

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.24 - phase: 0
(580 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
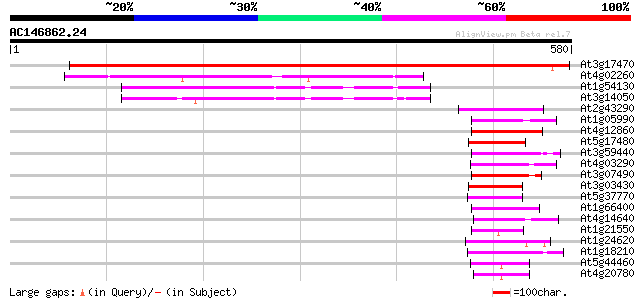
Score E
Sequences producing significant alignments: (bits) Value
At3g17470 hypothetical protein 710 0.0
At4g02260 RSH1 146 4e-35
At1g54130 RSH3 145 5e-35
At3g14050 putative GTP pyrophosphokinase 140 2e-33
At2g43290 putative calcium binding protein 58 1e-08
At1g05990 calcium-binding protein, putative 57 3e-08
At4g12860 putative calmodulin 56 6e-08
At5g17480 calcium-binding protein 53 4e-07
At3g59440 calmodulin-like protein 53 4e-07
At4g03290 putative calmodulin 52 1e-06
At3g07490 putative calmodulin 52 1e-06
At3g03430 pollen allergen Bra r II 50 3e-06
At5g37770 CALMODULIN-RELATED PROTEIN 2, TOUCH-INDUCED (TCH2) 49 1e-05
At1g66400 calmodulin-related protein 45 8e-05
At4g14640 calmodulin 45 1e-04
At1g21550 unknown protein 44 2e-04
At1g24620 putative calmodulin 44 2e-04
At1g18210 unknown protein 44 3e-04
At5g44460 calmodulin-like protein 43 5e-04
At4g20780 calcium-binding protein - like 43 5e-04
>At3g17470 hypothetical protein
Length = 570
Score = 710 bits (1832), Expect = 0.0
Identities = 363/523 (69%), Positives = 433/523 (82%), Gaps = 7/523 (1%)
Query: 63 GGKMVIELVGAFNDLTERMKV--LSTSSSGLLFKSLKLSIPVLQTSPLTPDGRSPLSKAL 120
GGKMV+ELVGAFN++TERM LSTSSS LLFK+LKLSIP+LQ+ PL DGRSPLSKAL
Sbjct: 45 GGKMVVELVGAFNEVTERMNSVWLSTSSSRLLFKALKLSIPILQSLPLASDGRSPLSKAL 104
Query: 121 SIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKNFASRV 180
S++++LADLQMDAEVISA IL EV++ ++++E+R IG+ TAHLLHE RVKN +V
Sbjct: 105 SLSIILADLQMDAEVISASILSEVVDANAISIYEVRDHIGTGTAHLLHEIFRVKNIPFKV 164
Query: 181 DILDDENAAALRKFCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQIISLQVMKIYAPL 240
D+LDDE AA+LRKF LTYYDIRA+I+DL KLD MRHL HLPRY+QQI+SL+V+KIY+PL
Sbjct: 165 DVLDDETAASLRKFYLTYYDIRAVIMDLVSKLDEMRHLDHLPRYRQQILSLEVLKIYSPL 224
Query: 241 AHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSDPI 300
AHAVG N++SLELED+SF+YLFP SY+Y+D+WLR E G LIDVYK++L SLK D +
Sbjct: 225 AHAVGANHLSLELEDISFRYLFPCSYIYLDSWLRGHENGSKPLIDVYKEQLHRSLKDDLV 284
Query: 301 LAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRENALEAGERAC 360
LAE+V+D+ +KGRYKSRYS MKKLL+DGR+PE+VNDVLGLRV+L P S N +E GE+AC
Sbjct: 285 LAEMVNDVYIKGRYKSRYSMMKKLLRDGRKPEEVNDVLGLRVILMPNSVVNDVEVGEKAC 344
Query: 361 YRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIRTTEMDR 420
YR +II+S+WKEIP RTKDYI+RPK NGYRSLHMAVDVS+ + RPLMEIQIRT +MD
Sbjct: 345 YRTSEIIRSLWKEIPHRTKDYIARPKENGYRSLHMAVDVSDSDQIRPLMEIQIRTMDMDG 404
Query: 421 LAVGGMASHSLYKAGLTNPEEAKRLKTIMLAAAELAALRLKDFPSANHKGIE--FDQRDR 478
A G ASHSLYK GLT+P+EAKRLK IMLAAA+LAA+RLKD S H+ + +QRDR
Sbjct: 405 SANAGTASHSLYKGGLTDPKEAKRLKAIMLAAADLAAIRLKDISSNKHQSFKTTTNQRDR 464
Query: 479 VFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMMLLLDSNSDGSLSSDEFQMFQKQVE 538
VF LLDKNGDG ISIEEL EV+EELGAPGEDA +MM LLDSNSDGSLSSDEF FQKQVE
Sbjct: 465 VFCLLDKNGDGMISIEELMEVMEELGAPGEDAEEMMQLLDSNSDGSLSSDEFDTFQKQVE 524
Query: 539 MVRNLEDRDDEYKKILDEKLH---MADDSGLIQVYNKEFGNRL 578
+R EDRD+EYK +LDEKLH D +GLIQ+YNKE +RL
Sbjct: 525 FMRKWEDRDNEYKSLLDEKLHDLPHQDTTGLIQLYNKELEDRL 567
>At4g02260 RSH1
Length = 883
Score = 146 bits (368), Expect = 4e-35
Identities = 109/390 (27%), Positives = 183/390 (45%), Gaps = 31/390 (7%)
Query: 57 SVEVPGGGKMVIELVGAFNDLTERMKVLSTSSSGLLFKSLKLSIPVLQTSPLTPDGRSPL 116
S E G + + + DL + L + K LKL+ G +
Sbjct: 115 SAESSSGASSDVTVETLWEDLFPSISYLPRKELEFVQKGLKLAFEA-HHGQKRRSGEPFI 173
Query: 117 SKALSIAMLLADLQMDAEVISAGILREVLE-VGELNLHEIRSQIGSATAHLLHESLRVKN 175
+++A +L +L++D E I AG+L + +E + +I + G+ H++ +V
Sbjct: 174 IHPVAVARILGELELDWESIVAGLLHDTVEDTNFITFEKIEEEFGATVRHIVEGETKVSK 233
Query: 176 FA-----SRVDILDDENAAALRKFCLTYYD-IRALILDLALKLDMMRHLGHLPRYQQQII 229
+ + + D A LR+ L D +R +I+ LA +L MR L H+P ++Q I
Sbjct: 234 LGKLKCKTESETIQDVKADDLRQMFLAMTDEVRVIIVKLADRLHNMRTLCHMPPHKQSSI 293
Query: 230 SLQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKD 289
+ + ++++APLA +G I ELE+LSF Y+ Y V + + ++YK+
Sbjct: 294 AGETLQVFAPLAKLLGMYSIKSELENLSFMYVSAEDYDRVTS----------RIANLYKE 343
Query: 290 ELLESLKSDPILAELVDD----------ISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLG 339
E +++ IL + ++D V+ K YS K LK D N +
Sbjct: 344 HEKELTEANRILVKKIEDDQFLDLVTVNTDVRSVCKETYSIYKAALKSKGSINDYNQIAQ 403
Query: 340 LRVVLNPKSRENA--LEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAV 397
LR+V+ PK L + ++ CY ++ +WK IP KDYI+ PK NGY+SLH V
Sbjct: 404 LRIVVKPKPSVGVGPLCSPQQICYHVLGLVHEIWKPIPRTVKDYIATPKPNGYQSLHTTV 463
Query: 398 DVSEIGRTRPLMEIQIRTTEMDRLAVGGMA 427
+ + + +E+QIRT EMD +A G+A
Sbjct: 464 -IPFLYESMFRLEVQIRTEEMDLIAERGIA 492
>At1g54130 RSH3
Length = 712
Score = 145 bits (367), Expect = 5e-35
Identities = 101/321 (31%), Positives = 169/321 (52%), Gaps = 22/321 (6%)
Query: 116 LSKALSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKN 175
L + AMLLAD+ ++ V+ AGIL + L+ ++ I GS A L+ ++
Sbjct: 238 LQHCVETAMLLADIGANSTVVVAGILHDTLDDSFMSYDYILRTFGSGVADLVEGLSQLSK 297
Query: 176 FASRVDIL-DDENAAALRKFCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQIISLQVM 234
A + A L L D RA+++ LA +L M L LP ++Q + + +
Sbjct: 298 LARENNTACKTVEADRLHTMFLAMADARAVLIKLADRLHNMMTLYALPPVKRQRFAKETL 357
Query: 235 KIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLES 294
+I+APLA+ +G + ++LE+L F++L P + + L +++ ++I ++L ++
Sbjct: 358 EIFAPLANRLGISSWKVKLENLCFKHLHPDQHHEMSDML--EDSFDEAMITSAIEKLEQA 415
Query: 295 LKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRENALE 354
LK + I +V GR+KS YS K+LK ++++D+ GLR++++
Sbjct: 416 LKKEGISYHVVS-----GRHKSLYSIYCKMLKKKLTMDEIHDIHGLRLIVD--------- 461
Query: 355 AGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIR 414
E+ CY+A ++ +W E+P + KDYIS PK NGY+SLH V +G +E+QIR
Sbjct: 462 -NEKDCYKALGVVHKLWSEVPGKLKDYISHPKFNGYQSLHTVV----MGDGTIPLEVQIR 516
Query: 415 TTEMDRLAVGGMASHSLYKAG 435
T EM A G A+H YK G
Sbjct: 517 TKEMHLQAEFGFAAHWRYKEG 537
>At3g14050 putative GTP pyrophosphokinase
Length = 709
Score = 140 bits (353), Expect = 2e-33
Identities = 102/328 (31%), Positives = 174/328 (52%), Gaps = 33/328 (10%)
Query: 116 LSKALSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKN 175
L + AMLLA++ ++ V+ AG+L + ++ ++ I G+ A L+ ++
Sbjct: 234 LQHCVETAMLLANIGANSTVVVAGLLHDTIDDSFMSYDYILRNFGAGVADLVEGVSKL-- 291
Query: 176 FASRVDILDDENAAA--------LRKFCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQ 227
S++ L EN A L L D RA+++ LA +L M+ L L +QQ
Sbjct: 292 --SQLSKLARENNTACKTVEADRLHTMFLAMADARAVLIKLADRLHNMKTLYALSPVKQQ 349
Query: 228 IISLQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVY 287
+ + ++I+APLA+ +G + ++LE+L F++L+P + + T L +++ ++I
Sbjct: 350 RFAKETLEIFAPLANRLGISTWKVQLENLCFKHLYPNQHNEMSTML--EDSFDEAMITSA 407
Query: 288 KDELLESLKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPK 347
++L ++LK I ++ GR+KS YS K+LK ++++D+ GLR++++
Sbjct: 408 IEKLEQALKKAGISYHVLC-----GRHKSLYSIYSKMLKKKLTVDEIHDIHGLRLIVD-- 460
Query: 348 SRENALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRP 407
E CY+A ++ S+W E+P + KDYI+ PK NGY+SLH V T P
Sbjct: 461 --------NEGDCYKALGVVHSLWSEVPGKLKDYITHPKFNGYQSLHTVV---MDNGTVP 509
Query: 408 LMEIQIRTTEMDRLAVGGMASHSLYKAG 435
L E+QIRT EM A G A+H YK G
Sbjct: 510 L-EVQIRTQEMHLQAEFGFAAHWRYKEG 536
>At2g43290 putative calcium binding protein
Length = 215
Score = 58.2 bits (139), Expect = 1e-08
Identities = 34/91 (37%), Positives = 54/91 (58%), Gaps = 3/91 (3%)
Query: 465 SANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSD 522
SA K I+ + RVF++ DKNGDG+I+ EEL + +E LG P +D M+ +D+N D
Sbjct: 55 SAPTKRIDPSELKRVFQMFDKNGDGRITKEELNDSLENLGIYIPDKDLTQMIHKIDANGD 114
Query: 523 GSLSSDEFQ-MFQKQVEMVRNLEDRDDEYKK 552
G + DEF+ ++ V+ N + ++E K
Sbjct: 115 GCVDIDEFESLYSSIVDEHHNDGETEEEDMK 145
Score = 38.9 bits (89), Expect = 0.008
Identities = 23/68 (33%), Positives = 37/68 (53%), Gaps = 5/68 (7%)
Query: 472 EFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHD----MMLLLDSNSDGSLSS 527
E D +D F + D++GDG I++EEL V+ LG D M++ +D++ DG ++
Sbjct: 141 EEDMKD-AFNVFDQDGDGFITVEELKSVMASLGLKQGKTLDGCKKMIMQVDADGDGRVNY 199
Query: 528 DEFQMFQK 535
EF K
Sbjct: 200 KEFLQMMK 207
>At1g05990 calcium-binding protein, putative
Length = 150
Score = 57.0 bits (136), Expect = 3e-08
Identities = 31/90 (34%), Positives = 52/90 (57%), Gaps = 8/90 (8%)
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQMFQK 535
RVF++ DKNGDG I+ +EL+E + LG P ++ M+ +D N DG + DEF
Sbjct: 8 RVFQMFDKNGDGTITGKELSETLRSLGIYIPDKELTQMIEKIDVNGDGCVDIDEFG---- 63
Query: 536 QVEMVRNLEDRDDEYKKILDEKLHMADDSG 565
E+ + + D +DE ++ + E ++ D +G
Sbjct: 64 --ELYKTIMDEEDEEEEDMKEAFNVFDQNG 91
Score = 40.0 bits (92), Expect = 0.004
Identities = 21/68 (30%), Positives = 35/68 (50%), Gaps = 4/68 (5%)
Query: 472 EFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPG----EDAHDMMLLLDSNSDGSLSS 527
E + F + D+NGDG I+++EL V+ LG +D M+ +D + DG ++
Sbjct: 76 EEEDMKEAFNVFDQNGDGFITVDELKAVLSSLGLKQGKTLDDCKKMIKKVDVDGDGRVNY 135
Query: 528 DEFQMFQK 535
EF+ K
Sbjct: 136 KEFRQMMK 143
Score = 33.9 bits (76), Expect = 0.26
Identities = 21/77 (27%), Positives = 38/77 (49%), Gaps = 6/77 (7%)
Query: 483 LDKNGDGKISIEELTE----VIEELGAPGEDAHDMMLLLDSNSDGSLSSDEFQ--MFQKQ 536
+D NGDG + I+E E +++E ED + + D N DG ++ DE + +
Sbjct: 49 IDVNGDGCVDIDEFGELYKTIMDEEDEEEEDMKEAFNVFDQNGDGFITVDELKAVLSSLG 108
Query: 537 VEMVRNLEDRDDEYKKI 553
++ + L+D KK+
Sbjct: 109 LKQGKTLDDCKKMIKKV 125
>At4g12860 putative calmodulin
Length = 152
Score = 55.8 bits (133), Expect = 6e-08
Identities = 28/77 (36%), Positives = 47/77 (60%), Gaps = 3/77 (3%)
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEF-QMFQ 534
RVF++ DKNGDGKI+ EL + + +G P + ++M+ +D N DG++ DEF ++Q
Sbjct: 8 RVFQMFDKNGDGKIAKNELKDFFKSVGIMVPENEINEMIAKMDVNGDGAMDIDEFGSLYQ 67
Query: 535 KQVEMVRNLEDRDDEYK 551
+ VE ED + ++
Sbjct: 68 EMVEEKEEEEDMREAFR 84
Score = 40.4 bits (93), Expect = 0.003
Identities = 24/68 (35%), Positives = 37/68 (54%), Gaps = 5/68 (7%)
Query: 472 EFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPG----EDAHDMMLLLDSNSDGSLSS 527
E D R+ FR+ D+NGDG I+ EEL V+ +G ED M+ +D + DG ++
Sbjct: 76 EEDMRE-AFRVFDQNGDGFITDEELRSVLASMGLKQGRTLEDCKKMISKVDVDGDGMVNF 134
Query: 528 DEFQMFQK 535
EF+ +
Sbjct: 135 KEFKQMMR 142
>At5g17480 calcium-binding protein
Length = 83
Score = 53.1 bits (126), Expect = 4e-07
Identities = 26/60 (43%), Positives = 38/60 (63%), Gaps = 1/60 (1%)
Query: 475 QRDRVFRLLDKNGDGKISIEELTEVIEELGA-PGEDAHDMMLLLDSNSDGSLSSDEFQMF 533
+ DR+F+ D NGDGKIS EL E ++ LG+ +D MM +D++ DG++S EF F
Sbjct: 9 EHDRIFKKFDANGDGKISAAELEEALKTLGSVTADDVKRMMAEIDTDGDGNISYQEFTDF 68
>At3g59440 calmodulin-like protein
Length = 195
Score = 53.1 bits (126), Expect = 4e-07
Identities = 32/94 (34%), Positives = 50/94 (53%), Gaps = 8/94 (8%)
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQMFQK 535
RVF++ DKNGDG+I+ EEL + +E LG P +D M+ +D+N DG + +EF+
Sbjct: 54 RVFQMFDKNGDGRITKEELNDSLENLGIFMPDKDLIQMIQKMDANGDGCVDINEFESLYG 113
Query: 536 QVEMVRNLEDRDDEYKKILDEKLHMADDSGLIQV 569
+ + D D + + D+ D G I V
Sbjct: 114 SIVEEKEEGDMRDAF-NVFDQ-----DGDGFITV 141
Score = 41.2 bits (95), Expect = 0.002
Identities = 24/68 (35%), Positives = 37/68 (54%), Gaps = 5/68 (7%)
Query: 472 EFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPG----EDAHDMMLLLDSNSDGSLSS 527
E D RD F + D++GDG I++EEL V+ LG E +M++ +D + DG ++
Sbjct: 121 EGDMRD-AFNVFDQDGDGFITVEELNSVMTSLGLKQGKTLECCKEMIMQVDEDGDGRVNY 179
Query: 528 DEFQMFQK 535
EF K
Sbjct: 180 KEFLQMMK 187
>At4g03290 putative calmodulin
Length = 154
Score = 51.6 bits (122), Expect = 1e-06
Identities = 29/91 (31%), Positives = 52/91 (56%), Gaps = 6/91 (6%)
Query: 477 DRVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQMFQ 534
+RVF++ DK+GDGKI+ +EL E + LG P ++ ++ +D N DG + +EF
Sbjct: 7 NRVFQMFDKDGDGKITTKELNESFKNLGIIIPEDELTQIIQKIDVNGDGCVDIEEFGELY 66
Query: 535 KQVEMVRNLEDRDDEYKKILDEKLHMADDSG 565
K + +ED D+ ++ + E ++ D +G
Sbjct: 67 KTI----MVEDEDEVGEEDMKEAFNVFDRNG 93
Score = 40.8 bits (94), Expect = 0.002
Identities = 20/61 (32%), Positives = 35/61 (56%), Gaps = 4/61 (6%)
Query: 480 FRLLDKNGDGKISIEELTEVIEELGAPG----EDAHDMMLLLDSNSDGSLSSDEFQMFQK 535
F + D+NGDG I+++EL V+ LG E+ M++ +D + DG ++ EF+ K
Sbjct: 86 FNVFDRNGDGFITVDELKAVLSSLGLKQGKTLEECRKMIMQVDVDGDGRVNYMEFRQMMK 145
Query: 536 Q 536
+
Sbjct: 146 K 146
Score = 35.0 bits (79), Expect = 0.11
Identities = 20/64 (31%), Positives = 34/64 (52%), Gaps = 6/64 (9%)
Query: 474 DQRDRVFRLLDKNGDGKISIEELTE-----VIEELGAPG-EDAHDMMLLLDSNSDGSLSS 527
D+ ++ + +D NGDG + IEE E ++E+ G ED + + D N DG ++
Sbjct: 40 DELTQIIQKIDVNGDGCVDIEEFGELYKTIMVEDEDEVGEEDMKEAFNVFDRNGDGFITV 99
Query: 528 DEFQ 531
DE +
Sbjct: 100 DELK 103
>At3g07490 putative calmodulin
Length = 153
Score = 51.6 bits (122), Expect = 1e-06
Identities = 27/74 (36%), Positives = 45/74 (60%), Gaps = 7/74 (9%)
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQMFQK 535
R+F++ D+NGDGKI+ +EL + +E LG P +D M+ +D N DG + +EF +
Sbjct: 8 RIFQMFDRNGDGKITKQELNDSLENLGIYIPDKDLVQMIEKIDLNGDGYVDIEEFGGLYQ 67
Query: 536 QVEMVRNLEDRDDE 549
+ +E+RD+E
Sbjct: 68 TI-----MEERDEE 76
Score = 39.3 bits (90), Expect = 0.006
Identities = 24/68 (35%), Positives = 36/68 (52%), Gaps = 5/68 (7%)
Query: 472 EFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPG----EDAHDMMLLLDSNSDGSLSS 527
E D R+ F + D+N DG I++EEL V+ LG ED M+ +D + DG ++
Sbjct: 76 EEDMRE-AFNVFDQNRDGFITVEELRSVLASLGLKQGRTLEDCKRMISKVDVDGDGMVNF 134
Query: 528 DEFQMFQK 535
EF+ K
Sbjct: 135 KEFKQMMK 142
Score = 34.3 bits (77), Expect = 0.20
Identities = 27/88 (30%), Positives = 42/88 (47%), Gaps = 7/88 (7%)
Query: 465 SANHKGIEFDQRDRVFRL--LDKNGDGKISIEE---LTEVIEELGAPGEDAHDMMLLLDS 519
S + GI +D V + +D NGDG + IEE L + I E ED + + D
Sbjct: 29 SLENLGIYIPDKDLVQMIEKIDLNGDGYVDIEEFGGLYQTIMEERDEEEDMREAFNVFDQ 88
Query: 520 NSDGSLSSDEFQ--MFQKQVEMVRNLED 545
N DG ++ +E + + ++ R LED
Sbjct: 89 NRDGFITVEELRSVLASLGLKQGRTLED 116
>At3g03430 pollen allergen Bra r II
Length = 83
Score = 50.1 bits (118), Expect = 3e-06
Identities = 25/57 (43%), Positives = 36/57 (62%), Gaps = 1/57 (1%)
Query: 475 QRDRVFRLLDKNGDGKISIEELTEVIEELGA-PGEDAHDMMLLLDSNSDGSLSSDEF 530
+ DR+F+ D NGDGKIS EL + ++ LG+ ED MM +D++ DG +S EF
Sbjct: 9 EHDRIFKKFDANGDGKISAAELGDALKNLGSVTHEDIKRMMAEIDTDGDGYISYQEF 65
>At5g37770 CALMODULIN-RELATED PROTEIN 2, TOUCH-INDUCED (TCH2)
Length = 161
Score = 48.5 bits (114), Expect = 1e-05
Identities = 27/59 (45%), Positives = 35/59 (58%), Gaps = 2/59 (3%)
Query: 474 DQRDRVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEF 530
D +VF+ DKNGDGKIS++EL EVI L A E+ MM D + +G + DEF
Sbjct: 16 DDIKKVFQRFDKNGDGKISVDELKEVIRALSPTASPEETVTMMKQFDLDGNGFIDLDEF 74
Score = 37.7 bits (86), Expect = 0.018
Identities = 20/54 (37%), Positives = 33/54 (61%), Gaps = 2/54 (3%)
Query: 480 FRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQ 531
F L D +G+G+IS +EL V++ LG +D M+ +D + DG ++ DEF+
Sbjct: 99 FELYDLDGNGRISAKELHSVMKNLGEKCSVQDCKKMISKVDIDGDGCVNFDEFK 152
>At1g66400 calmodulin-related protein
Length = 157
Score = 45.4 bits (106), Expect = 8e-05
Identities = 30/73 (41%), Positives = 40/73 (54%), Gaps = 3/73 (4%)
Query: 478 RVFRLLDKNGDGKISIEELTEVIEEL--GAPGEDAHDMMLLLDSNSDGSLSSDEF-QMFQ 534
+VF+ DKN DGKISI+EL +VI L A E+ MM D + +G + DEF +FQ
Sbjct: 18 KVFQRFDKNNDGKISIDELKDVIGALSPNASQEETKAMMKEFDLDGNGFIDLDEFVALFQ 77
Query: 535 KQVEMVRNLEDRD 547
+ N RD
Sbjct: 78 ISDQSSNNSAIRD 90
Score = 33.5 bits (75), Expect = 0.33
Identities = 25/80 (31%), Positives = 39/80 (48%), Gaps = 4/80 (5%)
Query: 454 ELAALRLKDFPSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPG--EDAH 511
E AL S+N+ I F L D + +G+IS EL V++ LG +D
Sbjct: 71 EFVALFQISDQSSNNSAIR--DLKEAFDLYDLDRNGRISANELHSVMKNLGEKCSIQDCQ 128
Query: 512 DMMLLLDSNSDGSLSSDEFQ 531
M+ +DS+ DG + +EF+
Sbjct: 129 RMINKVDSDGDGCVDFEEFK 148
>At4g14640 calmodulin
Length = 151
Score = 45.1 bits (105), Expect = 1e-04
Identities = 30/90 (33%), Positives = 48/90 (53%), Gaps = 7/90 (7%)
Query: 480 FRLLDKNGDGKISIEELTEVIEEL--GAPGEDAHDMMLLLDSNSDGSLSSDEFQMFQKQV 537
F L DK+GDG I++EEL VI L ++ HD++ +DS+S+G++ EF
Sbjct: 18 FCLFDKDGDGCITVEELATVIRSLDQNPTEQELHDIITEIDSDSNGTIEFAEFLNL---- 73
Query: 538 EMVRNLEDRDDEYKKILDEKLHMADDSGLI 567
M + L++ D E + K+ D +G I
Sbjct: 74 -MAKKLQESDAEEELKEAFKVFDKDQNGYI 102
Score = 35.4 bits (80), Expect = 0.088
Identities = 22/73 (30%), Positives = 39/73 (53%), Gaps = 8/73 (10%)
Query: 474 DQRDRVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQ 531
++ F++ DK+ +G IS EL+ V+ LG E+ M+ D + DG ++ DEF
Sbjct: 85 EELKEAFKVFDKDQNGYISASELSHVMINLGEKLTDEEVEQMIKEADLDGDGQVNYDEF- 143
Query: 532 MFQKQVEMVRNLE 544
V+M+ N++
Sbjct: 144 -----VKMMINID 151
>At1g21550 unknown protein
Length = 155
Score = 44.3 bits (103), Expect = 2e-04
Identities = 25/58 (43%), Positives = 33/58 (56%), Gaps = 4/58 (6%)
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELG----APGEDAHDMMLLLDSNSDGSLSSDEFQ 531
R F + D NGDG IS EEL +V+E LG A D M+ + D N DG + +EF+
Sbjct: 92 RAFNVFDVNGDGYISAEELRDVLERLGFEEEAKAWDCGRMIRVHDKNLDGFVDFEEFK 149
Score = 37.0 bits (84), Expect = 0.030
Identities = 19/56 (33%), Positives = 33/56 (58%), Gaps = 3/56 (5%)
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMMLLLDSNSDGSLSSDEFQMF 533
R+F+ LDKN DG ++++EL ++++LG ++ L++ SL DEF F
Sbjct: 13 RMFKTLDKNQDGLVTLDELLWILDKLGWAEHTPDELELIVGKQ---SLDLDEFLRF 65
>At1g24620 putative calmodulin
Length = 186
Score = 43.9 bits (102), Expect = 2e-04
Identities = 31/99 (31%), Positives = 48/99 (48%), Gaps = 11/99 (11%)
Query: 472 EFDQRDRVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDE 529
E + + VF+ D NGDGKIS +EL ++ LG P E+ + +D DG ++ +E
Sbjct: 34 EIRELEAVFKKFDVNGDGKISSKELGAIMTSLGHEVPEEELEKAITEIDRKGDGYINFEE 93
Query: 530 FQMF----QKQVEMVRNLEDRDDEYK-----KILDEKLH 559
F Q +++ NL+D Y I E+LH
Sbjct: 94 FVELNTKGMDQNDVLENLKDAFSVYDIDGNGSISAEELH 132
Score = 38.5 bits (88), Expect = 0.010
Identities = 26/83 (31%), Positives = 44/83 (52%), Gaps = 10/83 (12%)
Query: 458 LRLKDFPSANHKGIEFDQRDRV------FRLLDKNGDGKISIEELTEVIEELGAPGEDAH 511
+ ++F N KG+ DQ D + F + D +G+G IS EEL EV+ LG A
Sbjct: 89 INFEEFVELNTKGM--DQNDVLENLKDAFSVYDIDGNGSISAEELHEVLRSLGDECSIAE 146
Query: 512 DMMLL--LDSNSDGSLSSDEFQM 532
++ +D + DG++ +EF++
Sbjct: 147 CRKMIGGVDKDGDGTIDFEEFKI 169
>At1g18210 unknown protein
Length = 170
Score = 43.5 bits (101), Expect = 3e-04
Identities = 30/101 (29%), Positives = 49/101 (47%), Gaps = 5/101 (4%)
Query: 474 DQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMMLL--LDSNSDGSLSSDEFQ 531
++ +VF D NGDGKIS+ EL V + +G + +L +D++ DG ++ DEF
Sbjct: 22 EELKKVFDQFDSNGDGKISVLELGGVFKAMGTSYTETELNRVLEEVDTDRDGYINLDEFS 81
Query: 532 MFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLIQVYNK 572
+ + D D Y + +K + S L QV N+
Sbjct: 82 TLCRSSSSAAEIRDAFDLYDQ---DKNGLISASELHQVLNR 119
Score = 35.0 bits (79), Expect = 0.11
Identities = 20/54 (37%), Positives = 33/54 (61%), Gaps = 2/54 (3%)
Query: 480 FRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQ 531
F L D++ +G IS EL +V+ LG ED M+ +D++ DG+++ +EFQ
Sbjct: 97 FDLYDQDKNGLISASELHQVLNRLGMSCSVEDCTRMIGPVDADGDGNVNFEEFQ 150
>At5g44460 calmodulin-like protein
Length = 181
Score = 42.7 bits (99), Expect = 5e-04
Identities = 23/65 (35%), Positives = 37/65 (56%), Gaps = 4/65 (6%)
Query: 477 DRVFRLLDKNGDGKISIEELTEVIEELGAPG----EDAHDMMLLLDSNSDGSLSSDEFQM 532
+ F + D++GDG IS EL +V+++LG P E M++ +DSN DG + EF+
Sbjct: 113 EEAFNVFDEDGDGFISAVELQKVLKKLGLPEAGEIEQVEKMIVSVDSNHDGRVDFFEFKN 172
Query: 533 FQKQV 537
+ V
Sbjct: 173 MMQTV 177
Score = 37.4 bits (85), Expect = 0.023
Identities = 23/64 (35%), Positives = 33/64 (50%), Gaps = 4/64 (6%)
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMMLLLDS---NSDGSLSSDEFQMFQ 534
RVF L DKN DG I++EEL++ + LG D D+ +DS L D+F
Sbjct: 31 RVFDLFDKNNDGFITVEELSQALSRLGLDA-DFSDLKSTVDSFIKPDKTGLRFDDFAALH 89
Query: 535 KQVE 538
K ++
Sbjct: 90 KTLD 93
>At4g20780 calcium-binding protein - like
Length = 191
Score = 42.7 bits (99), Expect = 5e-04
Identities = 23/62 (37%), Positives = 35/62 (56%), Gaps = 4/62 (6%)
Query: 480 FRLLDKNGDGKISIEELTEVIEELGAPG----EDAHDMMLLLDSNSDGSLSSDEFQMFQK 535
F++ D+NGDG IS EL V+++LG P E M++ +D N DG + EF+ +
Sbjct: 125 FKVFDENGDGFISARELQTVLKKLGLPEGGEMERVEKMIVSVDRNQDGRVDFFEFKNMMR 184
Query: 536 QV 537
V
Sbjct: 185 TV 186
Score = 40.4 bits (93), Expect = 0.003
Identities = 28/90 (31%), Positives = 48/90 (53%), Gaps = 10/90 (11%)
Query: 452 AAELAALRLKDFPSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAH 511
A + ++ RL+ PS N ++ R+F L DKNGDG I++EEL++ + LG D
Sbjct: 12 ARQSSSFRLRS-PSLNALRLQ-----RIFDLFDKNGDGFITVEELSQALTRLGL-NADLS 64
Query: 512 DMMLLLDS---NSDGSLSSDEFQMFQKQVE 538
D+ ++S + L+ D+F K ++
Sbjct: 65 DLKSTVESYIQPGNTGLNFDDFSSLHKTLD 94
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,551,433
Number of Sequences: 26719
Number of extensions: 534652
Number of successful extensions: 2026
Number of sequences better than 10.0: 116
Number of HSP's better than 10.0 without gapping: 62
Number of HSP's successfully gapped in prelim test: 56
Number of HSP's that attempted gapping in prelim test: 1778
Number of HSP's gapped (non-prelim): 229
length of query: 580
length of database: 11,318,596
effective HSP length: 105
effective length of query: 475
effective length of database: 8,513,101
effective search space: 4043722975
effective search space used: 4043722975
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146862.24


