

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146794.6 + phase: 0
(1018 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
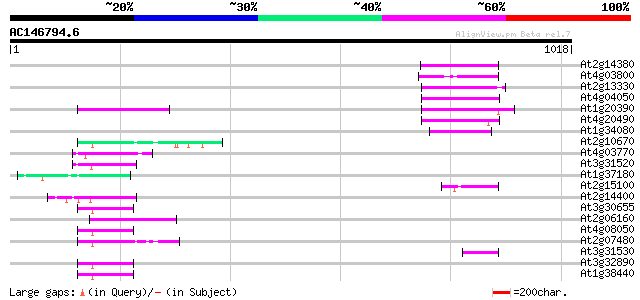
Score E
Sequences producing significant alignments: (bits) Value
At2g14380 putative retroelement pol polyprotein 81 3e-15
At4g03800 78 3e-14
At2g13330 F14O4.9 75 2e-13
At4g04050 putative transposon protein 73 7e-13
At1g20390 hypothetical protein 73 9e-13
At4g20490 putative protein 69 1e-11
At1g34080 putative protein 64 3e-10
At2g10670 pseudogene 57 7e-08
At4g03770 hypothetical protein 54 5e-07
At3g31520 hypothetical protein 50 7e-06
At1g37180 hypothetical protein 50 7e-06
At2g15100 putative retroelement pol polyprotein 49 2e-05
At2g14400 putative retroelement pol polyprotein 47 7e-05
At3g30655 hypothetical protein, 3' partial 46 9e-05
At2g06160 putative Athila retroelement ORF1 protein 45 2e-04
At4g08050 44 4e-04
At2g07480 F9A16.15 44 4e-04
At3g31530 hypothetical protein 44 5e-04
At3g32890 Athila ORF 1, putative 44 6e-04
At1g38440 hypothetical protein 44 6e-04
>At2g14380 putative retroelement pol polyprotein
Length = 764
Score = 80.9 bits (198), Expect = 3e-15
Identities = 47/142 (33%), Positives = 72/142 (50%)
Query: 746 LWFSEDELPEAGKHHNLALHISVNCKSDMLSNVLVDTGSSLNVMPKSTLDQLSYRGTPLR 805
L F+ ++L HN L I ++ ++ +L+DTGSS+NV+ K L ++ ++
Sbjct: 330 LTFTSEDLFGVDLPHNDPLVIELHIGESEVTRILIDTGSSVNVVFKDVLQKMKVHDRHIK 389
Query: 806 RSTFLVKAFDGTRKSVLGEIDLPITIGPETFLITFQVMDINASYSCLLGRPWIHDAGAVT 865
S + FDG G I LPI +G F V+D Y+ +LG PWIHD A+
Sbjct: 390 PSVRPLTGFDGNTMMTNGTIKLPIYLGGAATWHKFVVVDKPTIYNIILGTPWIHDMQAIP 449
Query: 866 STLHQKLKFAKSGKLVTIHGEE 887
S+ HQ +K S + TI G +
Sbjct: 450 SSYHQCIKIPTSIGIETIRGNQ 471
>At4g03800
Length = 637
Score = 77.8 bits (190), Expect = 3e-14
Identities = 50/144 (34%), Positives = 76/144 (52%), Gaps = 14/144 (9%)
Query: 743 CSNLWFSEDELPEAGKHHNLALHISVNCKSDMLSNVLVDTGSSLNVMPKSTLDQLSYRGT 802
CS++ F E+E + H+ AL I+++ + +S +LVDTGSS++++ + G
Sbjct: 182 CSSVSFDEEETRHIERPHDDALIITLDVANFKISRILVDTGSSVDLI---------FLGP 232
Query: 803 PLRRSTFLVKAFDGTRKSVLGEIDLPITIGPETFLITFQVMDINASYSCLLGRPWIHDAG 862
P + AF LG I LP+ G + ++ F V D A+Y+ +LG PWI
Sbjct: 233 PSP-----LVAFTSESAMSLGTIKLPVLAGKMSKIVDFVVFDKPATYNIILGTPWIFQMK 287
Query: 863 AVTSTLHQKLKFAKSGKLVTIHGE 886
AV ST HQ LKF S + TI G+
Sbjct: 288 AVPSTYHQWLKFPTSNGVETIWGD 311
>At2g13330 F14O4.9
Length = 889
Score = 75.1 bits (183), Expect = 2e-13
Identities = 49/152 (32%), Positives = 75/152 (49%), Gaps = 6/152 (3%)
Query: 748 FSEDELPEAGKHHNLALHISVNCKSDMLSNVLVDTGSSLNVMPKSTLDQLSYRGTPLRRS 807
F E E+ + K H+ AL I ++ + LS ++VDTGSS++V+ + + + L+
Sbjct: 49 FWESEITDLDKPHDDALVIRIDVGNYELSCIMVDTGSSVDVLFYDAFKRTGHLDSKLQGR 108
Query: 808 TFLVKAFDGTRKSVLGEIDLPITIGPETFLITFQVMDINASYSCLLGRPWIHDAGAVTST 867
+ F G +G I LP L F V+D A ++ +LGRPW+H AV ST
Sbjct: 109 KTPLTGFAGDTTFSIGTIQLPTIARGVRQLTNFLVVDKKAPFNAILGRPWLHVMKAVPST 168
Query: 868 LHQKLKFAKSGKLVTIHGEEAYLVSQLSSFSC 899
HQ +KF + ++G SQ SS C
Sbjct: 169 YHQCIKFPSYKGIAVVYG------SQRSSRKC 194
>At4g04050 putative transposon protein
Length = 681
Score = 73.2 bits (178), Expect = 7e-13
Identities = 43/141 (30%), Positives = 71/141 (49%)
Query: 748 FSEDELPEAGKHHNLALHISVNCKSDMLSNVLVDTGSSLNVMPKSTLDQLSYRGTPLRRS 807
F E E + K H+ AL I ++ + LS++++DTGSS++V+ ++ + + L+
Sbjct: 523 FWESETTDLNKPHDDALVIRIDVGNYELSHIMIDTGSSVDVLFYDAFKRMGHLDSELQGR 582
Query: 808 TFLVKAFDGTRKSVLGEIDLPITIGPETFLITFQVMDINASYSCLLGRPWIHDAGAVTST 867
+ F G LG I LP L +F V++ A ++ +LGRPW+H AV ST
Sbjct: 583 KTPLTGFAGDTTFSLGTIQLPTIARGVRRLTSFLVVNKKAPFNAILGRPWLHAMKAVPST 642
Query: 868 LHQKLKFAKSGKLVTIHGEEA 888
HQ +KF + + A
Sbjct: 643 YHQCIKFPSDKGIAVVSAANA 663
>At1g20390 hypothetical protein
Length = 1791
Score = 72.8 bits (177), Expect = 9e-13
Identities = 47/176 (26%), Positives = 81/176 (45%), Gaps = 7/176 (3%)
Query: 748 FSEDELPEAGKHHNLALHISVNCKSDMLSNVLVDTGSSLNVMPKSTLDQLSYRGTPLRRS 807
F++ +L HN L + + ++ VL+DTGSS++++ K L ++ ++
Sbjct: 555 FTDVDLEGLDTPHNDPLVVELIISDSRVTRVLIDTGSSVDLIFKDVLTAMNITDRQIKPV 614
Query: 808 TFLVKAFDGTRKSVLGEIDLPITIGPETFLITFQVMDINASYSCLLGRPWIHDAGAVTST 867
+ + FDG +G I LPI +G + F V+ A Y+ +LG PWIH A+ ST
Sbjct: 615 SKPLAGFDGDFVMTIGTIKLPIFVGGLIAWVKFVVIGKPAVYNVILGTPWIHQMQAIPST 674
Query: 868 LHQKLKFAKSGKLVTIHG-------EEAYLVSQLSSFSCIEAGSAEGTAFQGLTVE 916
HQ +KF + T+ +Y S+L + ++ T G+ E
Sbjct: 675 YHQCVKFPTHNGIFTLRAPKEAKTPSRSYEESELCRTEMVNIDESDPTRCVGVGAE 730
Score = 46.6 bits (109), Expect = 7e-05
Identities = 37/170 (21%), Positives = 71/170 (41%), Gaps = 4/170 (2%)
Query: 124 DRYNGLTCPQNHIIKY---VRKMGNYKDN-DSLMIHCFQDSLMEDATEWYTSLSKNDIHT 179
+ YNG P+ + + + + DN D+ F + L A W++ L N I +
Sbjct: 175 ESYNGRNDPKEFLTSFNVAINRAELTIDNFDAGRCQIFIEHLTGPAHNWFSRLKPNSIDS 234
Query: 180 FDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAARITPALDEEEMT 239
F +L ++F HY + + L S+SQ +ES R + R++ IT + +
Sbjct: 235 FHQLTSSFLKHYAPLIENQTSNADLWSISQGAKESLRSFVDRFKLVVTNITVPDEAAIVA 294
Query: 240 QTFLKTLKKDYVERMIIAAPNNFSEMVTMGTRLEEAVREGIIVFEKAESS 289
+ + + + AP+ + + +R E E +I+ K S+
Sbjct: 295 LRNAVWYDSRFRDDITLHAPSTLEDALHRASRFIELEEEKLILARKHNST 344
>At4g20490 putative protein
Length = 853
Score = 68.9 bits (167), Expect = 1e-11
Identities = 45/155 (29%), Positives = 72/155 (46%), Gaps = 14/155 (9%)
Query: 748 FSEDELPEAGKHHNLALHISVNCKSDMLSNVLVDTGSSLNVMPKSTLDQLSYRGTPLRRS 807
F+ ++ HN AL I + + + +L+DTGSS+N+M TL ++ + +
Sbjct: 465 FTSEDASGIQHPHNDALVIKLVMEDFDVERILIDTGSSVNLMFLKTLFKMGISESKIMPK 524
Query: 808 TFLVKAFDGTRKSVLGEIDLPITIGPETFLITFQVMDINASYSCLLGRPWIHDAGAVTST 867
+ +DG K +GEI L + +G T F V+D Y+ +LG PWI+ A+ ST
Sbjct: 525 IRPMTGYDGEAKMSIGEIKLQVQVGGITQKTKFVVIDSEPIYNAILGSPWIYSMKAIPST 584
Query: 868 --------------LHQKLKFAKSGKLVTIHGEEA 888
L + + AK G L+ I G A
Sbjct: 585 NLHTIRQPEGLTDMLRDREEAAKEGNLIAIKGRRA 619
>At1g34080 putative protein
Length = 811
Score = 64.3 bits (155), Expect = 3e-10
Identities = 37/112 (33%), Positives = 63/112 (56%)
Query: 763 ALHISVNCKSDMLSNVLVDTGSSLNVMPKSTLDQLSYRGTPLRRSTFLVKAFDGTRKSVL 822
AL IS++ + + +L+DTGSS++++ TL ++ ++ + + +F L
Sbjct: 476 ALVISLDVGNCEVQRILIDTGSSVDLIFLDTLVRMGISKKDIKGAPSPLVSFTIETSMSL 535
Query: 823 GEIDLPITIGPETFLITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKF 874
G I LP+T +I F V D A+Y+ +LG PW+++ AV ST HQ +KF
Sbjct: 536 GTITLPVTAQGVVKMIEFTVFDRPAAYNIILGTPWLYEMKAVPSTYHQCVKF 587
>At2g10670 pseudogene
Length = 929
Score = 56.6 bits (135), Expect = 7e-08
Identities = 79/318 (24%), Positives = 120/318 (36%), Gaps = 66/318 (20%)
Query: 124 DRYNGLTC--PQNHIIKYVRKMGNYKDN----DSLMIHCFQDSLMEDATEWYTSLSKNDI 177
++++GL P +H+ + R G K N D + F SL + A W +L KN I
Sbjct: 84 NKFHGLPMEDPLDHLDELERLCGLTKINGVSEDGFKLRLFPFSLGDKAHLWEKTLPKNSI 143
Query: 178 HTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAARITPALDEEE 237
T+D+ AF + + N+R R + + K+ ESF E +R++G T
Sbjct: 144 TTWDDCKMAFLAKFFSNSRTARLRNEISGFTLKQNESFCEAWERFKGYQ---TKCPHHGF 200
Query: 238 MTQTFLKTLKKDYVE--RMIIAAPNNFSEMVTMGTRLEEAVREGIIVFEK-AESSVNASK 294
+ L TL + + RM++ +N G L++ + EG + E A+S N ++
Sbjct: 201 SQASLLNTLYRGVLPKIRMLLDTASN-------GNFLKKDIEEGWELVENLAQSGGNYNE 253
Query: 295 RYGNG----------HHKK----------------------KETEVGMVSAGAGQSMATV 322
YG HH++ E E V G A +
Sbjct: 254 DYGRSIYTSSNTDDKHHREMKALNDKLDKIIQMQQKHVHFISEDEPFQVQEGENDQCAEI 313
Query: 323 A---------PINAAQMPPSYPYAPYSQHPFFPP------FYHQYPLPPGQPQVPVNAIA 367
P AA P+ P+ PY+Q F P Y Q PPG A A
Sbjct: 314 RYVHNQGPSLPSAAAAAEPTKPFVPYNQSLGFVPKQQFQGGYQQQQPPPGFTPHQQQAHA 373
Query: 368 QQMKQQLPVQQQQQHQQA 385
Q + V QQ QA
Sbjct: 374 PQNSDIMTVLQQLIQGQA 391
Score = 35.0 bits (79), Expect = 0.22
Identities = 28/102 (27%), Positives = 49/102 (47%), Gaps = 9/102 (8%)
Query: 776 SNVLVDTGSSLNVMPKSTLDQLSYRGTPLRRSTFLVKAFDGTRKSVLGEI-DLPITIG-- 832
+N L D G+S+++MP S + +L + + + D T K+ G + D+P+ I
Sbjct: 589 NNCLCDLGASVSLMPLSVVKKLGF--VHYKPCDLTLILADRTSKTPFGLLEDVPVMINGV 646
Query: 833 --PETFLITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKL 872
P F++ MD + +LGRP++ GAV K+
Sbjct: 647 EVPTDFVVL--EMDGESKDPLILGRPFLASVGAVIDVKQGKI 686
>At4g03770 hypothetical protein
Length = 464
Score = 53.9 bits (128), Expect = 5e-07
Identities = 41/148 (27%), Positives = 65/148 (43%), Gaps = 12/148 (8%)
Query: 115 PKKFKIPDFDRYNGLTCPQNH----IIKYVRKMGNYKDNDSLMIHCFQDSLMEDATEWYT 170
P K +I + +NG + P+ H II R ++ D+ + H F + L A W+T
Sbjct: 231 PGKLRI---EYFNGSSDPKGHLKLFIISVARAKFRPEERDAGLCHLFVEHLKGPALNWFT 287
Query: 171 SLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAARIT 230
L N + +F EL+ F Y + L SLSQ+ E R++ ++R A++
Sbjct: 288 RLKGNSVDSFQELSTLFLKQYSVLIDPSTSDADLWSLSQQPNEPLRDFLAKFRSTLAKVE 347
Query: 231 PALDEEEMTQTFLKTLKKDYVERMIIAA 258
D L TLKK R ++A
Sbjct: 348 GIND-----LAALSTLKKALCLRARLSA 370
>At3g31520 hypothetical protein
Length = 528
Score = 50.1 bits (118), Expect = 7e-06
Identities = 31/119 (26%), Positives = 56/119 (47%), Gaps = 7/119 (5%)
Query: 115 PKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYK----DNDSLMIHCFQDSLMEDATEWYT 170
P K +I + +NG + P+ H+ ++ + K + D+ + H F + L A W++
Sbjct: 239 PGKLRI---EYFNGSSDPKRHLKSFIISVARTKFRPEERDAGLCHLFVEHLKGPALCWFS 295
Query: 171 SLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAARI 229
L N + +F EL+ F Y L + L SLSQ+ E R++ + R A++
Sbjct: 296 RLEGNSMDSFQELSTLFLKQYSVLIDLGTSDADLWSLSQQPNEPLRDFLAKLRSTLAKV 354
>At1g37180 hypothetical protein
Length = 661
Score = 50.1 bits (118), Expect = 7e-06
Identities = 52/220 (23%), Positives = 89/220 (39%), Gaps = 25/220 (11%)
Query: 15 PSTAAGTSGTANNGPIPATNVVSINTTLPQTTAAVTEPLVHAI----------PQSVN-- 62
PS GT+ T N P +++ +T A PL AI P +VN
Sbjct: 205 PSVEVGTTSTQN--PTTSSSPAGFAAVSNETIARTLVPLDPAIRLPISDPRQEPSAVNSE 262
Query: 63 INTHHGNIPVIKTMEERMEELAKELRHEIKANQGNADSFKTQDLCLVPKVDVPKKFKIPD 122
+ + I + + A E+ I+A + + + L ++ + FK+P
Sbjct: 263 LQSLQDQIWAMNAKVHQATTSAPEVEKVIEATRRTPFTPRISKL----RIREFRDFKLPV 318
Query: 123 FDRYNGLTCPQNHIIKY---VRKMG-NYKDNDSLMIHCFQDSLMEDATEWYTSLSKNDIH 178
YNG P H+ + VR++ + D+ + F ++L A W+T L + I
Sbjct: 319 ---YNGKGDPNEHLTSFQVIVRRVPLEPYEEDAGLCKLFSENLSGPALTWFTQLEEGSID 375
Query: 179 TFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREY 218
F +L+ AF YG+ + L +LSQ ++ R Y
Sbjct: 376 NFKQLSTAFIKQYGYFIKSDITEAHLWNLSQSADDPLRTY 415
>At2g15100 putative retroelement pol polyprotein
Length = 1329
Score = 48.5 bits (114), Expect = 2e-05
Identities = 34/110 (30%), Positives = 53/110 (47%), Gaps = 12/110 (10%)
Query: 784 SSLNVMPKSTLDQLSYRGTPLRR------STFLVKAFDGTRKSVLGEIDLPITIGPETFL 837
S+ N PK+ + + T LR+ +T L + LG + LP+T +
Sbjct: 314 SASNAAPKAPQPATTKKPTELRKPAAGTCTTMLCET-----SMSLGTVVLPVTAQGVVKM 368
Query: 838 ITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKF-AKSGKLVTIHGE 886
+ F V D A+Y+ +LG PW+++ V ST HQ +KF GK+ I E
Sbjct: 369 VEFTVFDRPAAYNVILGTPWLYEMKVVPSTYHQCVKFPTPVGKMTGISTE 418
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 46.6 bits (109), Expect = 7e-05
Identities = 42/176 (23%), Positives = 75/176 (41%), Gaps = 18/176 (10%)
Query: 69 NIPVIKTMEERMEELAKELRHE-IKANQGNADSF----KTQDLCLVPKVDVPKKFKIPD- 122
N ++TM+E EL KE+R + ++A D K + P++ +I D
Sbjct: 102 NFDQLETMKEVSREL-KEMRSKFLQATSSEPDINRVIEKARQTPFTPQIT---SLRIRDS 157
Query: 123 ----FDRYNGLTCPQNHIIKYVRKMG----NYKDNDSLMIHCFQDSLMEDATEWYTSLSK 174
+ YNGL P+ ++ ++ G N D D F ++L A W+T L
Sbjct: 158 RKLNLESYNGLEDPKGYLAAFLIAAGRVDLNEADEDVRYCKLFSENLCGQALMWFTQLEP 217
Query: 175 NDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAARIT 230
I F+EL+ F Y + L +LSQ E+ R + +++ ++++
Sbjct: 218 GSISNFNELSVVFLKQYSILMDKSISDTDLWNLSQGPNETLRAFITKFKYVLSKLS 273
Score = 38.5 bits (88), Expect = 0.020
Identities = 24/89 (26%), Positives = 49/89 (54%), Gaps = 4/89 (4%)
Query: 744 SNLWFSEDELPEAGKHHNLALHISVNCKSDMLSNVLVDTGSSLNVMPKSTLDQLSYRGTP 803
+++ F E E + H+ AL ++++ + +S +L+DTGSS++++ STL+++
Sbjct: 407 TSILFDEKETQHLERSHDDALVVTLDVANFEVSRILIDTGSSVDLIFLSTLERMGISRAD 466
Query: 804 LRRSTFLVKAFDGTRKSVLGEIDLPITIG 832
+ R V+ K+V E+D + IG
Sbjct: 467 VNRRKLGVE----RAKAVNDEVDKLLKIG 491
>At3g30655 hypothetical protein, 3' partial
Length = 660
Score = 46.2 bits (108), Expect = 9e-05
Identities = 31/107 (28%), Positives = 54/107 (49%), Gaps = 6/107 (5%)
Query: 124 DRYNGLTC--PQNHIIKYVRKMGNYKDN----DSLMIHCFQDSLMEDATEWYTSLSKNDI 177
++++GL P +H+ ++ R G K N D + F SL + A W +L +N I
Sbjct: 52 NKFHGLPMEDPLDHLDEFERLYGLTKINGVSEDGFKLRLFPFSLGDKAHLWEKTLPQNSI 111
Query: 178 HTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRG 224
+D+ AF + + N+R R + +QK+ ESF E +R++G
Sbjct: 112 TAWDDCKKAFLAKFFSNSRTARLRNEISGFTQKQNESFGEAWERFKG 158
>At2g06160 putative Athila retroelement ORF1 protein
Length = 750
Score = 45.1 bits (105), Expect = 2e-04
Identities = 38/162 (23%), Positives = 70/162 (42%), Gaps = 3/162 (1%)
Query: 145 NYKDNDSLMIHCFQDSLMEDATEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFL 204
N D F SL + W +L +N I T+D+ AF + + N+R R +
Sbjct: 79 NGVSEDGFKFRLFPFSLGDKTHLWEKTLPQNSITTWDDCKKAFFAKFFSNSRTARLRNEI 138
Query: 205 RSLSQKKEESFREYAQRWRGAAARIT-PALDEEEMTQTFLKTLKKDYVERMIIAAPNNF- 262
+QK+ ESF E +R++G + P + + T + + + A+ NF
Sbjct: 139 SGFTQKQNESFCEAWERFKGYQTKCPHPGFKQASLLSTLYRGVLPKLRMLLDTASNGNFL 198
Query: 263 SEMVTMGTRLEEAVREGIIVF-EKAESSVNASKRYGNGHHKK 303
++ V G L E + + + E + S++ S + HH++
Sbjct: 199 NKDVEKGWELVENLAQSDGNYNENYDKSISTSSESDDKHHRE 240
>At4g08050
Length = 1428
Score = 44.3 bits (103), Expect = 4e-04
Identities = 33/107 (30%), Positives = 52/107 (47%), Gaps = 6/107 (5%)
Query: 124 DRYNGLTC--PQNHIIKYVRKMGNYKDN----DSLMIHCFQDSLMEDATEWYTSLSKNDI 177
++++GL P +H+ ++ R K N D + F SL + A W +LS + I
Sbjct: 52 NKFHGLPMEDPLDHLDEFDRLCNLTKINGVSEDGFKLRLFPFSLGDKAHIWEKNLSHDSI 111
Query: 178 HTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRG 224
T+D+ AF S + N R R + SQK ESF E +R++G
Sbjct: 112 TTWDDYKKAFLSKFFSNARTARLRNEIYGFSQKTGESFCEAWERFKG 158
Score = 37.7 bits (86), Expect = 0.033
Identities = 31/102 (30%), Positives = 48/102 (46%), Gaps = 9/102 (8%)
Query: 776 SNVLVDTGSSLNVMPKS---TLDQLSYRGTPLRRSTFLVKAFDGTRKSVLGEIDLPITIG 832
S+ L D G+S+N+MP S L+ + Y+ L L+ A +RK DLP+ I
Sbjct: 589 SDCLCDLGASVNLMPLSMARRLEFIQYKPCDLT----LILADRSSRKHFGMLKDLPVMIN 644
Query: 833 PETFLITFQVMDINASYS--CLLGRPWIHDAGAVTSTLHQKL 872
F V+D+ + +LGRP++ GAV K+
Sbjct: 645 GVEVPTDFVVLDMEVEHKDPLILGRPFLASVGAVIDVKEGKI 686
>At2g07480 F9A16.15
Length = 1012
Score = 44.3 bits (103), Expect = 4e-04
Identities = 48/191 (25%), Positives = 84/191 (43%), Gaps = 18/191 (9%)
Query: 124 DRYNGLTC--PQNHIIKYVRKMGNYKDN----DSLMIHCFQDSLMEDATEWYTSLSKNDI 177
++++GL P +H+ + R K N +S + F SL + A W +L I
Sbjct: 47 NKFHGLPMEDPLDHLDNFDRLCSLTKINGVSEESFKLRLFPFSLGDKAHLWEKTLPVKSI 106
Query: 178 HTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAARITPALDEEE 237
T+D+ AF + + N+R R + +QK ESF E +R++G + P +
Sbjct: 107 DTWDDCKKAFLAKFFSNSRKARLRSEISGFNQKNSESFSEAWERFKGYTTQ-CPHHESLP 165
Query: 238 MTQTFLKTLKKDYVERMIIAAPNNFSEMVTMGTRLEEAVREGIIVFEK-AESSVNASKRY 296
+ L+ L K RM++ +N G L + V EG + E A+S N ++ Y
Sbjct: 166 PQYSILRCLPK---IRMLLDTASN-------GNILNKDVAEGWELVENLAQSHRNYNEDY 215
Query: 297 GNGHHKKKETE 307
+ ++E
Sbjct: 216 DRTNRGSSDSE 226
Score = 33.1 bits (74), Expect = 0.82
Identities = 37/177 (20%), Positives = 70/177 (38%), Gaps = 17/177 (9%)
Query: 202 EFLRSLSQKKEESFREYAQRWRGAAARITPALDEEEMTQTFLKTLKKDYVERMIIAAPNN 261
E + +L+Q +Y + RG++ D E+ + +KTL D ++++++A N
Sbjct: 199 ELVENLAQSHRNYNEDYDRTNRGSS-------DSEDKHKKEIKTLN-DRIDKLVLAQQRN 250
Query: 262 FSEMVTMG-TRLEEAVREGIIVFEKAESSVNASKRYGNGHHKKKETEVGMVSAGAGQSMA 320
+ T+L++ E + + E + G ++K + S
Sbjct: 251 VYYITEEELTQLQDG--ENLTIEEVSYLQNQGGYNKGFNNYKPPHPNLSYRSNNVANPQD 308
Query: 321 TVAPINAAQMPPSYPYAPYSQ-----HPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQ 372
V P Q + P+ PY+Q F PP + Q P P + + QQ+ Q
Sbjct: 309 QVYPPQN-QPTQAKPFVPYNQGYNQKQNFGPPGFTQQPQQPSAQDSEMKTLPQQLVQ 364
>At3g31530 hypothetical protein
Length = 831
Score = 43.9 bits (102), Expect = 5e-04
Identities = 23/66 (34%), Positives = 33/66 (49%)
Query: 822 LGEIDLPITIGPETFLITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKFAKSGKLV 881
LG I LP+ T ++ F + D Y+ ++G PW++ AV ST H LKF S +
Sbjct: 3 LGTIKLPVRAKGVTKIVDFSITDQPTVYNAIIGTPWLNQFRAVASTYHLCLKFPTSDSVW 62
Query: 882 TIHGEE 887
G E
Sbjct: 63 EKLGHE 68
>At3g32890 Athila ORF 1, putative
Length = 755
Score = 43.5 bits (101), Expect = 6e-04
Identities = 32/107 (29%), Positives = 51/107 (46%), Gaps = 6/107 (5%)
Query: 124 DRYNGLTC--PQNHIIKYVRKMGNYKDN----DSLMIHCFQDSLMEDATEWYTSLSKNDI 177
++++GL P +H+ ++ R K N D + F SL + A W +L + I
Sbjct: 5 NKFHGLPMEDPLDHLDEFDRLYNLTKINGVSEDGFKLRLFPFSLGDKAHIWEKNLPHDSI 64
Query: 178 HTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRG 224
T+D+ AF S + N R R + SQK ESF E +R++G
Sbjct: 65 TTWDDCKKAFLSKFFSNARTARLRNEISGFSQKTRESFCEAWERFKG 111
Score = 33.9 bits (76), Expect = 0.48
Identities = 30/102 (29%), Positives = 46/102 (44%), Gaps = 9/102 (8%)
Query: 776 SNVLVDTGSSLNVMPKST---LDQLSYRGTPLRRSTFLVKAFDGTRKSVLGEIDLPITIG 832
S+ L D G+S+++MP S L+ + Y+ L L+ A RK DLP+ I
Sbjct: 541 SDCLCDLGASVSLMPLSVARRLEFIQYKPCDLT----LILADRSFRKPFGMLKDLPVMIN 596
Query: 833 PETFLITFQVMDINASYS--CLLGRPWIHDAGAVTSTLHQKL 872
F V+D+ + +LGRP + GAV K+
Sbjct: 597 GVEVPTDFVVLDMEVEHKDPLILGRPLLASVGAVIDVREGKI 638
>At1g38440 hypothetical protein
Length = 263
Score = 43.5 bits (101), Expect = 6e-04
Identities = 32/107 (29%), Positives = 51/107 (46%), Gaps = 6/107 (5%)
Query: 124 DRYNGLTC--PQNHIIKYVRKMGNYKDN----DSLMIHCFQDSLMEDATEWYTSLSKNDI 177
++++GL P +H+ ++ R K N D + F SL + A W +L + I
Sbjct: 5 NKFHGLPMEDPLDHLDEFDRLYNLTKINGVSEDGFKLRLFPFSLGDKAHIWEKNLPHDSI 64
Query: 178 HTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRG 224
T+D+ AF S + N R R + SQK ESF E +R++G
Sbjct: 65 TTWDDCKKAFLSKFFSNARTARLRNEISGFSQKTSESFCEAWERFKG 111
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,751,042
Number of Sequences: 26719
Number of extensions: 1092295
Number of successful extensions: 4206
Number of sequences better than 10.0: 138
Number of HSP's better than 10.0 without gapping: 47
Number of HSP's successfully gapped in prelim test: 93
Number of HSP's that attempted gapping in prelim test: 3782
Number of HSP's gapped (non-prelim): 356
length of query: 1018
length of database: 11,318,596
effective HSP length: 109
effective length of query: 909
effective length of database: 8,406,225
effective search space: 7641258525
effective search space used: 7641258525
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146794.6


