

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146866.2 + phase: 0
(504 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
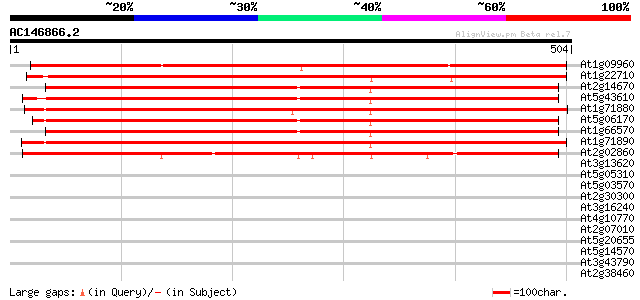
Score E
Sequences producing significant alignments: (bits) Value
At1g09960 putative sucrose/H+ symporter 700 0.0
At1g22710 putative sucrose transport protein, SUC2 530 e-150
At2g14670 putative sucrose-proton symporter 525 e-149
At5g43610 sucrose transporter protein 523 e-148
At1g71880 sucrose transport protein SUC1 522 e-148
At5g06170 sucrose transporter protein 521 e-148
At1g66570 hypothetical protein 519 e-147
At1g71890 putative sucrose transport protein 499 e-141
At2g02860 Sucrose transporter (suc3) 457 e-129
At3g13620 unknown protein 38 0.015
At5g05310 unknown protein 33 0.37
At5g03570 transporter like protein 33 0.48
At2g30300 hypothetical protein 31 1.4
At3g16240 delta tonoplast integral protein (delta-TIP) 31 1.8
At4g10770 oligopeptide transporter like protein 30 4.1
At2g07010 putative retroelement pol polyprotein 30 4.1
At5g20655 24 kDa vacuolar protein - like 29 5.3
At5g14570 high affinity nitrate transporter - like protein 29 5.3
At3g43790 transporter-like protein 29 5.3
At2g38460 unknown protein 29 5.3
>At1g09960 putative sucrose/H+ symporter
Length = 510
Score = 700 bits (1806), Expect = 0.0
Identities = 340/486 (69%), Positives = 403/486 (81%), Gaps = 6/486 (1%)
Query: 19 STSTSRPV-QPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIW 77
STS+SRPV P + + R LLRVASVA GIQFGWALQLSLLTPYVQ+LGIPH WAS+IW
Sbjct: 23 STSSSRPVVSPPRSKVSKRVLLRVASVACGIQFGWALQLSLLTPYVQELGIPHAWASVIW 82
Query: 78 LCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDI 137
LCGP+SGLFVQPLVGH SDRC+S++GRRRPFI+ GA +I ++V++IG+AADIG+ GD
Sbjct: 83 LCGPLSGLFVQPLVGHSSDRCTSKYGRRRPFIVAGAVAISISVMVIGHAADIGWAFGDR- 141
Query: 138 TQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGN 197
+P AIV FV+GFWILDVANN+TQGPCRALLADLT ND RRTRVAN YFSLFMAVGN
Sbjct: 142 EGKIKPRAIVAFVLGFWILDVANNMTQGPCRALLADLTENDNRRTRVANGYFSLFMAVGN 201
Query: 198 ILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSS 257
+LGYATGSY+GWYKIFTFT T AC++ CANLKSAF++DV FI +TT LS+ +AHEVPL+S
Sbjct: 202 VLGYATGSYNGWYKIFTFTKTVACNVECANLKSAFYIDVVFIAITTILSVSAAHEVPLAS 261
Query: 258 SGA---GESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREI 314
+ G++ +EAF+ E+FGTF+YF VWI+L VTALTWIGWFPF LFDTDWMGREI
Sbjct: 262 LASEAHGQTSGTDEAFLSEIFGTFRYFPGNVWIILLVTALTWIGWFPFILFDTDWMGREI 321
Query: 315 YGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICF 374
YGG+P G Y GV MGALGL+LNSV L +TS+LME+LCRK GAGFVWGISNI MAICF
Sbjct: 322 YGGEPNIGTSYSAGVSMGALGLMLNSVFLGITSVLMEKLCRKWGAGFVWGISNILMAICF 381
Query: 375 IAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPLG 434
+ M++ ++ A+ +GY+ Q PP IV AA+ IFTILG P+AITYSVPYALIS IE LG
Sbjct: 382 LGMIITSFVASHLGYIGHEQ-PPASIVFAAVLIFTILGIPLAITYSVPYALISIRIESLG 440
Query: 435 LGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIPR 494
LGQGLS+GVLNLAIV+PQ++VS+GSGPWDQLFGGGNSPA AV A + G++A+LA+PR
Sbjct: 441 LGQGLSLGVLNLAIVIPQVIVSVGSGPWDQLFGGGNSPALAVGAATGFIGGIVAILALPR 500
Query: 495 TRTQKP 500
TR QKP
Sbjct: 501 TRIQKP 506
>At1g22710 putative sucrose transport protein, SUC2
Length = 512
Score = 530 bits (1364), Expect = e-150
Identities = 262/494 (53%), Positives = 344/494 (69%), Gaps = 13/494 (2%)
Query: 16 SSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASI 75
S+ T T QP + LR+++ V+S+A+G+QFGWALQLSLLTPYVQ LGIPHKWAS+
Sbjct: 14 SALETQTGELDQPER----LRKIISVSSIAAGVQFGWALQLSLLTPYVQLLGIPHKWASL 69
Query: 76 IWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGD 135
IWLCGP+SG+ VQP+VG+ SDRC+SRFGRRRPFI+ GA + VAV +IGYAADIG+ +GD
Sbjct: 70 IWLCGPISGMLVQPIVGYHSDRCTSRFGRRRPFIVAGAGLVTVAVFLIGYAADIGHSMGD 129
Query: 136 DITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAV 195
+ + + AI +F +GFWILDVANN QGPCRA LADL+ +A++TR ANA+FS FMAV
Sbjct: 130 QLDKPPKTRAIAIFALGFWILDVANNTLQGPCRAFLADLSAGNAKKTRTANAFFSFFMAV 189
Query: 196 GNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPL 255
GN+LGYA GSY YK+ FT+T +C + CANLK+ FFL + +++ T++S+ E P
Sbjct: 190 GNVLGYAAGSYRNLYKVVPFTMTESCDLYCANLKTCFFLSITLLLIVTFVSLCYVKEKPW 249
Query: 256 SSSGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIY 315
+ + ++ F E+FG FK P+W++L VTAL WI WFPF LFDTDWMGRE+Y
Sbjct: 250 TPEPTADGKASNVPFFGEIFGAFKELKRPMWMLLIVTALNWIAWFPFLLFDTDWMGREVY 309
Query: 316 GGDPEGGL------IYDTGVRMGALGLLLNSVVLAVTSLLMERLCRK-RGAGFVWGISNI 368
GG+ + +Y+ GVR GALGL+LN++VL SL +E + RK GA +WGI N
Sbjct: 310 GGNSDATATAASKKLYNDGVRAGALGLMLNAIVLGFMSLGVEWIGRKLGGAKRLWGIVNF 369
Query: 369 FMAICFIAMLVLTYAANSIGYVSKGQP--PPTGIVIAALAIFTILGFPMAITYSVPYALI 426
+AIC +V+T A + G PP + AL +F ILG P AIT+S+P+AL
Sbjct: 370 ILAICLAMTVVVTKQAENHRRDHGGAKTGPPGNVTAGALTLFAILGIPQAITFSIPFALA 429
Query: 427 STHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGL 486
S G GQGLS+GVLNLAIVVPQ+V+S+G GP+D+LFGGGN PAF + A+AA +SG+
Sbjct: 430 SIFSTNSGAGQGLSLGVLNLAIVVPQMVISVGGGPFDELFGGGNIPAFVLGAIAAAVSGV 489
Query: 487 LALLAIPRTRTQKP 500
LAL +P P
Sbjct: 490 LALTVLPSPPPDAP 503
>At2g14670 putative sucrose-proton symporter
Length = 492
Score = 525 bits (1352), Expect = e-149
Identities = 257/467 (55%), Positives = 333/467 (71%), Gaps = 8/467 (1%)
Query: 33 TPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVG 92
+PLR+++ VAS+A+GIQFGWALQLSLLTPYVQ LG+PHKW+S IWLCGPVSGL VQP VG
Sbjct: 28 SPLRKMISVASIAAGIQFGWALQLSLLTPYVQLLGVPHKWSSFIWLCGPVSGLLVQPSVG 87
Query: 93 HLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAIVVFVIG 152
+ SDRC+SRFGRRRPFI GA + VAVV+IGYAAD G+ +GD I + + A+V+F +G
Sbjct: 88 YFSDRCTSRFGRRRPFIATGALLVAVAVVLIGYAADFGHSMGDKIDKPVKMRAVVIFALG 147
Query: 153 FWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSYSGWYKI 212
FWILDVANN QGPCRA L DL DA++TR ANA+FS FMAVGN+LGYA GSY+ YKI
Sbjct: 148 FWILDVANNTLQGPCRAFLGDLAAGDAKKTRTANAFFSFFMAVGNVLGYAAGSYTNLYKI 207
Query: 213 FTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSSGAGESGSAEEAFMW 272
F FT+T AC I CANLKS FFL + ++V T +++ + S +S + + F
Sbjct: 208 FPFTMTKACDIYCANLKSCFFLSITLLLVVTIIALWYVEDKQWSPK--ADSDNEKTPFFG 265
Query: 273 ELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGDPEGG----LIYDTG 328
E+FG FK P+W++L VTAL WI WFPF L+DTDWMGRE+YGGD +G +Y+ G
Sbjct: 266 EIFGAFKVMKRPMWMLLIVTALNWIAWFPFLLYDTDWMGREVYGGDSKGDDKMKKLYNQG 325
Query: 329 VRMGALGLLLNSVVLAVTSLLMERLCRK-RGAGFVWGISNIFMAICFIAMLVLTYAANSI 387
+ +GALGL+LNS+VL + SL +E + +K GA +WG NI +A+C +++T A
Sbjct: 326 IHVGALGLMLNSIVLGIVSLGIEGISKKIGGAKRLWGAVNIILAVCLAMTVLVTKKAEEH 385
Query: 388 GYVSKGQPPPT-GIVIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQGLSMGVLNL 446
++ PT GI AL +F +LG P+AIT+S+P+AL S G GQGLS+GVLN+
Sbjct: 386 RRIAGPMALPTDGIRAGALTLFALLGIPLAITFSIPFALASIISSSSGAGQGLSLGVLNM 445
Query: 447 AIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIP 493
AIV+PQ++VS G GP D LFGGGN P F V A+AA +S ++A +P
Sbjct: 446 AIVIPQMIVSFGVGPIDALFGGGNLPRFVVGAIAAAISSVVAFTVLP 492
>At5g43610 sucrose transporter protein
Length = 492
Score = 523 bits (1346), Expect = e-148
Identities = 260/488 (53%), Positives = 338/488 (68%), Gaps = 15/488 (3%)
Query: 12 SRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHK 71
+R SS+S + P +P+R+++ VAS+A+GIQFGWALQLSLLTPYVQ LG+PHK
Sbjct: 14 NRQSSSSSADLNGP-------SPMRKMISVASIAAGIQFGWALQLSLLTPYVQLLGVPHK 66
Query: 72 WASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGY 131
W+S IWLCGPVSGL VQP VG+ SDRC SRFGRRRPFI +GA + VAVV+IGYAAD G+
Sbjct: 67 WSSFIWLCGPVSGLLVQPSVGYFSDRCKSRFGRRRPFIAMGALLVAVAVVLIGYAADFGH 126
Query: 132 LIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSL 191
+GD + + + A+V+F +GFWILDVANN QGPCRA L DL DA++TR ANA+FS
Sbjct: 127 SMGDKVDEPVKMRAVVIFALGFWILDVANNTLQGPCRAFLGDLAAGDAKKTRTANAFFSF 186
Query: 192 FMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAH 251
FMAVGN+LGYA GSY+ YKIF FT+T AC I CANLKS FFL + ++V T +++
Sbjct: 187 FMAVGNVLGYAAGSYTNLYKIFPFTMTKACDIYCANLKSCFFLSITLLLVVTIIALWYVE 246
Query: 252 EVPLSSSGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMG 311
+ S +S + + F E+FG FK P+W++L VTAL WI WFPF L+DTDWMG
Sbjct: 247 DKQWSPK--ADSDNEKTPFFGEIFGAFKVMKRPMWMLLIVTALNWIAWFPFLLYDTDWMG 304
Query: 312 REIYGGDPEGG----LIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKR-GAGFVWGIS 366
RE+YGGD +G +Y+ G+ +G LGL+LNS+VL SL +E + RK GA +WG
Sbjct: 305 REVYGGDSKGDDKMKKLYNQGIHVGGLGLMLNSIVLGFMSLGIEGISRKMGGAKRLWGAV 364
Query: 367 NIFMAICFIAMLVLTYAANSIGYVSKGQPPPT-GIVIAALAIFTILGFPMAITYSVPYAL 425
NI +A+C +++T A ++ PT GI AL +F +LG P+AIT+S+P+AL
Sbjct: 365 NIILAVCLAMTVLVTKKAEEHRRIAGPMALPTDGIRAGALTLFALLGIPLAITFSIPFAL 424
Query: 426 ISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSG 485
S G GQGLS+GVLN+ IV+PQ+VVS G GP D LFGGGN P F V A+AA +S
Sbjct: 425 ASIISSSSGAGQGLSLGVLNMTIVIPQMVVSFGVGPIDALFGGGNLPGFVVGAIAAAISS 484
Query: 486 LLALLAIP 493
++A +P
Sbjct: 485 VVAFSVLP 492
>At1g71880 sucrose transport protein SUC1
Length = 513
Score = 522 bits (1344), Expect = e-148
Identities = 263/497 (52%), Positives = 342/497 (67%), Gaps = 10/497 (2%)
Query: 14 TRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWA 73
T+ + + T P QP +PLR+++ VAS+A+G+QFGWALQLSLLTPYVQ LGIPHKW+
Sbjct: 10 TKDAAALETQSPEDFDQP-SPLRKIISVASIAAGVQFGWALQLSLLTPYVQLLGIPHKWS 68
Query: 74 SIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLI 133
S+IWLCGPVSG+ VQP+VG SDRC S+FGRRRPFI GAA + VAV +IGYAAD GY +
Sbjct: 69 SLIWLCGPVSGMIVQPIVGFHSDRCRSKFGRRRPFIATGAALVAVAVFLIGYAADFGYKM 128
Query: 134 GDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFM 193
GD + + + AI +F +GFWILDVANN QGPCRA LADL DA+RTRVANA+FS FM
Sbjct: 129 GDKLEEKVKVRAIGIFALGFWILDVANNTLQGPCRAFLADLAAGDAKRTRVANAFFSFFM 188
Query: 194 AVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHE- 252
AVGN+LGYA GSY+ +K+F FT+T AC I CANLK+ FFL + +++ T S+ ++
Sbjct: 189 AVGNVLGYAAGSYTNLHKMFPFTMTKACDIYCANLKTCFFLSITLLLIVTVTSLWYVNDK 248
Query: 253 --VPLSSSGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWM 310
P + + ++ E+FG FK P+W++L VTAL WI WFPF LFDTDWM
Sbjct: 249 QWSPPPRNADDDEKTSSVPLFGEIFGAFKVMKRPMWMLLIVTALNWIAWFPFLLFDTDWM 308
Query: 311 GREIYGGDPEGG----LIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRK-RGAGFVWGI 365
GRE++GGD +G +Y GV+ GA+GL+ NS+VL SL +E + RK GA +WGI
Sbjct: 309 GREVFGGDSDGNERSKKLYSLGVQSGAMGLMFNSIVLGFMSLGVEWIGRKLGGAKRLWGI 368
Query: 366 SNIFMAI-CFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYA 424
N +A + +LV +A + P + AL++F +LG P+AIT+S P+A
Sbjct: 369 VNFILAAGLAMTVLVTKFAEDHRKTAGDLAGPSASVKAGALSLFAVLGIPLAITFSTPFA 428
Query: 425 LISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLS 484
L S G GQGLS+GVLNLAIV+PQ++VSLG GP+D LFGGGN PAF VAA+AA +S
Sbjct: 429 LASIFSSCSGAGQGLSLGVLNLAIVIPQMIVSLGGGPFDALFGGGNLPAFIVAAIAAAIS 488
Query: 485 GLLALLAIPRTRTQKPR 501
G+LAL +P P+
Sbjct: 489 GVLALTVLPSPPPDAPK 505
>At5g06170 sucrose transporter protein
Length = 491
Score = 521 bits (1341), Expect = e-148
Identities = 254/478 (53%), Positives = 340/478 (70%), Gaps = 8/478 (1%)
Query: 21 STSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCG 80
S+S V P +P +PLR+++ VAS+A+GIQFGWALQLSLLTPYVQ LG+PHKW+S IWLCG
Sbjct: 17 SSSSVVVPDEP-SPLRKMISVASIAAGIQFGWALQLSLLTPYVQLLGVPHKWSSFIWLCG 75
Query: 81 PVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQN 140
P+SGL VQP VG+ SDRC SRFGRRRPFI GA + +AV++IG+AAD G+ +GD + +
Sbjct: 76 PISGLLVQPTVGYFSDRCKSRFGRRRPFIATGALLVALAVILIGFAADFGHTMGDKLDEA 135
Query: 141 YRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILG 200
+ A+ FV+GFWILDVANN QGPCRA L DL DA++TR ANA FS FMAVGN+LG
Sbjct: 136 VKIRAVGFFVVGFWILDVANNTLQGPCRAFLGDLAAGDAKKTRTANAIFSFFMAVGNVLG 195
Query: 201 YATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSSGA 260
YA GSY+ +KIF FT+T AC I CANLKS F + + ++V T +++ + S +
Sbjct: 196 YAAGSYTNLHKIFPFTVTKACDIYCANLKSCFIISITLLIVLTIIALWYVEDKQWSPN-- 253
Query: 261 GESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGDPE 320
+S + + F E+FG FK P+W++L+VTAL WI WFPF L+DTDWMGRE+YGGD
Sbjct: 254 ADSDNEKTPFFGEIFGAFKVMKRPMWMLLAVTALNWIAWFPFLLYDTDWMGREVYGGDSA 313
Query: 321 GG----LIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICFIA 376
G +Y+ G+++G+LGL+LNS+VL V SL++ + +K GA +WG NI +A+C
Sbjct: 314 GDDKMKKLYNHGIQVGSLGLMLNSIVLGVMSLVIGVISKKIGAKRLWGAVNIILAVCLAM 373
Query: 377 MLVLTYAANSIGYVSKGQPPPTGIV-IAALAIFTILGFPMAITYSVPYALISTHIEPLGL 435
+++T A ++ PT + AL++F ILG P+AIT+S+P+AL S G
Sbjct: 374 TVLVTKKAEEHRKIAGRMALPTNAIRDGALSLFAILGIPLAITFSIPFALASIISSSSGA 433
Query: 436 GQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIP 493
GQGLS+GVLN+AIV+PQ++VS G GP D LFGGGN P F V A+AAL+S ++AL +P
Sbjct: 434 GQGLSLGVLNMAIVIPQMIVSFGVGPIDALFGGGNLPGFVVGAIAALISSVVALTVLP 491
>At1g66570 hypothetical protein
Length = 491
Score = 519 bits (1336), Expect = e-147
Identities = 256/467 (54%), Positives = 330/467 (69%), Gaps = 8/467 (1%)
Query: 33 TPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVG 92
+PLR+++ VAS+A+GIQFGWALQLSLLTPYVQ LG+PHKW S IWLCGPVSGL VQP VG
Sbjct: 27 SPLRKMISVASIAAGIQFGWALQLSLLTPYVQLLGVPHKWPSFIWLCGPVSGLLVQPSVG 86
Query: 93 HLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAIVVFVIG 152
+ SDRC+SRFGRRRPFI GA + V+VV+IGYAAD G+ +GD I + + A+V+F +G
Sbjct: 87 YFSDRCTSRFGRRRPFIATGALLVAVSVVLIGYAADFGHSMGDKIDKPVKMRAVVIFALG 146
Query: 153 FWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSYSGWYKI 212
FWILDVANN QGPCRA L DL DA++TR ANA+FS FMAVGN+LGYA GSY+ YKI
Sbjct: 147 FWILDVANNTLQGPCRAFLGDLAAGDAQKTRTANAFFSFFMAVGNVLGYAAGSYTNLYKI 206
Query: 213 FTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSSGAGESGSAEEAFMW 272
F FT+T AC I CANLKS FFL + ++V T +++ + S +S + + F
Sbjct: 207 FPFTMTKACDIYCANLKSCFFLSITLLLVVTIIALWYVEDKQWSPK--ADSDNEKTPFFG 264
Query: 273 ELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGDPEGG----LIYDTG 328
E+FG FK P+W++L VTAL WI WFPF L+DTDWMGRE+YGGD +G +Y+ G
Sbjct: 265 EIFGAFKVMKRPMWMLLIVTALNWIAWFPFLLYDTDWMGREVYGGDSKGDDKMKKLYNQG 324
Query: 329 VRMGALGLLLNSVVLAVTSLLMERLCRKR-GAGFVWGISNIFMAICFIAMLVLTYAANSI 387
+ +GALGL+LNS+VL V SL +E + RK GA +WG NI +A+C +++T A
Sbjct: 325 IHVGALGLMLNSIVLGVMSLGIEGISRKMGGAKRLWGAVNIILAVCLAMTVLVTKKAEEH 384
Query: 388 GYVSKGQPPPT-GIVIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQGLSMGVLNL 446
++ PT GI AL +F +LG P+AIT+S+P+AL S G GQ LS+GVLN+
Sbjct: 385 RRIAGPMALPTDGIRAGALTLFALLGIPLAITFSIPFALASIISSSSGAGQRLSLGVLNM 444
Query: 447 AIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIP 493
AIV+PQ++VS G GP D LFG GN P F V A+AA +S ++A +P
Sbjct: 445 AIVIPQMIVSFGVGPIDALFGDGNLPGFVVGAIAAAVSSIVAFTVLP 491
>At1g71890 putative sucrose transport protein
Length = 512
Score = 499 bits (1285), Expect = e-141
Identities = 249/497 (50%), Positives = 337/497 (67%), Gaps = 8/497 (1%)
Query: 11 RSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPH 70
R+ ++ + S P QP +PLR+++ VAS+A+G+QFGWALQLSLLTPY+Q LGIPH
Sbjct: 8 RAANNATALETQSSPEDLGQP-SPLRKIISVASIAAGVQFGWALQLSLLTPYIQLLGIPH 66
Query: 71 KWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIG 130
KW+S +WLCGP+SG+ VQP+VG+ SDRC SRFGRRRPFI G A + V+V +IG+AAD+G
Sbjct: 67 KWSSYMWLCGPISGMIVQPIVGYHSDRCESRFGRRRPFIAAGVALVAVSVFLIGFAADMG 126
Query: 131 YLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFS 190
+ GD + R AI++F+ GFW LDVANN QGPCRA LADL DA++TRVANA FS
Sbjct: 127 HSFGDKLENKVRTRAIIIFLTGFWFLDVANNTLQGPCRAFLADLAAGDAKKTRVANACFS 186
Query: 191 LFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSA 250
FMAVGN+LGYA GSY+ +K+F FT+T AC I CANLK+ FFL + +++ T+ S+
Sbjct: 187 FFMAVGNVLGYAAGSYTNLHKMFPFTMTKACDIYCANLKTCFFLSITLLLIVTFSSLWYV 246
Query: 251 HEVPLS-SSGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDW 309
+ S G E ++ F E+FG ++ P+ ++L VT + WI WFPF L+DTDW
Sbjct: 247 KDKQWSPPQGDKEEKTSSLFFFGEIFGAVRHMKRPMVMLLIVTVINWIAWFPFILYDTDW 306
Query: 310 MGREIYGGDPEGG----LIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKR-GAGFVWG 364
MGRE+YGG+ +G +YD GV+ GALGL+ NS++L SL +E + RK GA +WG
Sbjct: 307 MGREVYGGNSDGDERSKKLYDQGVQAGALGLMFNSILLGFVSLGVESIGRKMGGAKRLWG 366
Query: 365 ISNIFMAICFIAMLVLTYAANSIGYVSKG-QPPPTGIVIAALAIFTILGFPMAITYSVPY 423
N +AI +++T +A ++ P +GI ++FT+LG P+AITYS+P+
Sbjct: 367 CVNFILAIGLAMTVLVTKSAEHHREIAGPLAGPSSGIKAGVFSLFTVLGIPLAITYSIPF 426
Query: 424 ALISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALL 483
AL S G GQGLS+GVLN+AI +PQ++VS SGP D FGGGN P+F V A+AA +
Sbjct: 427 ALASIFSTNSGAGQGLSLGVLNIAICIPQMIVSFSSGPLDAQFGGGNLPSFVVGAIAAAV 486
Query: 484 SGLLALLAIPRTRTQKP 500
SG+LAL +P P
Sbjct: 487 SGVLALTVLPSPPPDAP 503
>At2g02860 Sucrose transporter (suc3)
Length = 594
Score = 457 bits (1176), Expect = e-129
Identities = 250/549 (45%), Positives = 339/549 (61%), Gaps = 72/549 (13%)
Query: 12 SRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHK 71
S + S ++ S S + V L L+ +VA+G+QFGWALQLSLLTPY+Q LGI H
Sbjct: 37 SESASPSNHSDSADGESVSKNCSLVTLVLSCTVAAGVQFGWALQLSLLTPYIQTLGISHA 96
Query: 72 WASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGY 131
++S IWLCGP++GL VQP VG SD+C+S++GRRRPFILVG+ I +AV+IIG++ADIGY
Sbjct: 97 FSSFIWLCGPITGLVVQPFVGIWSDKCTSKYGRRRPFILVGSFMISIAVIIIGFSADIGY 156
Query: 132 LIGD-----DITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVAN 186
L+GD + R A VVF+IGFW+LD+ANN QGP RALLADL+ D R T AN
Sbjct: 157 LLGDSKEHCSTFKGTRTRAAVVFIIGFWLLDLANNTVQGPARALLADLSGPDQRNT--AN 214
Query: 187 AYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLS 246
A F L+MA+GNILG++ G+ W + F F + AC +C NLK+AF L V F+ + T ++
Sbjct: 215 AVFCLWMAIGNILGFSAGASGKWQEWFPFLTSRACCAACGNLKAAFLLAVVFLTICTLVT 274
Query: 247 IVSAHEVPLSSS----------------------------------------------GA 260
I A E+P +S+ G
Sbjct: 275 IYFAKEIPFTSNKPTRIQDSAPLLDDLQSKGLEHSKLNNGTANGIKYERVERDTDEQFGN 334
Query: 261 GESGSAEEAF-------MWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGRE 313
E+ +E + + L + ++ + VL V ALTW+ WFPF LFDTDWMGRE
Sbjct: 335 SENEHQDETYVDGPGSVLVNLLTSLRHLPPAMHSVLIVMALTWLSWFPFFLFDTDWMGRE 394
Query: 314 IYGGDPEGGL----IYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIF 369
+Y GDP G +YD GVR GALGLLLNSVVL ++S L+E +C++ GA VW +SN
Sbjct: 395 VYHGDPTGDSLHMELYDQGVREGALGLLLNSVVLGISSFLIEPMCQRMGARVVWALSNFT 454
Query: 370 MAICF-----IAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYA 424
+ C I+++ L+ N I Y+ +G AA+ +F +LGFP+AITYSVP++
Sbjct: 455 VFACMAGTAVISLMSLSDDKNGIEYIMRGNETTR---TAAVIVFALLGFPLAITYSVPFS 511
Query: 425 LISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLS 484
+ + G GQGL++GVLNLAIV+PQ++VSLG+GPWDQLFGGGN PAF +A+VAA +
Sbjct: 512 VTAEVTADSGGGQGLAIGVLNLAIVIPQMIVSLGAGPWDQLFGGGNLPAFVLASVAAFAA 571
Query: 485 GLLALLAIP 493
G++AL +P
Sbjct: 572 GVIALQRLP 580
>At3g13620 unknown protein
Length = 478
Score = 37.7 bits (86), Expect = 0.015
Identities = 31/108 (28%), Positives = 49/108 (44%), Gaps = 12/108 (11%)
Query: 210 YKIFTFTLTPACSISCANLKSAFFLDVA----------FIVVTTYLSIVSAHEVPLSSSG 259
Y I F +T A S+ + ++ F + A +I + LS + E LSSS
Sbjct: 255 YLIPLFAVTGAVSVDQSRWENGFHAEAAEMIAGKWLKIWIEIGAVLSSIGLFEAQLSSSA 314
Query: 260 AGESGSAEEAFMWELFGT-FKYFSMPVWIVLSVTALTWIGWFPFNLFD 306
G AE F+ + FG K+F+ P W+ + ++AL +G N D
Sbjct: 315 YQLEGMAELGFLPKFFGVRSKWFNTP-WVGILISALMSLGLSYMNFTD 361
>At5g05310 unknown protein
Length = 496
Score = 33.1 bits (74), Expect = 0.37
Identities = 39/159 (24%), Positives = 65/159 (40%), Gaps = 28/159 (17%)
Query: 6 TTNPH-RSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWAL----QLSLLT 60
T N H R R + +TS P + P++P+ + QF WA+ +L L +
Sbjct: 248 TDNDHQRERKQEATSPKVGSP-KVASPKSPIS--------TTRPQF-WAILDGMRLILAS 297
Query: 61 PYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAV 120
PY+ + + +WL +S F V ++ S GRRR F + S V
Sbjct: 298 PYLLLVSL------FLWLGAVISSFFYFQKVNIIATTIKSSIGRRRLFAQIN--SFVAVF 349
Query: 121 VIIGYAADIGYLI---GDDITQNYRPFAIV--VFVIGFW 154
++IG G ++ G + + PF + + I W
Sbjct: 350 ILIGQLTLTGRILTVAGVTVAISASPFVALGNLVAIAIW 388
>At5g03570 transporter like protein
Length = 498
Score = 32.7 bits (73), Expect = 0.48
Identities = 23/66 (34%), Positives = 37/66 (55%), Gaps = 6/66 (9%)
Query: 401 VIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQGLSMGVLNLAIVVPQIVVS---- 456
V AL FT+L F +T ++ + I T+I +G+G+G+S GV A V+ ++ S
Sbjct: 315 VSLALLFFTVLSFGTLMTATLEWKGIPTYI--IGIGRGISAGVGLAATVLYPLMQSRISP 372
Query: 457 LGSGPW 462
L +G W
Sbjct: 373 LRTGVW 378
>At2g30300 hypothetical protein
Length = 500
Score = 31.2 bits (69), Expect = 1.4
Identities = 28/100 (28%), Positives = 44/100 (44%), Gaps = 4/100 (4%)
Query: 325 YDTGVRMGALG-LLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICFIAMLVLTYA 383
+ T R A G LLLNS+V V L+ + + G S + + FI + VLT A
Sbjct: 143 FHTSQREEASGYLLLNSLVPLVACLVTAPMLMRHGGDKTMSYSKD-VKVGFIVLFVLTIA 201
Query: 384 ANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPY 423
+ P +V+ +A+F + P+AI V +
Sbjct: 202 TGIYAVATSLVSVPAVLVLVGIALFLLA--PLAIPIGVGF 239
>At3g16240 delta tonoplast integral protein (delta-TIP)
Length = 250
Score = 30.8 bits (68), Expect = 1.8
Identities = 37/132 (28%), Positives = 59/132 (44%), Gaps = 18/132 (13%)
Query: 360 GFVWGISNIFMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITY 419
G + I+ +F I +L T A + YV+ G PT V A L + + IT+
Sbjct: 94 GQITVITGVFYWIA--QLLGSTAACFLLKYVTGGLAVPTHSVAAGLGSIEGVVMEIIITF 151
Query: 420 SVPYALISTHIEP----LGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPA-- 473
++ Y + +T +P LG L++G++ A + L +GP+ GG +PA
Sbjct: 152 ALVYTVYATAADPKKGSLGTIAPLAIGLIVGANI-------LAAGPFS---GGSMNPARS 201
Query: 474 FAVAAVAALLSG 485
F A A SG
Sbjct: 202 FGPAVAAGDFSG 213
>At4g10770 oligopeptide transporter like protein
Length = 766
Score = 29.6 bits (65), Expect = 4.1
Identities = 26/100 (26%), Positives = 48/100 (48%), Gaps = 15/100 (15%)
Query: 277 TFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGDPEGGLIYDTGVRMGALGL 336
+F Y+ P ++ +T+L+W+ WF F + M ++I G ++ GV GA+GL
Sbjct: 254 SFAYYVFPGYLFQIMTSLSWVCWF----FPSSVMAQQIGSG------LHGLGV--GAIGL 301
Query: 337 LLNSVVLAVTSLLMER--LCRKRGAGFVWGISNIFMAICF 374
+++ + S L G GFV + + + IC+
Sbjct: 302 DWSTISSYLGSPLASPWFATANVGVGFVL-VIYVLVPICY 340
>At2g07010 putative retroelement pol polyprotein
Length = 1413
Score = 29.6 bits (65), Expect = 4.1
Identities = 30/107 (28%), Positives = 44/107 (41%), Gaps = 20/107 (18%)
Query: 5 TTTNPHRSRTRSSTSTSTSRPVQPVQ-----------PRTPLRQLLRVASVASGIQFGWA 53
T+ P ++ T SS+STS S V P Q PR RQ+ + A + I + +
Sbjct: 869 TSVTPTQTPTNSSSSTSPSTNVSPPQQDTTPIIENTPPRQGKRQVQQPARLKDYILYNAS 928
Query: 54 LQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSS 100
+ TP+V +SI G PL ++SD C S
Sbjct: 929 CTPN--TPHVLSPSTSQSSSSI-------QGNLQYPLTDYISDECFS 966
>At5g20655 24 kDa vacuolar protein - like
Length = 616
Score = 29.3 bits (64), Expect = 5.3
Identities = 21/65 (32%), Positives = 36/65 (55%), Gaps = 6/65 (9%)
Query: 400 IVIAALAIFTILGFP-MAITYSVP----YALISTHIEPLGLGQGLSMGVLNLAIVVPQIV 454
+V+ AL + LG +A+ + VP Y L+ + P+ L + L + L +++ VP I+
Sbjct: 536 LVLLALGTYYKLGSTYLALVWLVPPAFAYGLLEATLSPIRLPKPLKLATLLISLAVP-IL 594
Query: 455 VSLGS 459
VS GS
Sbjct: 595 VSSGS 599
>At5g14570 high affinity nitrate transporter - like protein
Length = 493
Score = 29.3 bits (64), Expect = 5.3
Identities = 12/29 (41%), Positives = 17/29 (58%)
Query: 184 VANAYFSLFMAVGNILGYATGSYSGWYKI 212
VAN Y+ M GN++G A G +GW +
Sbjct: 146 VANQYWMSSMFSGNVIGLANGVSAGWANV 174
>At3g43790 transporter-like protein
Length = 478
Score = 29.3 bits (64), Expect = 5.3
Identities = 37/160 (23%), Positives = 65/160 (40%), Gaps = 27/160 (16%)
Query: 53 ALQLSLLTPYV----QQLGIPHKWASIIWLCGPVSGLFV--QPLVGHLSDRCSSRFGRRR 106
AL +S L PY+ + I + I + G V F+ + L + + R+GR+
Sbjct: 48 ALPISSLFPYIYFMIRDFHIAKQEEDIGFYAGFVGSSFMIGRALTSIFWGKLADRYGRK- 106
Query: 107 PFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGP 166
P IL+G S+++ + G + I V F++G + N G
Sbjct: 107 PIILIGTFSVIIFNTLFGLSTSFWLAIS------------VRFLLGCF------NCLLGV 148
Query: 167 CRALLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSY 206
RA +++ + ++ + S +G ILG A G Y
Sbjct: 149 IRAYASEVVSEE--YNALSLSVVSTSRGIGLILGPAIGGY 186
>At2g38460 unknown protein
Length = 524
Score = 29.3 bits (64), Expect = 5.3
Identities = 22/66 (33%), Positives = 37/66 (55%), Gaps = 6/66 (9%)
Query: 401 VIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQGLSMGV-LNLAIVVPQI---VVS 456
V AL FT+L F +T ++ + I T+I +G+G+G+S V L +V P + + +
Sbjct: 319 VSLALLFFTVLSFGTLMTATLQWEGIPTYI--IGIGRGISATVGLAATLVYPLMQSRLST 376
Query: 457 LGSGPW 462
L +G W
Sbjct: 377 LRTGLW 382
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.140 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,999,756
Number of Sequences: 26719
Number of extensions: 466901
Number of successful extensions: 1563
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 1512
Number of HSP's gapped (non-prelim): 36
length of query: 504
length of database: 11,318,596
effective HSP length: 104
effective length of query: 400
effective length of database: 8,539,820
effective search space: 3415928000
effective search space used: 3415928000
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146866.2


