

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146819.13 + phase: 0
(97 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
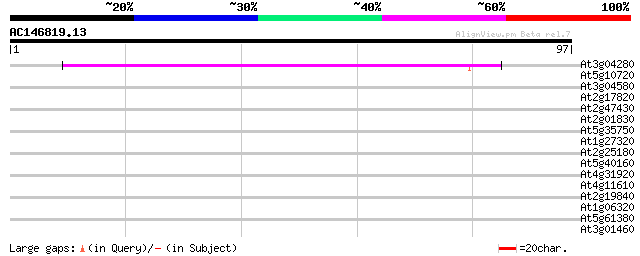
Score E
Sequences producing significant alignments: (bits) Value
At3g04280 putative response regulator protein (receiver component) 55 6e-09
At5g10720 histidine kinase - like protein 37 0.002
At3g04580 putative ethylene receptor 37 0.002
At2g17820 histidine kinase 1 35 0.006
At2g47430 putative histidine kinase 34 0.011
At2g01830 cytokinin receptor CRE1b (CRE1) 34 0.014
At5g35750 histidine kinase (AHK2) 33 0.024
At1g27320 histidine kinase (AHK3) 33 0.032
At2g25180 putative two-component response regulator protein 27 2.3
At5g40160 ankyrin repeat protein EMB506 26 3.0
At4g31920 predicted protein 26 3.9
At4g11610 phosphoribosylanthranilate transferase like protein 26 3.9
At2g19840 copia-like retroelement pol polyprotein 26 3.9
At1g06320 hypothetical protein 26 3.9
At5g61380 pseudo-response regulator 1 25 6.6
At3g01460 unknown protein 25 6.6
>At3g04280 putative response regulator protein (receiver component)
Length = 142
Score = 55.1 bits (131), Expect = 6e-09
Identities = 32/77 (41%), Positives = 46/77 (59%), Gaps = 1/77 (1%)
Query: 10 IGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSR 69
IG + KNG+E++ L E FDLILM + MP DG++ TKKLR+++ IVGV+
Sbjct: 44 IGGISQTAKNGEEAVILHRDGEASFDLILMDKEMPERDGVSTTKKLREMKVTSMIVGVTS 103
Query: 70 HLTNDEYAE-FCNAGLD 85
+E + F AGL+
Sbjct: 104 VADQEEERKAFMEAGLN 120
>At5g10720 histidine kinase - like protein
Length = 950
Score = 37.0 bits (84), Expect = 0.002
Identities = 20/62 (32%), Positives = 33/62 (52%), Gaps = 2/62 (3%)
Query: 1 MILFEWLIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEY 60
M+ + +G NG E++ I++ +DL+LM MPV+DG+ AT+ +R E
Sbjct: 819 MVAKSMMKQLGHTMDIANNGVEAITAINS--SSYDLVLMDVCMPVLDGLKATRLIRSYEE 876
Query: 61 PG 62
G
Sbjct: 877 TG 878
>At3g04580 putative ethylene receptor
Length = 766
Score = 36.6 bits (83), Expect = 0.002
Identities = 17/47 (36%), Positives = 27/47 (57%)
Query: 10 IGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLR 56
+G E AV +G E L+ +S VE + ++++ MP MDG K+R
Sbjct: 664 LGCEVTAVSSGFECLNALSNVEMSYRVVILDLQMPEMDGFEVAMKIR 710
>At2g17820 histidine kinase 1
Length = 1207
Score = 35.0 bits (79), Expect = 0.006
Identities = 21/56 (37%), Positives = 30/56 (53%), Gaps = 4/56 (7%)
Query: 34 FDLILMARIMPVMDGITATKKLRDLEYPGQ----IVGVSRHLTNDEYAEFCNAGLD 85
+DLILM MP MDG ATK +R E + IV ++ H + + A+ G+D
Sbjct: 1121 YDLILMDCQMPKMDGYEATKAIRRAEIGTELHIPIVALTAHAMSSDEAKCLEVGMD 1176
>At2g47430 putative histidine kinase
Length = 1122
Score = 34.3 bits (77), Expect = 0.011
Identities = 30/89 (33%), Positives = 40/89 (44%), Gaps = 16/89 (17%)
Query: 13 ETHAVKNGKESLDLI-----------STVERKFDLILMARIMPVMDGITATKKLRDLE-- 59
E +GKE+L L+ S + FD I M MP MDG AT+++R +E
Sbjct: 1012 EVEQCDSGKEALRLVTEGLTQREEQGSVDKLPFDYIFMDCQMPEMDGYEATREIRKVEKS 1071
Query: 60 --YPGQIVGVSRHLTNDEYA-EFCNAGLD 85
I+ VS H E A E AG+D
Sbjct: 1072 YGVRTPIIAVSGHDPGSEEARETIQAGMD 1100
>At2g01830 cytokinin receptor CRE1b (CRE1)
Length = 844
Score = 33.9 bits (76), Expect = 0.014
Identities = 23/85 (27%), Positives = 41/85 (48%), Gaps = 11/85 (12%)
Query: 11 GVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLR----------DLEY 60
G E ++G+ +L L+ + FD M MP MDG AT+++R +LE+
Sbjct: 732 GAEVVCAESGQVALGLLQ-IPHTFDACFMDIQMPQMDGFEATRQIRMMEKETKEKTNLEW 790
Query: 61 PGQIVGVSRHLTNDEYAEFCNAGLD 85
I+ ++ + + Y E +G+D
Sbjct: 791 HLPILAMTADVIHATYEECLKSGMD 815
>At5g35750 histidine kinase (AHK2)
Length = 1176
Score = 33.1 bits (74), Expect = 0.024
Identities = 18/49 (36%), Positives = 27/49 (54%), Gaps = 1/49 (2%)
Query: 11 GVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLE 59
G V++GK +L ++ FD M MP MDG AT+++R+LE
Sbjct: 1058 GAIVTCVESGKAALAMLKP-PHNFDACFMDLQMPEMDGFEATRRVRELE 1105
>At1g27320 histidine kinase (AHK3)
Length = 1036
Score = 32.7 bits (73), Expect = 0.032
Identities = 16/49 (32%), Positives = 28/49 (56%), Gaps = 1/49 (2%)
Query: 11 GVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLE 59
G + ++G +++ L+ +FD M MP MDG AT+++RD+E
Sbjct: 913 GADVVCAESGIKAISLLKP-PHEFDACFMDIQMPEMDGFEATRRIRDME 960
>At2g25180 putative two-component response regulator protein
Length = 573
Score = 26.6 bits (57), Expect = 2.3
Identities = 16/50 (32%), Positives = 28/50 (56%), Gaps = 1/50 (2%)
Query: 21 KESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSRH 70
+++L+L+ + KFDL++ MP MDG +L LE ++ +S H
Sbjct: 50 QKALELLRENKNKFDLVISDVDMPDMDGFKLL-ELVGLEMDLPVIMLSAH 98
>At5g40160 ankyrin repeat protein EMB506
Length = 315
Score = 26.2 bits (56), Expect = 3.0
Identities = 15/38 (39%), Positives = 21/38 (54%), Gaps = 3/38 (7%)
Query: 13 ETHAVKNGKESLDLISTVERKF---DLILMARIMPVMD 47
+T K+GK +LDL R F DL+ + +IMP D
Sbjct: 277 KTRRTKDGKLALDLALCFGRDFKSYDLVKLLKIMPTGD 314
>At4g31920 predicted protein
Length = 552
Score = 25.8 bits (55), Expect = 3.9
Identities = 16/48 (33%), Positives = 26/48 (53%), Gaps = 1/48 (2%)
Query: 23 SLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSRH 70
+L+L+ + KFDL++ MP MDG +L LE ++ +S H
Sbjct: 52 ALELLRENKNKFDLVISDVDMPDMDGFKLL-ELVGLEMDLPVIMLSAH 98
>At4g11610 phosphoribosylanthranilate transferase like protein
Length = 1011
Score = 25.8 bits (55), Expect = 3.9
Identities = 14/65 (21%), Positives = 30/65 (45%), Gaps = 1/65 (1%)
Query: 15 HAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSRHLTND 74
H K + DL+ + + ++ AR +P+MD + ++ G G++RH
Sbjct: 262 HKDKTATSTYDLVERMYFLYVRVVKARELPIMDITGSVDPFVEVRV-GNYKGITRHFEKR 320
Query: 75 EYAEF 79
++ E+
Sbjct: 321 QHPEW 325
>At2g19840 copia-like retroelement pol polyprotein
Length = 1137
Score = 25.8 bits (55), Expect = 3.9
Identities = 10/25 (40%), Positives = 15/25 (60%)
Query: 69 RHLTNDEYAEFCNAGLDDLHHEDGI 93
+HL D EFCN D++ ++GI
Sbjct: 356 KHLRTDNGLEFCNHKFDEVCKKEGI 380
>At1g06320 hypothetical protein
Length = 180
Score = 25.8 bits (55), Expect = 3.9
Identities = 9/18 (50%), Positives = 13/18 (72%)
Query: 66 GVSRHLTNDEYAEFCNAG 83
G + +T +EY EFCN+G
Sbjct: 4 GGEKKITVEEYVEFCNSG 21
>At5g61380 pseudo-response regulator 1
Length = 618
Score = 25.0 bits (53), Expect = 6.6
Identities = 17/60 (28%), Positives = 30/60 (49%), Gaps = 6/60 (10%)
Query: 3 LFEWLIDIGVETHAVKNGKESLDLISTVERKFDLIL------MARIMPVMDGITATKKLR 56
+F L + + AVK+ ++ +D ++ D+IL MA+ M ++ IT K LR
Sbjct: 34 VFTLLSECSYQVTAVKSARQVIDALNAEGPDIDIILAEIDLPMAKGMKMLRYITRDKDLR 93
>At3g01460 unknown protein
Length = 2176
Score = 25.0 bits (53), Expect = 6.6
Identities = 10/37 (27%), Positives = 22/37 (59%)
Query: 22 ESLDLISTVERKFDLILMARIMPVMDGITATKKLRDL 58
+ +DL++T+ KF + A ++P++ + +KL L
Sbjct: 1226 DCVDLVATLSEKFKSLYEAEVVPLVQKLKDYRKLECL 1262
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.142 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,204,444
Number of Sequences: 26719
Number of extensions: 77726
Number of successful extensions: 193
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 180
Number of HSP's gapped (non-prelim): 16
length of query: 97
length of database: 11,318,596
effective HSP length: 73
effective length of query: 24
effective length of database: 9,368,109
effective search space: 224834616
effective search space used: 224834616
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146819.13


