

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.7 - phase: 0
(198 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
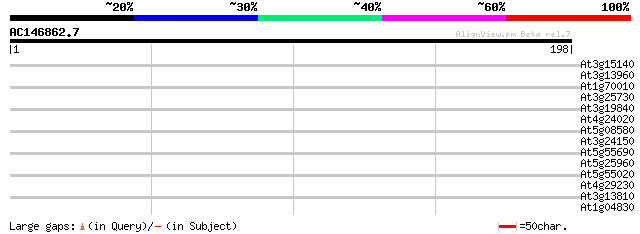
Score E
Sequences producing significant alignments: (bits) Value
At3g15140 unknown protein 38 0.004
At3g13960 hypothetical protein 33 0.13
At1g70010 hypothetical protein 33 0.13
At3g25730 AP2 domain transcription factor 30 1.1
At3g19840 unknown protein 29 1.4
At4g24020 unknown protein (At4g24020) 29 1.9
At5g08580 Unknown protein (MAH20.14) 28 2.4
At3g24150 hypothetical protein 28 3.2
At5g55690 unknown protein 27 5.4
At5g25960 putative protein 27 5.4
At5g55020 putative transcription factor MYB120 (MYB120) 27 7.1
At4g29230 putative protein 27 7.1
At3g13810 zinc finger protein, putative 27 9.2
At1g04830 unknown protein 27 9.2
>At3g15140 unknown protein
Length = 337
Score = 37.7 bits (86), Expect = 0.004
Identities = 22/56 (39%), Positives = 31/56 (55%), Gaps = 9/56 (16%)
Query: 122 REQYQEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPT 177
R++ QEF F+VID E EI+EFP +IV + T ++ F +VRPT
Sbjct: 120 RKKSQEFNFFLVIDLEGKV---------EILEFPILIVDAKTMEVVDLFHRFVRPT 166
>At3g13960 hypothetical protein
Length = 396
Score = 32.7 bits (73), Expect = 0.13
Identities = 18/64 (28%), Positives = 31/64 (48%), Gaps = 4/64 (6%)
Query: 31 FPELNNEPGNHPSGNVPEPNHRLGSEFLEPSNEFHNKPTYHHDYSTWTAC----HFHPHK 86
FPE + + P + +P + ++ + HNK +HH YS+ ++ H H H+
Sbjct: 261 FPEASRSFQDSPYHHHQQPLATVMNDPYHHCSTDHNKIDHHHTYSSSSSSQHLHHDHDHR 320
Query: 87 MQQC 90
QQC
Sbjct: 321 QQQC 324
>At1g70010 hypothetical protein
Length = 1315
Score = 32.7 bits (73), Expect = 0.13
Identities = 19/66 (28%), Positives = 32/66 (47%), Gaps = 13/66 (19%)
Query: 24 NGSSMEVFPELNNEPGNHPSGNVPEPNHRLGSEFLEPSNEFHNKPTYHHDYSTWTACHFH 83
+ SS+E+ P N P+ NVPEP+ ++ S+ KP Y DY +
Sbjct: 725 SSSSVEILPSAN------PTNNVPEPS-------VQTSHRKAKKPAYLQDYYCHSVVSST 771
Query: 84 PHKMQQ 89
PH++++
Sbjct: 772 PHEIRK 777
>At3g25730 AP2 domain transcription factor
Length = 333
Score = 29.6 bits (65), Expect = 1.1
Identities = 19/54 (35%), Positives = 24/54 (44%), Gaps = 1/54 (1%)
Query: 7 ASLKCLQGKGPPFTFQC-NGSSMEVFPELNNEPGNHPSGNVPEPNHRLGSEFLE 59
ASL G G NG +EV E P + G VP+PN R G++ E
Sbjct: 26 ASLLYRMGSGTSVVLDSENGVEVEVEAESRKLPSSRFKGVVPQPNGRWGAQIYE 79
>At3g19840 unknown protein
Length = 830
Score = 29.3 bits (64), Expect = 1.4
Identities = 20/73 (27%), Positives = 24/73 (32%), Gaps = 6/73 (8%)
Query: 38 PGNHPSGNVPEPNHRLGSEFLEPSNEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFEN 97
PG++P P P G + P H P YH T P M F +
Sbjct: 113 PGSNPFSTTPRPGMSAGPAQMNPGIHPHMYPPYHSLPGTPQGMWLQPPSMGGIPRAPFLS 172
Query: 98 H------FYPHPV 104
H YP PV
Sbjct: 173 HPTTFPGSYPFPV 185
>At4g24020 unknown protein (At4g24020)
Length = 959
Score = 28.9 bits (63), Expect = 1.9
Identities = 21/57 (36%), Positives = 24/57 (41%), Gaps = 4/57 (7%)
Query: 24 NGSSMEVFPELNNEPGNHPSGNVP-EPNHRLGSEFLEPSNEFHNKPTYHHDYSTWTA 79
NGS P NN P + S + P EPN GS L PSN T T T+
Sbjct: 686 NGSKPPELPNTNNSPNHWSSDHSPNEPN---GSPELPPSNGHKRSRTVDESAGTPTS 739
>At5g08580 Unknown protein (MAH20.14)
Length = 391
Score = 28.5 bits (62), Expect = 2.4
Identities = 16/48 (33%), Positives = 25/48 (51%)
Query: 115 MVSQGYPREQYQEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSV 162
++S+ +P E Y Q I +A DKD+ E+IE P V S++
Sbjct: 327 IISKIHPTEHYYAKQQADYIISQADSDKDRRLTLAEMIEHPYVFYSAI 374
>At3g24150 hypothetical protein
Length = 343
Score = 28.1 bits (61), Expect = 3.2
Identities = 10/22 (45%), Positives = 15/22 (67%)
Query: 53 LGSEFLEPSNEFHNKPTYHHDY 74
LG + EP + N+P+YHH+Y
Sbjct: 135 LGPKIQEPVSASTNEPSYHHEY 156
>At5g55690 unknown protein
Length = 277
Score = 27.3 bits (59), Expect = 5.4
Identities = 20/56 (35%), Positives = 24/56 (42%), Gaps = 8/56 (14%)
Query: 56 EFLEPSNEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFENHFYPHPVENQFQYA 111
E LE +N KPT Y TW K+ QC +N F VEN+ Q A
Sbjct: 94 ECLEKNNTKVEKPTIATKYPTW------DKKLDQCSLNDLYAVFM--AVENKIQEA 141
>At5g25960 putative protein
Length = 352
Score = 27.3 bits (59), Expect = 5.4
Identities = 11/29 (37%), Positives = 18/29 (61%)
Query: 20 TFQCNGSSMEVFPELNNEPGNHPSGNVPE 48
T NGSS ++ ++ ++ GN P G +PE
Sbjct: 83 TIPNNGSSEQITSQIWSKSGNCPKGTIPE 111
>At5g55020 putative transcription factor MYB120 (MYB120)
Length = 523
Score = 26.9 bits (58), Expect = 7.1
Identities = 14/45 (31%), Positives = 16/45 (35%)
Query: 68 PTYHHDYSTWTACHFHPHKMQQCQMNAFENHFYPHPVENQFQYAP 112
P Y D H HPH QQ Q N +H Q + P
Sbjct: 135 PLYPPDIIPNHQLHPHPHHQQQQQHNHHHHHHQQQQQHQQMYFQP 179
>At4g29230 putative protein
Length = 498
Score = 26.9 bits (58), Expect = 7.1
Identities = 20/97 (20%), Positives = 35/97 (35%), Gaps = 14/97 (14%)
Query: 47 PEPNHRLGSEFLEPSNEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFENHFYPHPV-- 104
P+ H + + + ++P+ + H H H+ QQ + +AF HP+
Sbjct: 328 PQTGHATCEDVMAEQHRHRHQPSSSTSHHMAHDHHHHHHQQQQQRHHAFNISQPTHPIST 387
Query: 105 ----ENQFQYAPINMVSQG--------YPREQYQEFQ 129
+A IN++ P E YQ Q
Sbjct: 388 IISPSTSLHHASINILDDNPYHVHRILLPNENYQTQQ 424
>At3g13810 zinc finger protein, putative
Length = 513
Score = 26.6 bits (57), Expect = 9.2
Identities = 16/78 (20%), Positives = 33/78 (41%), Gaps = 8/78 (10%)
Query: 35 NNEPGN---HPSGNVPEPNHRLGSEFLEPSNEFHNKPTYHHDYSTWTACHFHPHKMQQCQ 91
NN+P H S + P +H+ +P+ + + H+++ + HF +
Sbjct: 243 NNQPNPLLIHQSASHPHHHHQT-----QPTINVSSSSSSSHNHNIINSLHFDTNNGNTNN 297
Query: 92 MNAFENHFYPHPVENQFQ 109
N NH + P++ + Q
Sbjct: 298 SNNSNNHLHTFPMKKEQQ 315
>At1g04830 unknown protein
Length = 448
Score = 26.6 bits (57), Expect = 9.2
Identities = 11/26 (42%), Positives = 16/26 (61%)
Query: 27 SMEVFPELNNEPGNHPSGNVPEPNHR 52
S+ V P LN+ P + +VP P+HR
Sbjct: 43 SISVAPRLNSAPPPSSNTSVPSPSHR 68
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.135 0.441
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,285,606
Number of Sequences: 26719
Number of extensions: 244086
Number of successful extensions: 505
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 491
Number of HSP's gapped (non-prelim): 18
length of query: 198
length of database: 11,318,596
effective HSP length: 94
effective length of query: 104
effective length of database: 8,807,010
effective search space: 915929040
effective search space used: 915929040
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146862.7


