

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146817.2 + phase: 0
(145 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
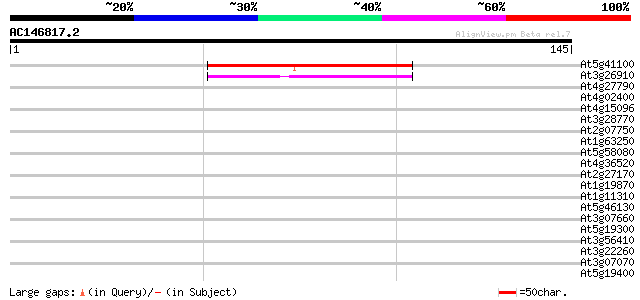
At5g41100 unknown protein 47 3e-06
At3g26910 unknown protein 45 1e-05
At4g27790 putative calcium binding protein 32 0.15
At4g02400 30 0.34
At4g15096 unknown protein 30 0.45
At3g28770 hypothetical protein 30 0.58
At2g07750 putative RNA helicase 29 0.76
At1g63250 putative RNA helicase 28 1.7
At5g58080 ARR2 - like protein 28 2.2
At4g36520 trichohyalin like protein 28 2.2
At2g27170 putative chromosome associated protein 28 2.2
At1g19870 unknown protein 28 2.2
At1g11310 Mlo like protein 28 2.2
At5g46130 putative protein 27 2.9
At3g07660 unknown protein 27 2.9
At5g19300 putative protein 27 3.8
At3g56410 putative protein 27 3.8
At3g22260 unknown protein 27 3.8
At3g07070 putative protein kinase 27 3.8
At5g19400 putative protein 27 4.9
>At5g41100 unknown protein
Length = 582
Score = 47.4 bits (111), Expect = 3e-06
Identities = 22/54 (40%), Positives = 37/54 (67%), Gaps = 1/54 (1%)
Query: 52 DNMKHHCDEKREVYEYMIIQP-KEKGKSNSGKGEHITSRPLQAAHDEYEEEATL 104
++MK C+EKR+V ++M+++ K+K + KGE + R L+ A DE ++EATL
Sbjct: 131 EDMKQQCEEKRDVVKHMLMEHVKDKVQVKGTKGERLIRRQLETARDELQDEATL 184
>At3g26910 unknown protein
Length = 621
Score = 45.4 bits (106), Expect = 1e-05
Identities = 22/53 (41%), Positives = 32/53 (59%), Gaps = 2/53 (3%)
Query: 52 DNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEATL 104
++MK CD KR VYE ++ KEKG+ S KGE + A+ E+ +EAT+
Sbjct: 132 EDMKQQCDGKRNVYEMSLV--KEKGRPKSSKGERHIPPESRPAYSEFHDEATM 182
>At4g27790 putative calcium binding protein
Length = 345
Score = 31.6 bits (70), Expect = 0.15
Identities = 21/92 (22%), Positives = 44/92 (47%), Gaps = 2/92 (2%)
Query: 16 VDKNIKKNGKFGPIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEK 75
++KN K +G+ G N F+ + SL ++E +N H D + + +++ +
Sbjct: 162 IEKNEKGHGEAGWWM--EQFKNSDFDHNGSLDIEEFNNFLHPEDSRNGDTQRWVLKERMT 219
Query: 76 GKSNSGKGEHITSRPLQAAHDEYEEEATLAKQ 107
G +G G+ ++ A++ Y+E A K+
Sbjct: 220 GMDTNGDGKLEYKEFVKNAYEMYKEFAKFEKE 251
>At4g02400
Length = 870
Score = 30.4 bits (67), Expect = 0.34
Identities = 18/63 (28%), Positives = 28/63 (43%)
Query: 18 KNIKKNGKFGPIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEKGK 77
K + KN K +P + I + L E D+ DE ++YEY P+E+ K
Sbjct: 10 KTLAKNKKRKGPHLPNSILKTIANEKRPLNSDEDDDEIDSDDENVDLYEYEEGVPEEESK 69
Query: 78 SNS 80
N+
Sbjct: 70 KNN 72
>At4g15096 unknown protein
Length = 679
Score = 30.0 bits (66), Expect = 0.45
Identities = 18/56 (32%), Positives = 25/56 (44%), Gaps = 2/56 (3%)
Query: 49 KEKDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEATL 104
KE + HH K ++I + SN G GE I R + H+E EE + L
Sbjct: 25 KEPEAPDHHLKHKN--LGLLMIYGNDDHLSNGGIGESIDHRSMNLVHEEREEHSKL 78
>At3g28770 hypothetical protein
Length = 2081
Score = 29.6 bits (65), Expect = 0.58
Identities = 21/92 (22%), Positives = 38/92 (40%)
Query: 16 VDKNIKKNGKFGPIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEK 75
++ + K+ GK + N+ + + K + +K D K+E + + KE+
Sbjct: 933 INTSSKQKGKDKKKKKKESKNSNMKKKEEDKKEYVNNELKKQEDNKKETTKSENSKLKEE 992
Query: 76 GKSNSGKGEHITSRPLQAAHDEYEEEATLAKQ 107
K N K E S EYEE+ + K+
Sbjct: 993 NKDNKEKKESEDSASKNREKKEYEEKKSKTKE 1024
>At2g07750 putative RNA helicase
Length = 845
Score = 29.3 bits (64), Expect = 0.76
Identities = 36/145 (24%), Positives = 62/145 (41%), Gaps = 20/145 (13%)
Query: 8 QIAALTSKVDKNIKKNGKFGPIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEY 67
+I + S ++ +N GP P G+N ++ + K +EK+N D+++++YE
Sbjct: 60 RITSERSLWNRIFSRNMGGGPRTFPGGLNKWQWKRMHEKKAREKENKL--LDQEKQLYEA 117
Query: 68 MI---IQPKEKGKSNSGKGEHITSRPLQAAH---DEYEEEATLA----KQGCPHIAKLGK 117
I I+ K G +SG+ + L+ +H E TLA K G +
Sbjct: 118 RIRTEIRAKMWGHPDSGE----KTAKLKQSHGPMSPKEHIKTLADRFMKAGADDLWNDND 173
Query: 118 GRRIKPLKHSNFNERNCRDKFRQDP 142
G P+K + R+C D P
Sbjct: 174 G----PVKKFDQGSRSCSDSIDSTP 194
>At1g63250 putative RNA helicase
Length = 798
Score = 28.1 bits (61), Expect = 1.7
Identities = 18/59 (30%), Positives = 32/59 (53%), Gaps = 5/59 (8%)
Query: 27 GPIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMI---IQPKEKGKSNSGK 82
GP P G+N ++ + K +EK+N D+++++YE I I+ K G +SG+
Sbjct: 29 GPRTFPGGLNKWQWKRMHEKKAREKENKL--LDQEKQLYEARIRTEIRAKMWGNPDSGE 85
>At5g58080 ARR2 - like protein
Length = 632
Score = 27.7 bits (60), Expect = 2.2
Identities = 12/30 (40%), Positives = 16/30 (53%)
Query: 23 NGKFGPIRVPRGVNNQIFEDSNSLKVKEKD 52
N FG I P +N +F D+NS + K D
Sbjct: 533 NNPFGNIEYPLPADNMVFRDNNSTRSKGLD 562
>At4g36520 trichohyalin like protein
Length = 1432
Score = 27.7 bits (60), Expect = 2.2
Identities = 16/42 (38%), Positives = 20/42 (47%)
Query: 1 MRPSKLHQIAALTSKVDKNIKKNGKFGPIRVPRGVNNQIFED 42
+R + L I T VD K N GP V RG + +FED
Sbjct: 209 IRVADLGAIPGYTVAVDGTTKLNKVTGPSSVGRGTSRFVFED 250
>At2g27170 putative chromosome associated protein
Length = 1163
Score = 27.7 bits (60), Expect = 2.2
Identities = 21/98 (21%), Positives = 45/98 (45%), Gaps = 8/98 (8%)
Query: 12 LTSKVDKNIKKNGKFGPIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQ 71
L +K D+ KK GP+ ++ F+ +KE M H C E+ + + ++ +
Sbjct: 881 LLAKQDEYTKKIRGLGPL------SSDAFDTYKRKNIKELQKMLHRCSEQLQQFSHVNKK 934
Query: 72 PKEKGKSNSGKGEHITSR--PLQAAHDEYEEEATLAKQ 107
++ + + + E + +R L A ++ +E T+ Q
Sbjct: 935 ALDQYVNFTEQREELQNRQAELDAGDEKIKELITVLDQ 972
>At1g19870 unknown protein
Length = 805
Score = 27.7 bits (60), Expect = 2.2
Identities = 22/111 (19%), Positives = 46/111 (40%), Gaps = 7/111 (6%)
Query: 24 GKFGPIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPK------EKGK 77
G P+ + + S+S+K ++ + K +KR+V + + PK E +
Sbjct: 633 GALTPMTITESQATPASQASSSVKARKGKSEKSGSSQKRKVSKKITSSPKQEIGTGEATE 692
Query: 78 SNSGKGEHITSRPLQAAHDEYEEEATLAKQGCPHIAKLGKGRRIKPLKHSN 128
GK E + R +D+ E++ K P + + + K +H++
Sbjct: 693 QEEGK-EQKSGRRTSFGYDQEARESSGGKNSLPRFMQPTQSAKAKVQEHNS 742
>At1g11310 Mlo like protein
Length = 573
Score = 27.7 bits (60), Expect = 2.2
Identities = 18/62 (29%), Positives = 26/62 (41%), Gaps = 3/62 (4%)
Query: 56 HHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEATLAKQGCPHIAKL 115
H C E +Y K+ GK + G G+ R L + Y +LA +G A+
Sbjct: 93 HPCSAAEEAKKY---GKKDAGKKDDGDGDKPGRRLLLELAESYIHRRSLATKGYDKCAEK 149
Query: 116 GK 117
GK
Sbjct: 150 GK 151
>At5g46130 putative protein
Length = 376
Score = 27.3 bits (59), Expect = 2.9
Identities = 13/48 (27%), Positives = 27/48 (56%), Gaps = 2/48 (4%)
Query: 43 SNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRP 90
S L KEKD +H +K+++ +YM+ + +E ++ + +H+ P
Sbjct: 211 SFGLNFKEKDEPTYHIIDKKDLPQYMLYELEE--MNSFTRTDHLVESP 256
>At3g07660 unknown protein
Length = 782
Score = 27.3 bits (59), Expect = 2.9
Identities = 21/74 (28%), Positives = 31/74 (41%), Gaps = 5/74 (6%)
Query: 26 FGPIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEKGKSN----SG 81
+ P N+ ED NSL++KE +N ++ E I P S+
Sbjct: 654 YNPTSAASAGNSTSNEDLNSLQLKE-NNGYSTTGQQSEALPVWITGPGRDVPSSFYGLQH 712
Query: 82 KGEHITSRPLQAAH 95
G+H+T P QA H
Sbjct: 713 HGQHVTYAPAQAGH 726
>At5g19300 putative protein
Length = 440
Score = 26.9 bits (58), Expect = 3.8
Identities = 14/41 (34%), Positives = 21/41 (51%)
Query: 42 DSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEKGKSNSGK 82
DS K+K+K K+ +E V E +Q +EKG G+
Sbjct: 57 DSKEEKIKKKRKNKNQEEEPELVTEKTKVQEEEKGNVEEGR 97
>At3g56410 putative protein
Length = 1535
Score = 26.9 bits (58), Expect = 3.8
Identities = 20/76 (26%), Positives = 37/76 (48%), Gaps = 3/76 (3%)
Query: 41 EDSNSLKVKEKDNMKHHCDEKREVYEYM-IIQPKEKGKSNSGKGEHITSRPLQAAHDEYE 99
++++S K E DN + CD + YE II+P+E SG+ + + + E
Sbjct: 554 QEAHSEKSTEMDN--NICDIEEPEYEKNEIIKPEEIVGEGSGESLEVPVNQNERMSESSE 611
Query: 100 EEATLAKQGCPHIAKL 115
E+ +A++ H +L
Sbjct: 612 EDERVAERSVSHSEEL 627
>At3g22260 unknown protein
Length = 240
Score = 26.9 bits (58), Expect = 3.8
Identities = 20/75 (26%), Positives = 39/75 (51%), Gaps = 6/75 (8%)
Query: 33 RGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKE---KGKSNSGKGEHITSR 89
R + +Q+F +++ K K +K +++ EY+ ++ + K K + G+H+T
Sbjct: 115 RALADQLFRNADYHKHVRKHVVKQLKQQRKLYEEYVPMKYRHYTRKMKKHGEWGDHVT-- 172
Query: 90 PLQAAHDEYEEEATL 104
LQAA D +E + L
Sbjct: 173 -LQAAADRFEAKICL 186
>At3g07070 putative protein kinase
Length = 414
Score = 26.9 bits (58), Expect = 3.8
Identities = 19/85 (22%), Positives = 37/85 (43%), Gaps = 3/85 (3%)
Query: 30 RVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSR 89
+VPR +N + +V +DN K H + + V E ++K +N+ + + R
Sbjct: 14 KVPRDSDNSYRRNG---EVTGRDNNKTHPENPKTVNEQNKNNDEDKEVTNNIAAQTFSFR 70
Query: 90 PLQAAHDEYEEEATLAKQGCPHIAK 114
L A + +E + + G + K
Sbjct: 71 ELATATKNFRQECLIGEGGFGRVYK 95
>At5g19400 putative protein
Length = 1092
Score = 26.6 bits (57), Expect = 4.9
Identities = 11/33 (33%), Positives = 19/33 (57%)
Query: 51 KDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKG 83
+DN+ D+ R+ YE + + K+ K +GKG
Sbjct: 254 RDNLIVAFDKNRQSYEKLFVPSKDSSKRLTGKG 286
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,530,844
Number of Sequences: 26719
Number of extensions: 153858
Number of successful extensions: 337
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 316
Number of HSP's gapped (non-prelim): 35
length of query: 145
length of database: 11,318,596
effective HSP length: 90
effective length of query: 55
effective length of database: 8,913,886
effective search space: 490263730
effective search space used: 490263730
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146817.2


