

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146866.12 + phase: 0
(273 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
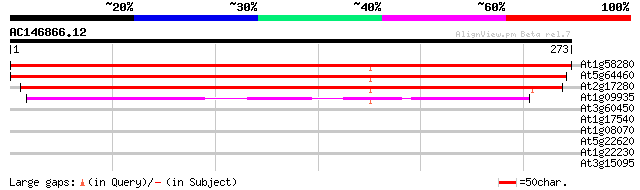
At1g58280 putative protein 382 e-107
At5g64460 putative phosphoglycerate mutase 377 e-105
At2g17280 unknown protein 362 e-100
At1g09935 putative protein 192 1e-49
At3g60450 unknown protein 35 0.056
At1g17540 protein kinase, putative 33 0.16
At1g08070 unknown protein 28 4.0
At5g22620 phosphoglycerate mutase like protein 28 5.3
At1g22230 hypothetical protein 28 6.9
At3g15095 unknown protein, 3' partial 27 9.0
>At1g58280 putative protein
Length = 276
Score = 382 bits (982), Expect = e-107
Identities = 180/274 (65%), Positives = 212/274 (76%), Gaps = 1/274 (0%)
Query: 1 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 60
M+T P Q LYPLH KTIHLVRHAQG+HNVEGEKNH AYLS D FDA+LTPLGWQQV+NL
Sbjct: 1 METKPSQGLYPLHRCKTIHLVRHAQGIHNVEGEKNHKAYLSEDLFDAHLTPLGWQQVDNL 60
Query: 61 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 120
KHV A G+S +IELVVVSPLLRT+QTAVG FGGE DGVN P LM G+SD PA+S
Sbjct: 61 HKHVNASGISNRIELVVVSPLLRTLQTAVGTFGGEGYKDGVNTPLLMTAGAGNSDRPAIS 120
Query: 121 SLNCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKE 179
LN PPF+AVE CRE +G+HPCD+R +++YR +FP IDFSLIETD+D WKP+ RE+ +
Sbjct: 121 RLNRPPFIAVESCREHLGVHPCDRRSNITKYRELFPAIDFSLIETDEDVLWKPDIREEDK 180
Query: 180 EVTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRS 239
++ RG+KF WL TRKEKEIAVVTHS FL+ TL++FGNDC P++K E+ F NCELRS
Sbjct: 181 DIATRGVKFFNWLSTRKEKEIAVVTHSGFLYQTLNSFGNDCDPSVKNEISKKFVNCELRS 240
Query: 240 MVIVDKCMIGSNNSTTNYPGKIPHGPDLPSDATD 273
V+VDKCM S+ TNYPG I G D SD D
Sbjct: 241 FVLVDKCMSSSDPPMTNYPGTILTGEDASSDIAD 274
>At5g64460 putative phosphoglycerate mutase
Length = 282
Score = 377 bits (968), Expect = e-105
Identities = 174/272 (63%), Positives = 221/272 (80%), Gaps = 1/272 (0%)
Query: 1 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 60
M+T G LYPLH KTI+LVRHAQG+HNV+GEKN+ AY+S+D+FDA LT LGW+QV++L
Sbjct: 1 METGAGIGLYPLHRCKTIYLVRHAQGIHNVDGEKNYKAYMSHDYFDAELTQLGWKQVDSL 60
Query: 61 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 120
+KHV + GL KKIELV+ SPL+RT+QTAVGVFGGE TD + PLM+ N G+S A+S
Sbjct: 61 RKHVHSSGLHKKIELVISSPLMRTLQTAVGVFGGEGYTDMSDVLPLMVANAGNSSRAAIS 120
Query: 121 SLNCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKE 179
SLNCPP + E CRE +G+HPCD+RR++S+Y+ +FP +DFSLIE+++D WK + RE E
Sbjct: 121 SLNCPPVITEESCREHLGVHPCDQRRSISDYQFLFPAVDFSLIESEEDKLWKADVRETIE 180
Query: 180 EVTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRS 239
E+ RG KFL WL TRKEKEIA+VTHS FLF+TL+A N+CHP++K E+C HFANCELRS
Sbjct: 181 ELAARGKKFLNWLWTRKEKEIAIVTHSGFLFHTLNALQNECHPDVKKEICGHFANCELRS 240
Query: 240 MVIVDKCMIGSNNSTTNYPGKIPHGPDLPSDA 271
MVIVD+ M+GS++S T+YPGKIP G DLPSDA
Sbjct: 241 MVIVDRSMLGSDSSVTDYPGKIPKGIDLPSDA 272
>At2g17280 unknown protein
Length = 271
Score = 362 bits (928), Expect = e-100
Identities = 170/266 (63%), Positives = 213/266 (79%), Gaps = 2/266 (0%)
Query: 6 GQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVK 65
G LYPLH KTIHLVRHAQG+HNV GEK+H AY S D+FDA+LTPLGWQQV+NL+ HV+
Sbjct: 5 GIGLYPLHRCKTIHLVRHAQGIHNVAGEKDHSAYSSEDYFDAHLTPLGWQQVDNLRNHVR 64
Query: 66 AIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCP 125
A L K+ELV+VSP+LRT+QTAVG FGGE +T+G + PLM+ N G SD PA+SSLN P
Sbjct: 65 AAQLLNKVELVIVSPMLRTIQTAVGAFGGEEDTNGADATPLMVANAGSSDRPAISSLNSP 124
Query: 126 PFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKEEVTGR 184
PF+AVELCRE MG HPCD+RR+V+EY+ +FP IDFS+IETD+D WKP RE EEV R
Sbjct: 125 PFLAVELCRETMGDHPCDRRRSVTEYKALFPAIDFSIIETDNDVLWKPSPRESLEEVAAR 184
Query: 185 GLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRSMVIVD 244
G++F++W+ TRKEKEIA+V+HS FL LS+FG DC ++K E+ H +NCELRSMVIVD
Sbjct: 185 GVEFIKWIWTRKEKEIAIVSHSGFLHGLLSSFGKDCCDDLKKELSIHLSNCELRSMVIVD 244
Query: 245 KCMIGSNNS-TTNYPGKIPHGPDLPS 269
+ +G++++ TTNYPGK+P G D PS
Sbjct: 245 RGNLGTDSAETTNYPGKVPEGLDNPS 270
>At1g09935 putative protein
Length = 212
Score = 192 bits (489), Expect = 1e-49
Identities = 109/246 (44%), Positives = 146/246 (59%), Gaps = 40/246 (16%)
Query: 9 LYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAIG 68
LYPL K IHL+RH Q +HNVE EK+ +A LS FDA LT G QQVENL++ V + G
Sbjct: 6 LYPLESCKIIHLLRHGQALHNVEAEKDRNALLSPHLFDAPLTDHGHQQVENLRERVVSSG 65
Query: 69 LSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCPPFV 128
L K++ELVV SPL RTMQTAVGVFG E + S + PP +
Sbjct: 66 LLKRVELVVTSPLFRTMQTAVGVFGNE--------------------YKQSSMTSSPPIL 105
Query: 129 AVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKEEVTGRGLK 187
A+E+ R++ G+ P D RR IE+++D W+P+ RE +EE+ RGL+
Sbjct: 106 ALEVARDRNGVRPPDMRRN---------------IESEEDNLWRPDVRESEEEILARGLE 150
Query: 188 FLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRSMVIVDKCM 247
F++W EKE+AVV+H L + L F NDC +I+ E+C FANCE+R++VIVDK M
Sbjct: 151 FMKW----PEKEVAVVSHGIVLQHMLYVFANDCDLSIRHELCKRFANCEIRTVVIVDKGM 206
Query: 248 IGSNNS 253
S +
Sbjct: 207 ASSTEN 212
>At3g60450 unknown protein
Length = 274
Score = 34.7 bits (78), Expect = 0.056
Identities = 44/159 (27%), Positives = 68/159 (42%), Gaps = 21/159 (13%)
Query: 73 IELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCPPFVAVEL 132
I V VSP LR +QTA V A VN P + D P++ + + L
Sbjct: 66 IHRVFVSPFLRCLQTASEVV---AALSAVNVDP---NAMSSKDVPSIDKSKLKVSIELGL 119
Query: 133 CRE------QMGLHPCDKR--RTVSEYRHMFPGIDFSLIETDDDTWWKPEREKKEEVTG- 183
C + L P D + TVS+ MFP +++ + D +K + +E V G
Sbjct: 120 CEMLNSVAIRRELAPKDGKFDFTVSDIETMFPE---GMVDHNVDMVYKELPKWEESVEGC 176
Query: 184 --RGLKFLEWLCTR-KEKEIAVVTHSSFLFNTLSAFGND 219
R +K ++ L + E+ + +VTH + T S F D
Sbjct: 177 RDRYVKVVKALADKYPEENLLLVTHGEGVGTTFSTFYKD 215
>At1g17540 protein kinase, putative
Length = 740
Score = 33.1 bits (74), Expect = 0.16
Identities = 27/109 (24%), Positives = 45/109 (40%), Gaps = 1/109 (0%)
Query: 66 AIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCP 125
A+GLS ++E + LR + E T + + L + D P ++S+ P
Sbjct: 623 AMGLSHRVEKAIEKKKLREVLDPKISDWPEEETMVLAQLALQCCELRKKDRPDLASVLLP 682
Query: 126 PFVAV-ELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKP 173
+ E E +H D+ VS ++ P S +T+DD W P
Sbjct: 683 ALSKLREFATEDHEVHNSDRTFHVSRAHNLVPLSPISSCQTEDDAWENP 731
>At1g08070 unknown protein
Length = 741
Score = 28.5 bits (62), Expect = 4.0
Identities = 25/88 (28%), Positives = 39/88 (43%), Gaps = 12/88 (13%)
Query: 56 QVENLQKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSD 115
Q+ + H LSK IE ++SP + A+ VF + +P L+I N
Sbjct: 55 QMIKIGLHNTNYALSKLIEFCILSPHFEGLPYAISVF------KTIQEPNLLIWNTMFRG 108
Query: 116 HPAVSSLNCPPFVAVEL--CREQMGLHP 141
H +L+ P A++L C +GL P
Sbjct: 109 H----ALSSDPVSALKLYVCMISLGLLP 132
>At5g22620 phosphoglycerate mutase like protein
Length = 482
Score = 28.1 bits (61), Expect = 5.3
Identities = 25/87 (28%), Positives = 40/87 (45%), Gaps = 10/87 (11%)
Query: 7 QSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKA 66
Q + + +K + LVRH Q N EG S DF + LT G Q E ++ +
Sbjct: 39 QDQFTVETTKRVVLVRHGQSTWNEEGRIQG----SSDF--SVLTKKGESQAEISRQML-- 90
Query: 67 IGLSKKIELVVVSPLLRTMQTAVGVFG 93
+ ++ SPL R+ +TA ++G
Sbjct: 91 --IDDSFDVCFTSPLKRSKKTAEIIWG 115
>At1g22230 hypothetical protein
Length = 314
Score = 27.7 bits (60), Expect = 6.9
Identities = 13/45 (28%), Positives = 23/45 (50%)
Query: 155 FPGIDFSLIETDDDTWWKPEREKKEEVTGRGLKFLEWLCTRKEKE 199
FP +D + ++D + E E+++E G F +WL EK+
Sbjct: 146 FPPVDIISDDEEEDEEEEEEDEEEDEDESSGTVFSKWLMVLHEKQ 190
>At3g15095 unknown protein, 3' partial
Length = 417
Score = 27.3 bits (59), Expect = 9.0
Identities = 17/42 (40%), Positives = 24/42 (56%), Gaps = 8/42 (19%)
Query: 143 DKRRTVSEYRHMFPGIDFSLIETDDDTWWKPEREKKEEVTGR 184
D+RR S RH+F G+D S IE K E++++ E GR
Sbjct: 259 DRRR--SRRRHVFEGLDLSEIE------MKTEKKERGEEVGR 292
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.135 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,715,877
Number of Sequences: 26719
Number of extensions: 286729
Number of successful extensions: 625
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 614
Number of HSP's gapped (non-prelim): 10
length of query: 273
length of database: 11,318,596
effective HSP length: 98
effective length of query: 175
effective length of database: 8,700,134
effective search space: 1522523450
effective search space used: 1522523450
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146866.12


