

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146864.6 - phase: 0 /pseudo
(180 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
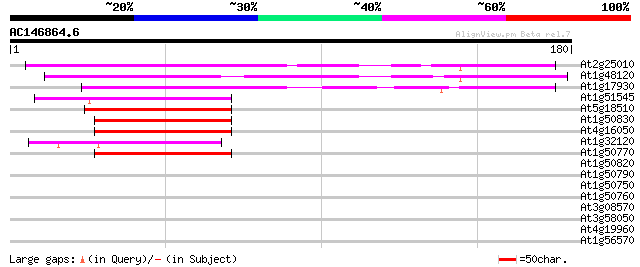
Score E
Sequences producing significant alignments: (bits) Value
At2g25010 unknown protein 71 3e-13
At1g48120 serine/threonine phosphatase PP7, putative 66 9e-12
At1g17930 unknown protein 63 7e-11
At1g51545 unknown protein 45 2e-05
At5g18510 putative protein 44 5e-05
At1g50830 hypothetical protein 44 6e-05
At4g16050 hypothetical protein 42 1e-04
At1g32120 hypothetical protein 41 4e-04
At1g50770 hypothetical protein 40 9e-04
At1g50820 hypothetical protein 39 0.001
At1g50790 hypothetical protein 37 0.007
At1g50750 hypothetical protein 34 0.049
At1g50760 hypothetical protein 34 0.049
At3g08570 hypothetical protein 28 2.7
At3g58050 putative protein 27 4.5
At4g19960 potassium transporter-like protein 27 5.9
At1g56570 hypothetical protein 27 7.7
>At2g25010 unknown protein
Length = 509
Score = 71.2 bits (173), Expect = 3e-13
Identities = 47/171 (27%), Positives = 85/171 (49%), Gaps = 17/171 (9%)
Query: 6 NVIRASGLYLLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPV 65
N++ +G + +++ L+ A ERW ET++FHLP+GEMTITLD+V+ +L + +
Sbjct: 53 NLVDKAGFGYFRKIGPMSLNNSLISALVERWRRETNTFHLPLGEMTITLDEVALVLGLEI 112
Query: 66 GGNLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKS 125
G+ + + LG + +A K + + + + L ++
Sbjct: 113 DGDPIVGSKVGDEVAMDMCGRLLG---KLPSAANKEVNCSRVKLNWL----------KRT 159
Query: 126 YANQPEEEDSMEWYRTRC-IRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDD 175
++ PE+ + +C RA+LLYL+G T+F+ G+ V YL F+D
Sbjct: 160 FSECPED---ASFDVVKCHTRAYLLYLIGSTIFATTDGDKVSVKYLPLFED 207
>At1g48120 serine/threonine phosphatase PP7, putative
Length = 1338
Score = 66.2 bits (160), Expect = 9e-12
Identities = 49/169 (28%), Positives = 80/169 (46%), Gaps = 21/169 (12%)
Query: 12 GLYLLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLF 71
GLY + + + Q+D+ L+ A ERW ET +FHLP GE+T+TL DV+ LL + V G
Sbjct: 68 GLYGVYKVAFIQLDYALITALVERWRPETHTFHLPAGEITVTLQDVNILLGLRVDGP--- 124
Query: 72 HESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPE 131
++ T+Y L + K L +H++ L +++ N P
Sbjct: 125 ----AVTGSTKYNWADLCEDLLGHRPGPKDLHGSHVSLAWL----------RENFRNLPA 170
Query: 132 EEDSMEWYRTRC-IRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDDLTTV 179
+ D + +C RAF+L L+ L+ DK+ + + +L D V
Sbjct: 171 DPDEV---TLKCHTRAFVLALMSGFLYGDKSKHDVALTFLPLLRDFDEV 216
>At1g17930 unknown protein
Length = 478
Score = 63.2 bits (152), Expect = 7e-11
Identities = 46/153 (30%), Positives = 75/153 (48%), Gaps = 20/153 (13%)
Query: 24 VDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQY 83
+++ L+ A ERW ET++FH P GEMTITLD+VS +L + V G + +Q
Sbjct: 62 LNNSLISALVERWRRETNTFHFPCGEMTITLDEVSLILGLAVDGKPVVGVKEKDEDPSQV 121
Query: 84 LVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSME-WYRTR 142
+ LG +L ++ + + + + +S+A P+ E Y T
Sbjct: 122 CLRLLG-----------KLPKGELSGNRVTAKWLK-----ESFAECPKGATMKEIEYHT- 164
Query: 143 CIRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDD 175
RA+L+Y+VG T+F+ + V YL F+D
Sbjct: 165 --RAYLIYIVGSTIFATTDPSKISVDYLILFED 195
>At1g51545 unknown protein
Length = 629
Score = 45.4 bits (106), Expect = 2e-05
Identities = 25/65 (38%), Positives = 39/65 (59%), Gaps = 2/65 (3%)
Query: 9 RASGLYLLLETNYGQV--DHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVG 66
R +G++ ++ + ++ + LL+A E+W ET SF P GE TITL+DV LL V
Sbjct: 10 RKAGIFEAIKASMYKIRKNQSLLLALVEKWCPETKSFLFPWGEATITLEDVLVLLGFSVQ 69
Query: 67 GNLLF 71
G+ +F
Sbjct: 70 GSPVF 74
>At5g18510 putative protein
Length = 702
Score = 43.9 bits (102), Expect = 5e-05
Identities = 22/47 (46%), Positives = 30/47 (63%)
Query: 25 DHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLF 71
D +++ +E+W SET SF P GE TITL+DV LL V G+ +F
Sbjct: 94 DTSSILSIAEKWCSETKSFIFPWGEATITLEDVMVLLGFSVLGSPVF 140
>At1g50830 hypothetical protein
Length = 768
Score = 43.5 bits (101), Expect = 6e-05
Identities = 22/44 (50%), Positives = 29/44 (65%)
Query: 28 LLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLF 71
L+++ SE+W ET SF P GE TITL+DV LL V G+ +F
Sbjct: 119 LILSVSEKWCPETKSFVFPWGEATITLEDVMVLLGFSVLGSPVF 162
>At4g16050 hypothetical protein
Length = 900
Score = 42.4 bits (98), Expect = 1e-04
Identities = 18/44 (40%), Positives = 30/44 (67%)
Query: 28 LLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLF 71
L+++ ++ W ET++F P GE TITL+DV+ LL + G+ +F
Sbjct: 363 LILSLAQNWCPETNTFVFPWGEATITLEDVNVLLGFSISGSSVF 406
>At1g32120 hypothetical protein
Length = 1206
Score = 40.8 bits (94), Expect = 4e-04
Identities = 25/64 (39%), Positives = 38/64 (59%), Gaps = 2/64 (3%)
Query: 7 VIRASGLY-LLLETNYGQVDHG-LLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIP 64
V + SG+Y +L + Y H L++A E+W ET++F P GE T+TL+D+ L +
Sbjct: 98 VWKKSGVYDAILASRYQIKRHDDLIVALVEKWCIETNTFVFPWGEATLTLEDMIVLGGLS 157
Query: 65 VGGN 68
V GN
Sbjct: 158 VTGN 161
>At1g50770 hypothetical protein
Length = 632
Score = 39.7 bits (91), Expect = 9e-04
Identities = 19/44 (43%), Positives = 28/44 (63%)
Query: 28 LLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLF 71
L++ +E+W +T +F P GE TITL+DV LL V G+ +F
Sbjct: 100 LVLGIAEKWCPDTKTFVFPWGETTITLEDVMLLLGFSVLGSPVF 143
>At1g50820 hypothetical protein
Length = 528
Score = 39.3 bits (90), Expect = 0.001
Identities = 20/47 (42%), Positives = 28/47 (59%)
Query: 25 DHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLF 71
D L++ +E+W +T +F P GE TITL+DV LL V G +F
Sbjct: 94 DTDLVLGLAEKWCPDTKTFIFPWGEATITLEDVMVLLGFSVLGLPVF 140
>At1g50790 hypothetical protein
Length = 812
Score = 36.6 bits (83), Expect = 0.007
Identities = 18/44 (40%), Positives = 28/44 (62%)
Query: 28 LLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLF 71
L++ +E+W +T++F GE TITL+DV LL V G+ +F
Sbjct: 101 LVMGIAEKWCPDTNTFVFSWGEATITLEDVMVLLGFSVLGSPVF 144
>At1g50750 hypothetical protein
Length = 816
Score = 33.9 bits (76), Expect = 0.049
Identities = 16/44 (36%), Positives = 25/44 (56%)
Query: 28 LLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLF 71
L++ +E+W T +F P GE +TL+DV L V G+ +F
Sbjct: 76 LILGIAEKWCPYTKTFVFPWGETAVTLEDVMVLSGFSVLGSPVF 119
>At1g50760 hypothetical protein
Length = 649
Score = 33.9 bits (76), Expect = 0.049
Identities = 17/44 (38%), Positives = 25/44 (56%)
Query: 28 LLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLF 71
L++ +E+W +T +F GE TITL+DV L V G+ F
Sbjct: 90 LVLGVAEKWSPDTKTFVFSWGEATITLEDVMVLSGFSVLGSPAF 133
>At3g08570 hypothetical protein
Length = 612
Score = 28.1 bits (61), Expect = 2.7
Identities = 12/36 (33%), Positives = 21/36 (58%)
Query: 133 EDSMEWYRTRCIRAFLLYLVGCTLFSDKAGNSCCVV 168
+ +++ RCI+ L + C++FSD AG+ VV
Sbjct: 2 QSRIDFLLIRCIKDSFLPYINCSIFSDVAGDITIVV 37
>At3g58050 putative protein
Length = 1209
Score = 27.3 bits (59), Expect = 4.5
Identities = 14/59 (23%), Positives = 30/59 (50%)
Query: 98 QTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSMEWYRTRCIRAFLLYLVGCTL 156
+T L +A ++ DTL+ +++ + + + +EED ME R R +++ C +
Sbjct: 197 ETCALHTARLSCDTLVDFWSALSEDTRQSLLRMKEEDFMERLRYRICYHSSYHILNCKM 255
>At4g19960 potassium transporter-like protein
Length = 842
Score = 26.9 bits (58), Expect = 5.9
Identities = 17/58 (29%), Positives = 34/58 (58%), Gaps = 3/58 (5%)
Query: 108 TYDTLLSIYTSYLTEAKSYANQPEEE--DSMEWYRTRCIRAFLLYLVGCTLFSDKAGN 163
T DT++S + + T + S N EEE D +E+ +T C + +++++G T+ + G+
Sbjct: 744 TLDTIVSAESLHNTVSFSQDNTVEEEETDELEFLKT-CKESGVVHIMGNTVVKARTGS 800
>At1g56570 hypothetical protein
Length = 541
Score = 26.6 bits (57), Expect = 7.7
Identities = 11/31 (35%), Positives = 21/31 (67%), Gaps = 2/31 (6%)
Query: 110 DTLLSIYT--SYLTEAKSYANQPEEEDSMEW 138
+++L +Y YL+EAK Y ++ E++D + W
Sbjct: 182 NSILDLYCRCGYLSEAKHYFHEMEDKDLITW 212
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.137 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,076,218
Number of Sequences: 26719
Number of extensions: 160540
Number of successful extensions: 350
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 332
Number of HSP's gapped (non-prelim): 17
length of query: 180
length of database: 11,318,596
effective HSP length: 93
effective length of query: 87
effective length of database: 8,833,729
effective search space: 768534423
effective search space used: 768534423
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146864.6


