

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146853.5 - phase: 0 /pseudo
(235 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
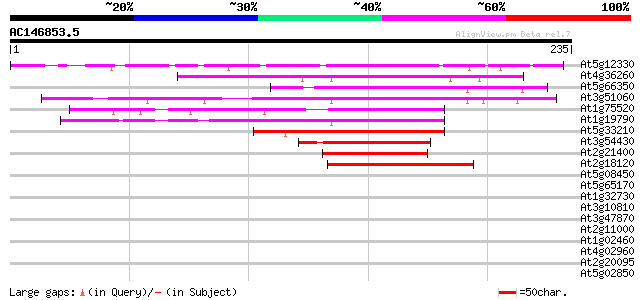
Score E
Sequences producing significant alignments: (bits) Value
At5g12330 lateral root primordia (LRP1) 138 3e-33
At4g36260 STYLISH2 protein (STY2) 103 1e-22
At5g66350 zinc finger protein SHI-like 93 1e-19
At3g51060 STYLISH 1 protein (STY1) 92 2e-19
At1g75520 lateral root primordium protein LRP1 like 92 2e-19
At1g19790 hypothetical protein 92 2e-19
At5g33210 putative protein 91 7e-19
At3g54430 putative protein 88 4e-18
At2g21400 unknown protein 80 9e-16
At2g18120 unknown protein 78 4e-15
At5g08450 unknown protein 38 0.004
At5g65170 putative protein 36 0.015
At1g32730 unknown protein 35 0.034
At3g10810 putative RING zinc finger protein 35 0.045
At3g47870 putative protein 34 0.077
At2g11000 unknown protein 33 0.13
At1g02460 polygalacturonase PG1, putative 33 0.13
At4g02960 putative polyprotein of LTR transposon 33 0.17
At2g20095 bHLH - like protein (bHLH133) 32 0.22
At5g02850 unknown protein 32 0.29
>At5g12330 lateral root primordia (LRP1)
Length = 320
Score = 138 bits (347), Expect = 3e-33
Identities = 100/241 (41%), Positives = 133/241 (54%), Gaps = 36/241 (14%)
Query: 1 MSMLGLRDLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDH-NSNHSLPSSASLSVGFGI 59
M M+GLRD+ L+AP+ +HHQ N DH NSN ++A+ ++G G+
Sbjct: 1 MGMVGLRDVFLVAPA-----YHHQ-------NAGVISGSDHMNSN----AAAAAALGVGV 44
Query: 60 FPLLTATPCMPQQQSQNNEVQENPSNNNNFW-NLRMCPEVVNLPKKGVITEEENHGKIAM 118
PLLTA PQQ +++++ N NN W N E L K T + G +
Sbjct: 45 IPLLTAGT--PQQNVEDSDI--NFLGNNRRWQNNNNNHETQYLHFKS--TNQTTVGTSSN 98
Query: 119 MESEENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRRER 178
+G G+ CQDCGN+AKK+C RRCRTCCK RG+DCSTH+KSTW+ + RRRER
Sbjct: 99 NSGSGSGASGTA--TCQDCGNQAKKECKQRRCRTCCKSRGFDCSTHVKSTWVSAARRRER 156
Query: 179 EVEMFAGGGDGEG-----CSGVKRQKGLLGS--SQNAAATSHSSNSNGTTPKSFATSSCH 231
+V M G G SG K+ + ++GS Q ATSH+S SN T P+SF TSS
Sbjct: 157 QV-MPTGANPTAGSSLSTSSGTKKPR-IVGSQQQQQQQATSHTSTSN-TPPQSFETSSSR 213
Query: 232 Q 232
Q
Sbjct: 214 Q 214
>At4g36260 STYLISH2 protein (STY2)
Length = 322
Score = 103 bits (256), Expect = 1e-22
Identities = 59/167 (35%), Positives = 87/167 (51%), Gaps = 22/167 (13%)
Query: 71 QQQSQNNEVQENPSNNNNFWNLRMCPEVVNLPKKGVITEEENHGKIAMMES------EEN 124
QQQ + N V + N N N +V +P + ++ +H + ++ + +
Sbjct: 22 QQQQKTNWVWYRSNANTNNINPSSSQQVWQIPPEQMLMHHHSHPQQQSLDLYPGHQIDVS 81
Query: 125 GVYGSEYRV---CQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVE 181
+ S + C+DCGN+AKKDC RCRTCCK RG+DCSTH++STWIP RRRER+ +
Sbjct: 82 DLATSSRSITISCRDCGNQAKKDCTHMRCRTCCKSRGFDCSTHVRSTWIPVARRRERQQQ 141
Query: 182 MF---AGGGDGEGCSGV----------KRQKGLLGSSQNAAATSHSS 215
+ +GGG G G G R L G+S ++ SHS+
Sbjct: 142 LHMSTSGGGGGSGSGGAGGGGSSIPKRHRDPTLPGTSSSSRLPSHSA 188
>At5g66350 zinc finger protein SHI-like
Length = 331
Score = 93.2 bits (230), Expect = 1e-19
Identities = 50/119 (42%), Positives = 71/119 (59%), Gaps = 7/119 (5%)
Query: 110 EENHGKIAMMESEENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTW 169
E + G + MM S GS CQDCGN++KKDC RCRTCCK RG DC TH+KSTW
Sbjct: 100 ETSGGALMMMRSGS----GSGGPSCQDCGNQSKKDCSHMRCRTCCKSRGLDCPTHVKSTW 155
Query: 170 IPSTRRREREVEMFAGGGDGE-GCSGVKRQKGLLGSSQNAAATSH--SSNSNGTTPKSF 225
+P+ +RRER+ ++ G + G S KRQ+ + + + A + S+N++G +F
Sbjct: 156 VPAAKRRERQQQLSTGQQPQQLGGSVPKRQRERIPARSTSMAYTRIPSNNTSGLEVGNF 214
>At3g51060 STYLISH 1 protein (STY1)
Length = 252
Score = 92.0 bits (227), Expect = 2e-19
Identities = 74/234 (31%), Positives = 104/234 (43%), Gaps = 36/234 (15%)
Query: 14 PSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVG-FGIFPLLTATPCMPQQ 72
P P S +N N N N N N N + PSS++ ++G ++ M Q
Sbjct: 31 PPPVSEAWLWYRNPNVNANANT------NVNANAPSSSNAALGTLELWQNHNQQEIMFQH 84
Query: 73 QSQNNEVQ---------ENPSNNNNFWNLRMCPEVVNLPKKGVITEEENHGKIAMMESEE 123
Q + PSN+N F G + AMM
Sbjct: 85 QQHQQRLDLYSSAAGLGVGPSNHNQF------------DISGETSTAGAGRAAAMMMIRS 132
Query: 124 NGVYGSEYRV-CQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEM 182
G G V CQDCGN+AKKDC RCRTCCK RG++CSTH++STW+P+ +RRER+ ++
Sbjct: 133 GGSGGGSGGVSCQDCGNQAKKDCSHMRCRTCCKSRGFECSTHVRSTWVPAAKRRERQQQL 192
Query: 183 FAGGGDGE---GCSGVKR-QKGLLGSSQNAAAT---SHSSNSNGTTPKSFATSS 229
+ G S KR ++ L +S + T SHS G P ++S+
Sbjct: 193 ATVQPQTQLPRGESVPKRHRENLPATSSSLVCTRIPSHSGLEVGNFPAEVSSSA 246
>At1g75520 lateral root primordium protein LRP1 like
Length = 346
Score = 92.0 bits (227), Expect = 2e-19
Identities = 58/175 (33%), Positives = 84/175 (47%), Gaps = 32/175 (18%)
Query: 26 NQNQNQNQNQPISHDHN----SNHSLPSSASL---SVGFGIFPLLTATPCMPQQQS--QN 76
N N N+ + + DH+ SN+ + + GF I+P P QQQ Q
Sbjct: 13 NSNNNKQDHHQVDKDHHHQDKSNYLYLYKDEIYNNNKGFEIWP-----PQYFQQQEHQQQ 67
Query: 77 NEVQENPSNNNNFWNLRMCPEVVNLPKKG---------VITEEENHGKIAMMESEENGVY 127
+ Q++ S NF++ M P + V+++ E G + N
Sbjct: 68 QQQQQHASAPANFYSFGMVPSGSSSNNNNNRSRSLYFNVVSDHEPGGFTVTRQGGMN--- 124
Query: 128 GSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEM 182
CQDCGN+AKKDC RCRTCCK RG+ C TH+KSTW+P+ +RRER ++
Sbjct: 125 ------CQDCGNQAKKDCPHMRCRTCCKSRGFHCQTHVKSTWVPAAKRRERLAQL 173
>At1g19790 hypothetical protein
Length = 345
Score = 92.0 bits (227), Expect = 2e-19
Identities = 54/166 (32%), Positives = 83/166 (49%), Gaps = 15/166 (9%)
Query: 22 HHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPCMPQQQSQNNEVQE 81
H++Q+ +Q ++ N+ S+++ + + + GF IFP P QQQ Q N
Sbjct: 12 HNKQDHHQEKDHNEDKSNNYLYLYK-DEIYNNNKGFEIFP-----PQYFQQQQQQNHA-- 63
Query: 82 NPSNNNNFWNLRMCPEVVNLPKKGVITEEENHGKIAMMESEENGVYGSEYRV-----CQD 136
+ N ++ M P N+ ++ E + G CQD
Sbjct: 64 --AAPTNLYSFGMVPSGGNINNNRSTNRSLYFNVVSDHEPVRSSTGGFTVTRQGNMNCQD 121
Query: 137 CGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEM 182
CGN+AKKDC RCRTCCK RG+DC TH+KSTW+ + +RRER+ ++
Sbjct: 122 CGNQAKKDCPHMRCRTCCKSRGFDCQTHVKSTWVSAAKRRERQAQL 167
>At5g33210 putative protein
Length = 173
Score = 90.5 bits (223), Expect = 7e-19
Identities = 41/81 (50%), Positives = 55/81 (67%), Gaps = 1/81 (1%)
Query: 103 KKGVITEEENHG-KIAMMESEENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDC 161
K+G T + + AMM G GS CQD GN+AKKDC RCRTCCK RG++C
Sbjct: 20 KRGCATHPRSIAERAAMMMIRSGGSGGSGGVSCQDFGNQAKKDCSHMRCRTCCKSRGFEC 79
Query: 162 STHLKSTWIPSTRRREREVEM 182
STH++STW+P+T+RRER+ ++
Sbjct: 80 STHVRSTWVPATKRRERQQQL 100
>At3g54430 putative protein
Length = 183
Score = 87.8 bits (216), Expect = 4e-18
Identities = 36/55 (65%), Positives = 44/55 (79%), Gaps = 2/55 (3%)
Query: 122 EENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRR 176
E+N G +VC+DCGNRAKK+C+F RCRTCCK RGY+C TH+KSTWIPS+ R
Sbjct: 31 EDNNTVGE--KVCRDCGNRAKKECLFERCRTCCKSRGYNCVTHVKSTWIPSSATR 83
>At2g21400 unknown protein
Length = 174
Score = 80.1 bits (196), Expect = 9e-16
Identities = 29/44 (65%), Positives = 37/44 (83%)
Query: 132 RVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRR 175
R C+DCGN+AKKDCV+ RCRTCCK + + C TH+KSTW+P+ RR
Sbjct: 7 RKCEDCGNQAKKDCVYMRCRTCCKSKAFHCQTHIKSTWVPAYRR 50
>At2g18120 unknown protein
Length = 222
Score = 78.2 bits (191), Expect = 4e-15
Identities = 32/61 (52%), Positives = 41/61 (66%)
Query: 134 CQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEMFAGGGDGEGCS 193
CQ+CGN+AKK C RCRTCCK G C TH++STWIP +RRER+ ++ + G S
Sbjct: 72 CQECGNQAKKGCTHGRCRTCCKSNGLHCPTHVRSTWIPIAKRRERQQQLQTPTSNPTGGS 131
Query: 194 G 194
G
Sbjct: 132 G 132
>At5g08450 unknown protein
Length = 918
Score = 38.1 bits (87), Expect = 0.004
Identities = 42/168 (25%), Positives = 72/168 (42%), Gaps = 25/168 (14%)
Query: 12 IAPSPSSLQHHHQQ----NQNQNQNQNQP----ISHDHNSNHS---LPSSASLSVGF--- 57
+ P P+ + H+HQQ Q+Q+Q+Q QP + H H+ +HS L ++AS S +
Sbjct: 41 VTPPPAQVHHNHQQPHQHPQSQSQSQPQPHLQALPHPHSHSHSHSPLAAAASASAPYEVE 100
Query: 58 --GIFPLLTATPCMPQQQSQNNEVQENPSNNNNFWNLRMCPEVVNLPKKGVITEEENHGK 115
+ + + P +++S V +PS + P + + P + G+
Sbjct: 101 SRTVVKVARSEPRDGERRSPLPLVYRSPSLPTTVSS--SDPHLTHAPVPMEPRDGAKDGR 158
Query: 116 IAMMESEEN-----GVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRG 158
+ES EN +YG R Q G + +D F R G+G
Sbjct: 159 EIRVESRENRSDGREIYGETKREIQ--GPKGDRDVKFERSVDDFSGKG 204
>At5g65170 putative protein
Length = 362
Score = 36.2 bits (82), Expect = 0.015
Identities = 26/61 (42%), Positives = 30/61 (48%), Gaps = 13/61 (21%)
Query: 11 LIAPS--------PSSLQHHHQQNQNQNQNQNQPISHDHNSNHSL-----PSSASLSVGF 57
LI+PS PSS HHH Q+QN N N P + N+SL PSS S GF
Sbjct: 209 LISPSTLNHHYLPPSSEYHHHHQHQNLLLNMNTPNISNPFLNNSLTDQPKPSSLRTSNGF 268
Query: 58 G 58
G
Sbjct: 269 G 269
>At1g32730 unknown protein
Length = 327
Score = 35.0 bits (79), Expect = 0.034
Identities = 17/54 (31%), Positives = 24/54 (43%), Gaps = 8/54 (14%)
Query: 134 CQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHL--------KSTWIPSTRRRERE 179
C CGN A+ C F+ C+ CC C H+ + T PST E++
Sbjct: 31 CIQCGNVARSRCPFQSCKGCCSRAENPCPIHVLKVASTSGEKTQAPSTPSSEQK 84
>At3g10810 putative RING zinc finger protein
Length = 684
Score = 34.7 bits (78), Expect = 0.045
Identities = 19/82 (23%), Positives = 37/82 (44%), Gaps = 5/82 (6%)
Query: 13 APSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPCMPQQ 72
+PSP S HHH + + + + H HN +H + S + + P+ + P ++
Sbjct: 512 SPSPHSKHHHHHHHHHHHHHH-----HHHNHHHHHHHNLSPKMAPEVSPVASPAPHRSRK 566
Query: 73 QSQNNEVQENPSNNNNFWNLRM 94
++ + NP N +F R+
Sbjct: 567 RAPSAPPPCNPGNRVHFKEKRV 588
>At3g47870 putative protein
Length = 328
Score = 33.9 bits (76), Expect = 0.077
Identities = 18/43 (41%), Positives = 22/43 (50%), Gaps = 3/43 (6%)
Query: 20 QHHHQQNQNQNQNQNQPI---SHDHNSNHSLPSSASLSVGFGI 59
Q + QNQNQN N NQ I DHN +S L +G G+
Sbjct: 142 QPQNGQNQNQNHNHNQMIHELGSDHNKQQEDVTSQQLDLGMGL 184
>At2g11000 unknown protein
Length = 695
Score = 33.1 bits (74), Expect = 0.13
Identities = 16/51 (31%), Positives = 30/51 (58%), Gaps = 1/51 (1%)
Query: 25 QNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPCMPQQQSQ 75
Q+ ++++ + PI HD ++ S+PS + SV + PLL+A C Q+ +
Sbjct: 2 QSVREDEDSSSPIHHDSKTSSSIPSGDNNSVWADVSPLLSAA-CSDLQEGE 51
>At1g02460 polygalacturonase PG1, putative
Length = 491
Score = 33.1 bits (74), Expect = 0.13
Identities = 23/91 (25%), Positives = 42/91 (45%), Gaps = 9/91 (9%)
Query: 3 MLGLRDLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPL 62
+LG L+L S S ++HH + ++++ N H+H+S+ P S+S+S P
Sbjct: 11 VLGFMTLILTLMSLSEARNHHHKEKHKHNN------HNHHSSKPKPPSSSISQPPTPPP- 63
Query: 63 LTATPCMPQQQSQNNEVQENPSNNNNFWNLR 93
P P + + +NN +N+R
Sbjct: 64 --GPPDSPAPSLPPSPSDDPADDNNGIYNVR 92
>At4g02960 putative polyprotein of LTR transposon
Length = 1456
Score = 32.7 bits (73), Expect = 0.17
Identities = 22/75 (29%), Positives = 39/75 (51%), Gaps = 3/75 (4%)
Query: 13 APSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPCMPQQ 72
APS + Q Q +Q QN N N PI ++ N N P+S + + P+ ++P +P
Sbjct: 812 APSHNGPQPTAQPHQTQNSNSNSPILNNPNPNSPSPNSPNQNSPLPQSPI--SSPHIPTP 869
Query: 73 QSQNNEVQENPSNNN 87
+ +E +PS+++
Sbjct: 870 STSISE-PNSPSSSS 883
>At2g20095 bHLH - like protein (bHLH133)
Length = 362
Score = 32.3 bits (72), Expect = 0.22
Identities = 22/76 (28%), Positives = 35/76 (45%), Gaps = 13/76 (17%)
Query: 19 LQHHHQQNQNQN------QNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPCMPQQ 72
+ HHH QNQ+Q+ QN + + H ++ SS + S G + ++P P
Sbjct: 114 MNHHHHQNQHQSYQAKRIQNWEEQVLR-HQASMKQESSNNNSYG------IMSSPNSPPN 166
Query: 73 QSQNNEVQENPSNNNN 88
+S + N NNNN
Sbjct: 167 KSCATIINTNEDNNNN 182
>At5g02850 unknown protein
Length = 426
Score = 32.0 bits (71), Expect = 0.29
Identities = 19/64 (29%), Positives = 32/64 (49%), Gaps = 4/64 (6%)
Query: 15 SPSSLQHHHQQNQNQNQNQNQPISHDHN----SNHSLPSSASLSVGFGIFPLLTATPCMP 70
+P+ +H H + + +Q+Q+Q I N S + SSAS S LL+ P +P
Sbjct: 25 NPTPTRHGHPTSSSSSQSQHQQIQQQPNLLPSSTVAAASSASASAAVSSSALLSLLPPLP 84
Query: 71 QQQS 74
+ Q+
Sbjct: 85 RAQA 88
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.128 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,014,730
Number of Sequences: 26719
Number of extensions: 280589
Number of successful extensions: 1332
Number of sequences better than 10.0: 83
Number of HSP's better than 10.0 without gapping: 48
Number of HSP's successfully gapped in prelim test: 35
Number of HSP's that attempted gapping in prelim test: 1150
Number of HSP's gapped (non-prelim): 161
length of query: 235
length of database: 11,318,596
effective HSP length: 96
effective length of query: 139
effective length of database: 8,753,572
effective search space: 1216746508
effective search space used: 1216746508
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146853.5


