +FR+E +N+ +
Sbjct: 71 FFKILLLGLLEPTIDQNLYYTGMKYTSATFTAAMTNVLPAFAFIMAWIFRLEKVNVKKIH 130
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYI--AQYHSFHSVAAH---KTQMLRGTLFLIGAC 192
+AK +G I+ V G + + KG + A H H +++ K + +G + C
Sbjct: 131 SQAKILGTIVTVGGAMLMTVVKGPLIPLPWANPHDIHQDSSNTGVKQDLTKGASLIAIGC 190
Query: 193 FSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVD-SSKEAWRLEWNLQLIT 251
++ + +Q ++ +P+ C + +I+S ++ ++ + AW + + +L+
Sbjct: 191 ICWAGFINLQAITLKSYPVELSLTAYICFLGSIESTIVALFIERGNPSAWAIHLDSKLLA 250
Query: 252 ILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVG 305
+Y G + + + +Q M +GP + + FNPL++V VA ++IL E + +G
Sbjct: 251 AVYGGVICSGIGYYVQGVIMKTRGPVFVTAFNPLSMVIVAILGSIILAEVMFLG 304
>At4g30420 nodulin-like protein
Length = 373
Score = 130 bits (326), Expect = 1e-30
Identities = 87/298 (29%), Positives = 138/298 (46%), Gaps = 10/298 (3%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPK----NF 73
+M +QL G + +R LV G YR A + + P +Y R + K +
Sbjct: 2 AMTMIQLCYAGVTLFARATLVHGLSPRVFILYRQAFATIFIFPF-LYLSRRKSKIAISSL 60
Query: 74 SCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNI 133
+ + IFL +G+++ LY G+ TS++ N+IP TFL + L E LN+
Sbjct: 61 DLKSFSLIFLVSLIGITINQNLYLEGLYLTSSSMGSAVGNIIPAITFLISFLAGYEKLNL 120
Query: 134 HTWNGRAKCVGAILCVAGTLAARLYKGKEFYIAQ--YHSFHSVAAH---KTQMLRGTLFL 188
G AK G ILCVAG ++ L +G + ++ SV H + L G LFL
Sbjct: 121 RDIRGLAKIAGTILCVAGAISMTLLRGPKILNSESALPIAKSVLGHLKDQNTWLIGCLFL 180
Query: 189 IGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQ 248
+ +S W +QV + +P C+ IQ AV+ ++ AW L +
Sbjct: 181 FSSTLCWSFWLILQVPISAYYPDNLSLSAWMCLFGTIQCAVVTFFLEKDPNAWILHSYSE 240
Query: 249 LITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
T LY+G ++A F +Q+WA+ +GP + ++FNPL V V A+ E + G+
Sbjct: 241 FATCLYAGIGASALSFTVQAWAIAKRGPVFSALFNPLCTVIVTILAALFFHEEIYTGS 298
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.332 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,563,726
Number of Sequences: 26719
Number of extensions: 242867
Number of successful extensions: 728
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 49
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 616
Number of HSP's gapped (non-prelim): 60
length of query: 328
length of database: 11,318,596
effective HSP length: 100
effective length of query: 228
effective length of database: 8,646,696
effective search space: 1971446688
effective search space used: 1971446688
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146941.4


