

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.18 - phase: 0 /pseudo
(155 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
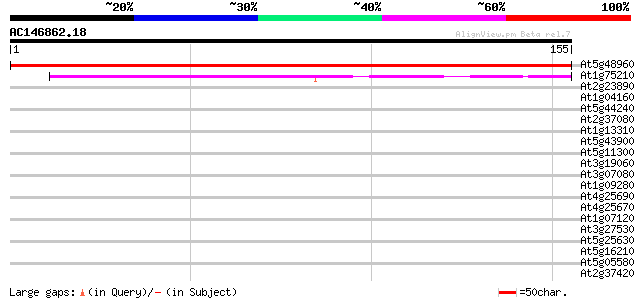
Score E
Sequences producing significant alignments: (bits) Value
At5g48960 unknown protein 270 2e-73
At1g75210 cytosolic IMP-GMP specific 5'-nucleotidase like protein 60 4e-10
At2g23890 hypothetical protein 39 0.001
At1g04160 myosin heavy chain MYA2 31 0.23
At5g44240 ATPase, calcium-transporting 31 0.30
At2g37080 putative myosin heavy chain 30 0.40
At1g13310 hypothetical protein 30 0.68
At5g43900 myosin heavy chain MYA2 (pir||S51824) 28 2.0
At5g11300 cyclin 3b 27 4.4
At3g19060 hypothetical protein 27 4.4
At3g07080 putative integral membrane protein 27 4.4
At1g09280 unknown protein 27 4.4
At4g25690 unknown protein 27 5.7
At4g25670 unknown protein 27 5.7
At1g07120 hypothetical protein 27 5.7
At3g27530 unknown protein 26 7.5
At5g25630 unknown protein 26 9.8
At5g16210 unknown protein 26 9.8
At5g05580 temperature-sensitive omega-3 fatty acid desaturase, c... 26 9.8
At2g37420 putative kinesin heavy chain 26 9.8
>At5g48960 unknown protein
Length = 642
Score = 270 bits (691), Expect = 2e-73
Identities = 133/155 (85%), Positives = 143/155 (91%)
Query: 1 MVENSLGIHGDEILYVGDHIYTDVSQSKVHLRWRTALICRELEDEYSALIRCRGDRESLV 60
M+E+SL +HGDEILYVGDHIYTDVS SKVHLRWRTALICRELE+EY ALI RG RE L+
Sbjct: 447 MIESSLNVHGDEILYVGDHIYTDVSVSKVHLRWRTALICRELEEEYMALIGSRGHREELI 506
Query: 61 ELINQKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQRLDEKI 120
ELINQKEVVGDLFNQLRLALQRRSK RPAQTLAATN+DD++LTE+MQKLLIVMQRLD+KI
Sbjct: 507 ELINQKEVVGDLFNQLRLALQRRSKGRPAQTLAATNLDDQELTETMQKLLIVMQRLDDKI 566
Query: 121 APMLEADGELFNSRWGFLSRAGLWDKSHLMRQIEK 155
MLE DGELFN RWGFLSRAGLWDKSHLMRQIEK
Sbjct: 567 GLMLETDGELFNKRWGFLSRAGLWDKSHLMRQIEK 601
>At1g75210 cytosolic IMP-GMP specific 5'-nucleotidase like protein
Length = 642
Score = 60.5 bits (145), Expect = 4e-10
Identities = 43/146 (29%), Positives = 72/146 (48%), Gaps = 14/146 (9%)
Query: 12 EILYVGDHIYTDVSQSKVHLRWRTALICRELEDEYSALIRCRGDRESLVELINQKEVVGD 71
++LYVGDHIY D+ +SK L WRT L+ ELE E L R R+ L+ + N+++ V D
Sbjct: 472 QVLYVGDHIYGDILRSKKILGWRTMLVVPELEKEVELLWELRNMRKDLILMRNERDSVED 531
Query: 72 LFNQLRLALQRR--SKDRPAQTLAATNMDDEDLTESMQKLLIVMQRLDEKIAPMLEADGE 129
+ L +L+ +++ + L+A +DL K+ Q+ + +
Sbjct: 532 KIHHLNWSLKFEDINENNKHEMLSAL----KDLESKRDKVRQSHQQAQREC-------HQ 580
Query: 130 LFNSRWGFLSRAGLWDKSHLMRQIEK 155
F+ WG L + G + S Q+E+
Sbjct: 581 KFHKVWGQLMKTG-YQSSRFAHQVER 605
>At2g23890 hypothetical protein
Length = 546
Score = 38.5 bits (88), Expect = 0.001
Identities = 17/37 (45%), Positives = 24/37 (63%), Gaps = 1/37 (2%)
Query: 9 HGDEILYVGDHIYTDVSQSKVHLRWRTALICRELEDE 45
HG E++Y GDH+++D+ + WRTA I ELE E
Sbjct: 374 HGPEVIYFGDHLFSDL-RGPSKAGWRTAAIIHELERE 409
>At1g04160 myosin heavy chain MYA2
Length = 1519
Score = 31.2 bits (69), Expect = 0.23
Identities = 24/79 (30%), Positives = 38/79 (47%), Gaps = 18/79 (22%)
Query: 55 DRESLVELINQKEVVGDLFNQLRLALQ----------RRSKDRPAQTLAATNMDDEDLTE 104
D+E + +L N+ E + + + L + + R S+DR Q LAA +
Sbjct: 968 DQELMEKLTNENEKLKGMVSSLEIKIDETAKELHETARISQDRLKQALAAES-------- 1019
Query: 105 SMQKLLIVMQRLDEKIAPM 123
+ KL MQRL+EKI+ M
Sbjct: 1020 KVAKLKTAMQRLEEKISDM 1038
>At5g44240 ATPase, calcium-transporting
Length = 1078
Score = 30.8 bits (68), Expect = 0.30
Identities = 29/108 (26%), Positives = 49/108 (44%), Gaps = 14/108 (12%)
Query: 21 YTDVSQSKVHLRWRTALICRELE-DEYSALIRCRGDR------ESLVEL----INQKEVV 69
+ + S V WR A +C+ LE D Y + DR E++ L IN +
Sbjct: 548 FKEASSLLVDREWRIAEVCQRLEHDLYILGVTAIEDRLQDGVPETIETLRKAGINFWMLT 607
Query: 70 GDLFN---QLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQ 114
GD N Q+ L+ S + Q L +ED++ S++++L+ M+
Sbjct: 608 GDKQNTAIQIALSCNFISPEPKGQLLMIDGKTEEDVSRSLERVLLTMR 655
>At2g37080 putative myosin heavy chain
Length = 583
Score = 30.4 bits (67), Expect = 0.40
Identities = 20/60 (33%), Positives = 32/60 (53%), Gaps = 1/60 (1%)
Query: 22 TDVSQSKVHLRWRTALICRELEDEYSALIRCRGDRESLVELINQKEVVGDLFNQLRLALQ 81
T++ QSK +R L+ R+LE+E A GD S+ EL + V +QL+ A++
Sbjct: 249 TELEQSKSEVRSLEQLV-RQLEEEDEARGNANGDSSSVEELKEEINVARQEISQLKSAVE 307
>At1g13310 hypothetical protein
Length = 323
Score = 29.6 bits (65), Expect = 0.68
Identities = 22/79 (27%), Positives = 37/79 (45%), Gaps = 6/79 (7%)
Query: 36 ALICRELEDEYSALIRCRGDRES---LVELINQKEVVGDLFNQLRLALQRRSKDRPAQT- 91
+L+C EY + + +S L EL+ ++ VGD LR AL K P++
Sbjct: 199 SLLCLAEGQEYVTTMGALHELKSQKYLAELLESEDRVGDAVGVLRRALAAAKKSTPSKDD 258
Query: 92 --LAATNMDDEDLTESMQK 108
+A + ED+ ++M K
Sbjct: 259 KWIAIFKKEREDVAKNMAK 277
>At5g43900 myosin heavy chain MYA2 (pir||S51824)
Length = 1505
Score = 28.1 bits (61), Expect = 2.0
Identities = 20/82 (24%), Positives = 41/82 (49%), Gaps = 3/82 (3%)
Query: 55 DRESLVELINQKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLT--ESMQKLLIV 112
D+E + ++ N+ E + + + L + + K T + + ++ L + KL
Sbjct: 967 DQELMDKITNENEKLKSMVSSLEMKIGETEKKLQETTKISQDRLNQALEAESKLVKLKTA 1026
Query: 113 MQRLDEKIAPMLEADGELFNSR 134
MQRL+EKI M EA+ ++ + +
Sbjct: 1027 MQRLEEKILDM-EAEKKIMHQQ 1047
>At5g11300 cyclin 3b
Length = 434
Score = 26.9 bits (58), Expect = 4.4
Identities = 20/91 (21%), Positives = 45/91 (48%), Gaps = 13/91 (14%)
Query: 59 LVELINQKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQRLDE 118
LV++ +K + + +++R+A AQ ++ +N DE++TE + VM+ L
Sbjct: 105 LVDMHTEKSKLAEDLSKIRMA--------EAQDVSLSNFKDEEITEQQEDGSGVMELLQ- 155
Query: 119 KIAPMLEADGELFNSRWGFLSRAGLWDKSHL 149
+++ D + + + L A ++D H+
Sbjct: 156 ----VVDIDSNVEDPQCCSLYAADIYDNIHV 182
>At3g19060 hypothetical protein
Length = 1647
Score = 26.9 bits (58), Expect = 4.4
Identities = 22/95 (23%), Positives = 44/95 (46%), Gaps = 11/95 (11%)
Query: 37 LICRELEDEYSALI----RCRGDRESLVELINQKEVVGDLFNQLRLALQRRSKDRPAQTL 92
L+ + EYS+ + R D +VE + +K +V + +L+ + D +T+
Sbjct: 981 LVLAQTAKEYSSCLAVVDRELLDHHVIVEDLKEKLIVSQVEGELK---DQCLVDNKLETV 1037
Query: 93 AATNMDDEDLTESMQKLLIVMQRLDEKIAPMLEAD 127
+ E+LTE+ K+ ++ LD + + E D
Sbjct: 1038 SVK----EELTEAQSKIKVLSSDLDRSVQKIAEID 1068
>At3g07080 putative integral membrane protein
Length = 438
Score = 26.9 bits (58), Expect = 4.4
Identities = 13/32 (40%), Positives = 19/32 (58%)
Query: 38 ICRELEDEYSALIRCRGDRESLVELINQKEVV 69
I R LED Y +L+ R R L+EL+ ++ V
Sbjct: 59 IGRYLEDAYGSLLFWRSKRSHLMELVESEKAV 90
>At1g09280 unknown protein
Length = 581
Score = 26.9 bits (58), Expect = 4.4
Identities = 15/39 (38%), Positives = 21/39 (53%), Gaps = 1/39 (2%)
Query: 35 TALICRELEDEYSALIRCRGDRESLVELINQKEVVGDLF 73
+ L+C D+YS RCR R LV + N V GD++
Sbjct: 282 SCLLCNNTFDDYSPRCRCRLCR-MLVLVCNHCRVKGDIY 319
>At4g25690 unknown protein
Length = 191
Score = 26.6 bits (57), Expect = 5.7
Identities = 19/56 (33%), Positives = 30/56 (52%), Gaps = 2/56 (3%)
Query: 71 DLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQRLDEKIAPMLEA 126
D F++++ A + K R + LA + +L + QK L QRLDE+ A + EA
Sbjct: 23 DEFDRIKQA--EKKKRRLEKALATSAAIRAELEKKKQKRLEEQQRLDEEGAAIAEA 76
>At4g25670 unknown protein
Length = 188
Score = 26.6 bits (57), Expect = 5.7
Identities = 19/56 (33%), Positives = 30/56 (52%), Gaps = 2/56 (3%)
Query: 71 DLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQRLDEKIAPMLEA 126
D F++++ A + K R + LA + +L + QK L QRLDE+ A + EA
Sbjct: 23 DEFDRIKQA--EKKKRRLEKALATSAAIRAELEKKKQKRLEEQQRLDEEGAAIAEA 76
>At1g07120 hypothetical protein
Length = 392
Score = 26.6 bits (57), Expect = 5.7
Identities = 21/102 (20%), Positives = 50/102 (48%), Gaps = 16/102 (15%)
Query: 21 YTDVSQSKVHLRWRTALICRELEDEYSALIRCRGDRESLVELINQKEVVGDLFNQLRLAL 80
+TD+S+ + ++W +++E S+L+ R + + +K + LR A
Sbjct: 203 FTDISEVETFVKW--------IDEELSSLVDERAVLKHFPKWPERK------VDSLREAA 248
Query: 81 --QRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQRLDEKI 120
+R K+ + L+ + + LT+++Q++ + RL+E +
Sbjct: 249 CNYKRPKNLGNEILSFKDNPKDSLTQALQRIQSLQDRLEESV 290
>At3g27530 unknown protein
Length = 938
Score = 26.2 bits (56), Expect = 7.5
Identities = 12/27 (44%), Positives = 17/27 (62%)
Query: 40 RELEDEYSALIRCRGDRESLVELINQK 66
+E EDE + L+ C G ES VE ++ K
Sbjct: 884 KESEDELNDLLVCLGQEESKVEKLSAK 910
>At5g25630 unknown protein
Length = 574
Score = 25.8 bits (55), Expect = 9.8
Identities = 15/51 (29%), Positives = 27/51 (52%), Gaps = 1/51 (1%)
Query: 33 WRTALICRELEDEYSALIRCRGDRESLVELINQKEVVGDLFNQLRLALQRR 83
WR A + E +AL +C+ + +E + QK+ G FN L++ + +R
Sbjct: 479 WRVAGLTDESNKAINAL-KCKDIEIAKLEKLYQKQSSGSSFNLLQIPVGKR 528
>At5g16210 unknown protein
Length = 1180
Score = 25.8 bits (55), Expect = 9.8
Identities = 17/72 (23%), Positives = 36/72 (49%), Gaps = 1/72 (1%)
Query: 42 LEDEYSALIRCRGDRESLVELINQKEVVGDLFNQLRLALQRRSKDRPAQTLA-ATNMDDE 100
L E A ++ G ++S+ + +E+ D F R +QR+ KD + N + +
Sbjct: 93 LAQEDIARLKTEGQKKSVPSIDKSEEMDSDEFGGNRPEIQRKKKDFSFTDIGPLKNNERQ 152
Query: 101 DLTESMQKLLIV 112
DL ++++ L++
Sbjct: 153 DLNCAVKEYLLL 164
>At5g05580 temperature-sensitive omega-3 fatty acid desaturase,
chloroplast precursor (sp|P48622)
Length = 435
Score = 25.8 bits (55), Expect = 9.8
Identities = 11/26 (42%), Positives = 14/26 (53%)
Query: 119 KIAPMLEADGELFNSRWGFLSRAGLW 144
K P L+ L NSR+GF S+ W
Sbjct: 34 KFNPPLKPPSSLLNSRYGFYSKTRNW 59
>At2g37420 putative kinesin heavy chain
Length = 1022
Score = 25.8 bits (55), Expect = 9.8
Identities = 24/103 (23%), Positives = 45/103 (43%), Gaps = 10/103 (9%)
Query: 28 KVHLRWRTALICRELEDEYSALIRCRGDRESLVELINQKEVVGDLFNQLRLALQRRSKDR 87
+V L+++T +I R L + L+ + + +V + +KEV+ +L R+K
Sbjct: 477 RVFLKFQTFMIQRNLHNSNKDLLDLKENYIQVVSKLKEKEVIVSRMKASETSLIDRAKGL 536
Query: 88 PAQTLAATN--------MDDEDLTES--MQKLLIVMQRLDEKI 120
A+N +D +D ES LL +LD+ +
Sbjct: 537 RCDLQHASNDINSLFTRLDQKDKLESDNQSMLLKFGSQLDQNL 579
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.136 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,261,892
Number of Sequences: 26719
Number of extensions: 121108
Number of successful extensions: 366
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 356
Number of HSP's gapped (non-prelim): 22
length of query: 155
length of database: 11,318,596
effective HSP length: 91
effective length of query: 64
effective length of database: 8,887,167
effective search space: 568778688
effective search space used: 568778688
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146862.18


