

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146792.3 + phase: 0
(518 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
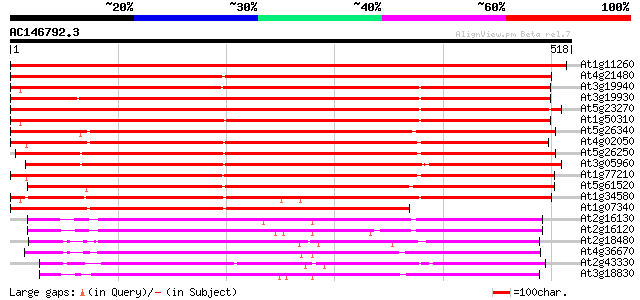
Score E
Sequences producing significant alignments: (bits) Value
At1g11260 glucose transporter 863 0.0
At4g21480 glucose transporter 801 0.0
At3g19940 putative monosaccharide transport protein, STP4 661 0.0
At3g19930 monosaccharide transport protein, STP4 656 0.0
At5g23270 monosaccharide transporter 655 0.0
At1g50310 hexose transporter, putative 653 0.0
At5g26340 hexose transporter - like protein 614 e-176
At4g02050 putative hexose transporter 563 e-161
At5g26250 hexose transporter - like protein 536 e-152
At3g05960 putative hexose transporter 532 e-151
At1g77210 unknown protein 530 e-150
At5g61520 monosaccharide transporter STP3 525 e-149
At1g34580 monosaccharide transporter like protein 516 e-146
At1g07340 hexose transporter like protein 376 e-104
At2g16130 putative sugar transporter 226 2e-59
At2g16120 putative sugar transporter 224 7e-59
At2g18480 putative sugar transporter 214 1e-55
At4g36670 sugar transporter like protein 210 1e-54
At2g43330 membrane transporter like protein 209 3e-54
At3g18830 sugar transporter like protein 206 2e-53
>At1g11260 glucose transporter
Length = 522
Score = 863 bits (2230), Expect = 0.0
Identities = 416/514 (80%), Positives = 460/514 (88%)
Query: 1 MAGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPA 60
M GG +G G K YPG LTPFV TC+VAAMGGLIFGYDIGISGGVTSM FLK+FFP+
Sbjct: 1 MPAGGFVVGDGQKAYPGKLTPFVLFTCVVAAMGGLIFGYDIGISGGVTSMPSFLKRFFPS 60
Query: 61 VYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLF 120
VYRK+ +D STNQYCQYDS TLTMFTSSLYLAAL+SSLVAST+TR+FGR+LSMLFGG+LF
Sbjct: 61 VYRKQQEDASTNQYCQYDSPTLTMFTSSLYLAALISSLVASTVTRKFGRRLSMLFGGILF 120
Query: 121 LVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITI 180
GALINGFA HVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITI
Sbjct: 121 CAGALINGFAKHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITI 180
Query: 181 GILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKA 240
GILVA VLNYFFAKIKGGWGWRLSLGGA+VPALIITIGSLVLPDTPNSMIERG + AK
Sbjct: 181 GILVAEVLNYFFAKIKGGWGWRLSLGGAVVPALIITIGSLVLPDTPNSMIERGQHEEAKT 240
Query: 241 QLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGIN 300
+L+RIRG++DV +EF+DLVAAS+ S +E+PWRNLL+RKYRP LTMAV+IPFFQQ TGIN
Sbjct: 241 KLRRIRGVDDVSQEFDDLVAASKESQSIEHPWRNLLRRKYRPHLTMAVMIPFFQQLTGIN 300
Query: 301 VIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLI 360
VIMFYAPVLFN+IGF DASLMSAV+TG VNV AT VSIYGVD+WGRR LFLEGG QMLI
Sbjct: 301 VIMFYAPVLFNTIGFTTDASLMSAVVTGSVNVAATLVSIYGVDRWGRRFLFLEGGTQMLI 360
Query: 361 CQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLE 420
CQ VAA IGAKFG G PG LP+WYAIVVV FICIYVAGFAWSWGPLGWLVPSEIFPLE
Sbjct: 361 CQAVVAACIGAKFGVDGTPGELPKWYAIVVVTFICIYVAGFAWSWGPLGWLVPSEIFPLE 420
Query: 421 IRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGI 480
IRSAAQS+ VSVNM+FTF++AQ+FL MLCH+KFGLFL FAFFV+VMSI+V+ LPETKGI
Sbjct: 421 IRSAAQSITVSVNMIFTFIIAQIFLTMLCHLKFGLFLVFAFFVVVMSIFVYIFLPETKGI 480
Query: 481 PIEEMDRVWKSHPFWSRFVEHGDHGNGVEMGKGA 514
PIEEM +VW+SH +WSRFVE G++GN +EMGK +
Sbjct: 481 PIEEMGQVWRSHWYWSRFVEDGEYGNALEMGKNS 514
>At4g21480 glucose transporter
Length = 508
Score = 801 bits (2068), Expect = 0.0
Identities = 390/500 (78%), Positives = 446/500 (89%), Gaps = 2/500 (0%)
Query: 1 MAGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPA 60
M GI IG G KEYPG LT +VT+TCIVAAMGGLIFGYDIGISGGVT+MD F +KFFP+
Sbjct: 1 MPSVGIVIGDGKKEYPGKLTLYVTVTCIVAAMGGLIFGYDIGISGGVTTMDSFQQKFFPS 60
Query: 61 VYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLF 120
VY K+ KD +NQYC++DS +LT+FTSSLYLAAL SSLVAS +TR+FGRK+SML GG+LF
Sbjct: 61 VYEKQKKDHDSNQYCRFDSVSLTLFTSSLYLAALCSSLVASYVTRQFGRKISMLLGGVLF 120
Query: 121 LVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITI 180
GAL+NGFA VWMLIVGR+LLGFGIGF NQ+VPLYLSEMAPYKYRGALNIGFQLSITI
Sbjct: 121 CAGALLNGFATAVWMLIVGRLLLGFGIGFTNQSVPLYLSEMAPYKYRGALNIGFQLSITI 180
Query: 181 GILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKA 240
GILVANVLN+FF+KI WGWRLSLGGA+VPALIIT+GSL+LPDTPNSMIERG A+A
Sbjct: 181 GILVANVLNFFFSKIS--WGWRLSLGGAVVPALIITVGSLILPDTPNSMIERGQFRLAEA 238
Query: 241 QLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGIN 300
+L++IRG++D+D+E NDL+ ASEAS VE+PWRNLLQRKYRP LTMA+LIP FQQ TGIN
Sbjct: 239 KLRKIRGVDDIDDEINDLIIASEASKLVEHPWRNLLQRKYRPHLTMAILIPAFQQLTGIN 298
Query: 301 VIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLI 360
VIMFYAPVLF +IGF DA+L+SAV+TG+VNV AT VSIYGVDKWGRR LFLEGG QMLI
Sbjct: 299 VIMFYAPVLFQTIGFGSDAALISAVVTGLVNVGATVVSIYGVDKWGRRFLFLEGGFQMLI 358
Query: 361 CQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLE 420
QVAVAAAIGAKFG G PG LP+WYAIVVVLFICIYVA FAWSWGPLGWLVPSEIFPLE
Sbjct: 359 SQVAVAAAIGAKFGVDGTPGVLPKWYAIVVVLFICIYVAAFAWSWGPLGWLVPSEIFPLE 418
Query: 421 IRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGI 480
IRSAAQS+ VSVNM+FTFL+AQVFL+MLCH+KFGLF+FFAFFV+VMSI+V+ LPET+G+
Sbjct: 419 IRSAAQSITVSVNMIFTFLIAQVFLMMLCHLKFGLFIFFAFFVVVMSIFVYLFLPETRGV 478
Query: 481 PIEEMDRVWKSHPFWSRFVE 500
PIEEM+RVW+SH +WS+FV+
Sbjct: 479 PIEEMNRVWRSHWYWSKFVD 498
>At3g19940 putative monosaccharide transport protein, STP4
Length = 514
Score = 661 bits (1705), Expect = 0.0
Identities = 324/501 (64%), Positives = 397/501 (78%), Gaps = 4/501 (0%)
Query: 1 MAGGGIPI--GGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFF 58
MAGG GGG + Y G +T FV +TCIVAAMGGL+FGYD+GISGGVTSM+ FL KFF
Sbjct: 1 MAGGAFVSEGGGGGRSYEGGVTAFVIMTCIVAAMGGLLFGYDLGISGGVTSMEEFLTKFF 60
Query: 59 PAVYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGL 118
P V + K K YC++D+Q L +FTSSLYLAAL++S +AS ITR+ GRK+SM GGL
Sbjct: 61 PQVESQMKKAKHDTAYCKFDNQMLQLFTSSLYLAALVASFMASVITRKHGRKVSMFIGGL 120
Query: 119 LFLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSI 178
FL+GAL N FA +V MLI+GR+LLG G+GFANQ+ P+YLSEMAP K RGALNIGFQ++I
Sbjct: 121 AFLIGALFNAFAVNVSMLIIGRLLLGVGVGFANQSTPVYLSEMAPAKIRGALNIGFQMAI 180
Query: 179 TIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGA 238
TIGILVAN++NY +K+ GWR+SLG A VPA+++ IGS +LPDTPNSM+ERG + A
Sbjct: 181 TIGILVANLINYGTSKM-AQHGWRVSLGLAAVPAVVMVIGSFILPDTPNSMLERGKNEEA 239
Query: 239 KAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTG 298
K LK+IRG ++VD EF DL+ A EA+ +VENPW+N+++ KYRP L IPFFQQ TG
Sbjct: 240 KQMLKKIRGADNVDHEFQDLIDAVEAAKKVENPWKNIMESKYRPALIFCSAIPFFQQITG 299
Query: 299 INVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQM 358
INVIMFYAPVLF ++GF DDA+LMSAVITGVVN+++T VSIY VD++GRR LFLEGG QM
Sbjct: 300 INVIMFYAPVLFKTLGFGDDAALMSAVITGVVNMLSTFVSIYAVDRYGRRLLFLEGGIQM 359
Query: 359 LICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFP 418
ICQ+ V + IGA+FGTSG G L A ++ FIC+YVAGFAWSWGPLGWLVPSEI P
Sbjct: 360 FICQLLVGSFIGARFGTSGT-GTLTPATADWILAFICVYVAGFAWSWGPLGWLVPSEICP 418
Query: 419 LEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETK 478
LEIR A Q++NVSVNM FTFL+ Q FL MLCHMKFGLF FFA V +M+++++FLLPETK
Sbjct: 419 LEIRPAGQAINVSVNMFFTFLIGQFFLTMLCHMKFGLFYFFASMVAIMTVFIYFLLPETK 478
Query: 479 GIPIEEMDRVWKSHPFWSRFV 499
G+PIEEM RVWK H FW +++
Sbjct: 479 GVPIEEMGRVWKQHWFWKKYI 499
>At3g19930 monosaccharide transport protein, STP4
Length = 514
Score = 656 bits (1693), Expect = 0.0
Identities = 316/499 (63%), Positives = 399/499 (79%), Gaps = 2/499 (0%)
Query: 1 MAGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPA 60
MAGG + G + Y LTP V +TC + A GGLIFGYD+GISGGVTSM+PFL++FFP
Sbjct: 1 MAGGFVSQTPGVRNYNYKLTPKVFVTCFIGAFGGLIFGYDLGISGGVTSMEPFLEEFFPY 60
Query: 61 VYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLF 120
VY KK K N+YC++DSQ LT+FTSSLY+AAL+SSL ASTITR FGRK SM GG F
Sbjct: 61 VY-KKMKSAHENEYCRFDSQLLTLFTSSLYVAALVSSLFASTITRVFGRKWSMFLGGFTF 119
Query: 121 LVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITI 180
+G+ NGFA ++ ML++GRILLGFG+GFANQ+VP+YLSEMAP RGA N GFQ++I
Sbjct: 120 FIGSAFNGFAQNIAMLLIGRILLGFGVGFANQSVPVYLSEMAPPNLRGAFNNGFQVAIIF 179
Query: 181 GILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKA 240
GI+VA ++NYF A++KG GWR+SLG A VPA++I IG+L+LPDTPNS+IERG + AK
Sbjct: 180 GIVVATIINYFTAQMKGNIGWRISLGLACVPAVMIMIGALILPDTPNSLIERGYTEEAKE 239
Query: 241 QLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGIN 300
L+ IRG +VDEEF DL+ ASE S QV++PW+N++ +YRPQL M IPFFQQ TGIN
Sbjct: 240 MLQSIRGTNEVDEEFQDLIDASEESKQVKHPWKNIMLPRYRPQLIMTCFIPFFQQLTGIN 299
Query: 301 VIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLI 360
VI FYAPVLF ++GF ASL+SA++TG++ ++ T VS++ VD++GRR LFL+GG QML+
Sbjct: 300 VITFYAPVLFQTLGFGSKASLLSAMVTGIIELLCTFVSVFTVDRFGRRILFLQGGIQMLV 359
Query: 361 CQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLE 420
Q+A+ A IG KFG +G GN+ + A ++V ICIYVAGFAWSWGPLGWLVPSEI PLE
Sbjct: 360 SQIAIGAMIGVKFGVAGT-GNIGKSDANLIVALICIYVAGFAWSWGPLGWLVPSEISPLE 418
Query: 421 IRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGI 480
IRSAAQ++NVSVNM FTFLVAQ+FL MLCHMKFGLF FFAFFV++M+I+++ +LPETK +
Sbjct: 419 IRSAAQAINVSVNMFFTFLVAQLFLTMLCHMKFGLFFFFAFFVVIMTIFIYLMLPETKNV 478
Query: 481 PIEEMDRVWKSHPFWSRFV 499
PIEEM+RVWK+H FW +F+
Sbjct: 479 PIEEMNRVWKAHWFWGKFI 497
>At5g23270 monosaccharide transporter
Length = 514
Score = 655 bits (1690), Expect = 0.0
Identities = 326/511 (63%), Positives = 406/511 (78%), Gaps = 4/511 (0%)
Query: 1 MAGGG-IPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFP 59
MAGG I G +Y G +T FV ITCIVAAMGGL+FGYDIGISGGV SM+ FL KFFP
Sbjct: 1 MAGGAFIDESGHGGDYEGRVTAFVMITCIVAAMGGLLFGYDIGISGGVISMEDFLTKFFP 60
Query: 60 AVYRK-KNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGL 118
V R+ +NK +YC+YD++ LT+FTSSLYLAAL +S +ASTITR FGRK+SM+ G L
Sbjct: 61 DVLRQMQNKRGRETEYCKYDNELLTLFTSSLYLAALFASFLASTITRLFGRKVSMVIGSL 120
Query: 119 LFLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSI 178
FL GAL+NG A ++ MLI+GR+ LG G+GFANQ+VPLYLSEMAP K RGALNIGFQL+I
Sbjct: 121 AFLSGALLNGLAINLEMLIIGRLFLGVGVGFANQSVPLYLSEMAPAKIRGALNIGFQLAI 180
Query: 179 TIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGA 238
TIGIL AN++NY K++ G GWRLSLG A VPA+++ +G LPDTPNS++ERG+++ A
Sbjct: 181 TIGILAANIVNYVTPKLQNGIGWRLSLGLAGVPAVMMLVGCFFLPDTPNSILERGNKEKA 240
Query: 239 KAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTG 298
K L++IRG +V+ EFN+L A EA+ +V++PW N++Q +YRPQLT IPFFQQ TG
Sbjct: 241 KEMLQKIRGTMEVEHEFNELCNACEAAKKVKHPWTNIMQARYRPQLTFCTFIPFFQQLTG 300
Query: 299 INVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQM 358
INVIMFYAPVLF +IGF +DASL+SAVITG+VNV++T VSIY VDK+GRRALFL+GG QM
Sbjct: 301 INVIMFYAPVLFKTIGFGNDASLISAVITGLVNVLSTIVSIYSVDKFGRRALFLQGGFQM 360
Query: 359 LICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFP 418
++ Q+AV + IG KFG +G GNL A +++ IC+YVAGFAWSWGPLGWLVPSEI P
Sbjct: 361 IVTQIAVGSMIGWKFGFNGE-GNLSGVDADIILALICLYVAGFAWSWGPLGWLVPSEICP 419
Query: 419 LEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETK 478
LEIRSA QS+NVSVNM FTF + Q FL MLCHMKFGLF FFA VL+M+I+++FLLPETK
Sbjct: 420 LEIRSAGQSLNVSVNMFFTFFIGQFFLTMLCHMKFGLFYFFAGMVLIMTIFIYFLLPETK 479
Query: 479 GIPIEEMDRVWKSHPFWSRFVEHGDHGNGVE 509
G+PIEEM +VWK H +W ++ + D G+ V+
Sbjct: 480 GVPIEEMGKVWKEHRYWGKY-SNNDDGDDVD 509
>At1g50310 hexose transporter, putative
Length = 517
Score = 653 bits (1684), Expect = 0.0
Identities = 321/502 (63%), Positives = 396/502 (77%), Gaps = 5/502 (0%)
Query: 1 MAGGGIPI--GGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFF 58
MAGG GGG Y G +T FV +TCIVAAMGGL+FGYD+GISGGVTSM+ FL KFF
Sbjct: 1 MAGGAFVSEGGGGGNSYEGGVTVFVIMTCIVAAMGGLLFGYDLGISGGVTSMEEFLSKFF 60
Query: 59 PAVYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGL 118
P V ++ ++ + YC++D+Q L +FTSSLYLAAL SS VAS +TR++GRK+SM GG+
Sbjct: 61 PEVDKQMHEARRETAYCKFDNQLLQLFTSSLYLAALASSFVASAVTRKYGRKISMFVGGV 120
Query: 119 LFLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSI 178
FL+G+L N FA +V MLIVGR+LLG G+GFANQ+ P+YLSEMAP K RGALNIGFQ++I
Sbjct: 121 AFLIGSLFNAFATNVAMLIVGRLLLGVGVGFANQSTPVYLSEMAPAKIRGALNIGFQMAI 180
Query: 179 TIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGA 238
TIGIL+AN++NY +++ GWR+SLG A VPA+I+ IGS VLPDTPNSM+ERG + A
Sbjct: 181 TIGILIANLINYGTSQMAKN-GWRVSLGLAAVPAVIMVIGSFVLPDTPNSMLERGKYEQA 239
Query: 239 KAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQR-KYRPQLTMAVLIPFFQQFT 297
+ L++IRG ++VDEEF DL A EA+ +V+NPW+N+ Q+ KYRP L IPFFQQ T
Sbjct: 240 REMLQKIRGADNVDEEFQDLCDACEAAKKVDNPWKNIFQQAKYRPALVFCSAIPFFQQIT 299
Query: 298 GINVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQ 357
GINVIMFYAPVLF ++GF DDASL+SAVITG VNVV+T VSIY VD++GRR LFLEGG Q
Sbjct: 300 GINVIMFYAPVLFKTLGFADDASLISAVITGAVNVVSTLVSIYAVDRYGRRILFLEGGIQ 359
Query: 358 MLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIF 417
M++ Q+ V IG KFGT+G+ G L A ++ FIC+YVAGFAWSWGPLGWLVPSEI
Sbjct: 360 MIVSQIVVGTLIGMKFGTTGS-GTLTPATADWILAFICLYVAGFAWSWGPLGWLVPSEIC 418
Query: 418 PLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPET 477
PLEIR A Q++NVSVNM FTFL+ Q FL MLCHMKFGLF FF V VM+++++FLLPET
Sbjct: 419 PLEIRPAGQAINVSVNMFFTFLIGQFFLTMLCHMKFGLFYFFGGMVAVMTVFIYFLLPET 478
Query: 478 KGIPIEEMDRVWKSHPFWSRFV 499
KG+PIEEM RVWK HPFW R++
Sbjct: 479 KGVPIEEMGRVWKQHPFWKRYM 500
>At5g26340 hexose transporter - like protein
Length = 526
Score = 614 bits (1583), Expect = e-176
Identities = 306/508 (60%), Positives = 386/508 (75%), Gaps = 7/508 (1%)
Query: 1 MAGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPA 60
M GGG E+ +TP V I+CI+AA GGL+FGYD+G+SGGVTSM FL+KFFP
Sbjct: 1 MTGGGFATSANGVEFEAKITPIVIISCIMAATGGLMFGYDVGVSGGVTSMPDFLEKFFPV 60
Query: 61 VYRK--KNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGL 118
VYRK DK +N YC+YD+Q L +FTSSLYLA L ++ AS TR GR+L+ML G+
Sbjct: 61 VYRKVVAGADKDSN-YCKYDNQGLQLFTSSLYLAGLTATFFASYTTRTLGRRLTMLIAGV 119
Query: 119 LFLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSI 178
F++G +N A + MLI GRILLG G+GFANQAVPL+LSE+AP + RG LNI FQL++
Sbjct: 120 FFIIGVALNAGAQDLAMLIAGRILLGCGVGFANQAVPLFLSEIAPTRIRGGLNILFQLNV 179
Query: 179 TIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGA 238
TIGIL AN++NY AKIKGGWGWRLSLG A +PAL++T+G+L++ +TPNS++ERG D
Sbjct: 180 TIGILFANLVNYGTAKIKGGWGWRLSLGLAGIPALLLTVGALLVTETPNSLVERGRLDEG 239
Query: 239 KAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTG 298
KA L+RIRG ++V+ EF DL+ AS + +V++P+RNLLQR+ RPQL +AV + FQQ TG
Sbjct: 240 KAVLRRIRGTDNVEPEFADLLEASRLAKEVKHPFRNLLQRRNRPQLVIAVALQIFQQCTG 299
Query: 299 INVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQM 358
IN IMFYAPVLF+++GF DASL SAV+TG VNV++T VSIY VDK GRR L LE G QM
Sbjct: 300 INAIMFYAPVLFSTLGFGSDASLYSAVVTGAVNVLSTLVSIYSVDKVGRRVLLLEAGVQM 359
Query: 359 LICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFP 418
QV +A +G K + NL + +AI+VV+ IC YVA FAWSWGPLGWL+PSE FP
Sbjct: 360 FFSQVVIAIILGVK--VTDTSTNLSKGFAILVVVMICTYVAAFAWSWGPLGWLIPSETFP 417
Query: 419 LEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETK 478
LE RSA QSV V VN+LFTF++AQ FL MLCH KFG+F+FF+ +VL+MS++V FLLPETK
Sbjct: 418 LETRSAGQSVTVCVNLLFTFIIAQAFLSMLCHFKFGIFIFFSAWVLIMSVFVMFLLPETK 477
Query: 479 GIPIEEM-DRVWKSHPFWSRFV-EHGDH 504
IPIEEM +RVWK H FW+RF+ +H DH
Sbjct: 478 NIPIEEMTERVWKKHWFWARFMDDHNDH 505
>At4g02050 putative hexose transporter
Length = 513
Score = 563 bits (1452), Expect = e-161
Identities = 282/501 (56%), Positives = 373/501 (74%), Gaps = 9/501 (1%)
Query: 1 MAGGGIPIGGGNKE----YPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKK 56
MAGG G KE Y G +T +V I C+VAA+GG IFGYDIGISGGVTSMD FL++
Sbjct: 1 MAGGSFGPTGVAKERAEQYQGKVTSYVIIACLVAAIGGSIFGYDIGISGGVTSMDEFLEE 60
Query: 57 FFPAVYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFG 116
FF VY KK + +N YC+YD+Q L FTSSLYLA L+S+LVAS ITR +GR+ S++ G
Sbjct: 61 FFHTVYEKKKQAHESN-YCKYDNQGLAAFTSSLYLAGLVSTLVASPITRNYGRRASIVCG 119
Query: 117 GLLFLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQL 176
G+ FL+G+ +N A ++ ML+ GRI+LG GIGF NQAVPLYLSE+AP RG LN+ FQL
Sbjct: 120 GISFLIGSGLNAGAVNLAMLLAGRIMLGVGIGFGNQAVPLYLSEVAPTHLRGGLNMMFQL 179
Query: 177 SITIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRD 236
+ TIGI AN++NY ++K WGWRLSLG A PAL++T+G LP+TPNS++ERG +
Sbjct: 180 ATTIGIFTANMVNYGTQQLKP-WGWRLSLGLAAFPALLMTLGGYFLPETPNSLVERGLTE 238
Query: 237 GAKAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQF 296
+ L ++RG E+V+ E D+V ASE + +++P+RN+LQ+++RPQL MA+ +P FQ
Sbjct: 239 RGRRVLVKLRGTENVNAELQDMVDASELANSIKHPFRNILQKRHRPQLVMAICMPMFQIL 298
Query: 297 TGINVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGA 356
TGIN I+FYAPVLF ++GF +ASL S+ +TG V V++T +SI VD+ GRRAL + GG
Sbjct: 299 TGINSILFYAPVLFQTMGFGGNASLYSSALTGAVLVLSTFISIGLVDRLGRRALLITGGI 358
Query: 357 QMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEI 416
QM+ICQV VA +G KFG + L + Y+++VV+FIC++V F WSWGPLGW +PSEI
Sbjct: 359 QMIICQVIVAVILGVKFGDN---QELSKGYSVIVVIFICLFVVAFGWSWGPLGWTIPSEI 415
Query: 417 FPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPE 476
FPLE RSA QS+ V+VN+LFTF++AQ FL +LC KFG+FLFFA +V VM+I+V+FLLPE
Sbjct: 416 FPLETRSAGQSITVAVNLLFTFIIAQAFLGLLCAFKFGIFLFFAGWVTVMTIFVYFLLPE 475
Query: 477 TKGIPIEEMDRVWKSHPFWSR 497
TKG+PIEEM +W H FW +
Sbjct: 476 TKGVPIEEMTLLWSKHWFWKK 496
>At5g26250 hexose transporter - like protein
Length = 507
Score = 536 bits (1380), Expect = e-152
Identities = 264/501 (52%), Positives = 360/501 (71%), Gaps = 7/501 (1%)
Query: 6 IPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKK 65
I G +K + +T +V I I+AA+GGLIFGYDIGISGGVT+MD FLK+FFP+VY +K
Sbjct: 5 ISSNGNSKSFDAKMTVYVFICVIIAAVGGLIFGYDIGISGGVTAMDDFLKEFFPSVYERK 64
Query: 66 NKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGAL 125
K N YC+YD+Q L +FTSSLYLAAL++S AS + GR+ +M + FL+G
Sbjct: 65 -KHAHENNYCKYDNQFLQLFTSSLYLAALVASFFASATCSKLGRRPTMQLASIFFLIGVG 123
Query: 126 INGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVA 185
+ A +++MLI+GRILLGFG+GF NQAVPL+LSE+AP + RG LNI FQL +TIGIL+A
Sbjct: 124 LAAGAVNIYMLIIGRILLGFGVGFGNQAVPLFLSEIAPARLRGGLNIVFQLMVTIGILIA 183
Query: 186 NVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRI 245
N++NYF + I +GWR++LGGA +PALI+ GSL++ +TP S+IER K LK+I
Sbjct: 184 NIVNYFTSSIHP-YGWRIALGGAGIPALILLFGSLLICETPTSLIERNKTKEGKETLKKI 242
Query: 246 RGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIMFY 305
RG+EDVDEE+ +V A + + QV++P+ L++ RP + +L+ FFQQFTGIN IMFY
Sbjct: 243 RGVEDVDEEYESIVHACDIARQVKDPYTKLMKPASRPPFVIGMLLQFFQQFTGINAIMFY 302
Query: 306 APVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAV 365
APVLF ++GF +DA+L+SAV+TG +NV++T V I+ VDK GRR L L+ MLICQ+ +
Sbjct: 303 APVLFQTVGFGNDAALLSAVVTGTINVLSTFVGIFLVDKTGRRFLLLQSSVHMLICQLVI 362
Query: 366 AAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAA 425
+ + G L A+VVV+F+C+YV GFAWSWGPLGWL+PSE FPLE R+
Sbjct: 363 GIILAKDLDVT---GTLARPQALVVVIFVCVYVMGFAWSWGPLGWLIPSETFPLETRTEG 419
Query: 426 QSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGIPIEEM 485
++ VS NM FTF++AQ FL MLC MK G+F FF+ +++VM ++ F +PETKG+ I++M
Sbjct: 420 FALAVSCNMFFTFVIAQAFLSMLCAMKSGIFFFFSGWIVVMGLFALFFVPETKGVSIDDM 479
Query: 486 -DRVWKSHPFWSRF-VEHGDH 504
D VWK H +W RF +E +H
Sbjct: 480 RDSVWKLHWYWKRFMLEEDEH 500
>At3g05960 putative hexose transporter
Length = 507
Score = 532 bits (1371), Expect = e-151
Identities = 261/496 (52%), Positives = 362/496 (72%), Gaps = 6/496 (1%)
Query: 15 YPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQY 74
+ +T +V I ++AA+GGLIFGYDIGISGGV++MD FLK+FFPAV+ +K K N Y
Sbjct: 13 FEAKMTVYVFICVMIAAVGGLIFGYDIGISGGVSAMDDFLKEFFPAVWERK-KHVHENNY 71
Query: 75 CQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVW 134
C+YD+Q L +FTSSLYLAAL++S VAS + GR+ +M F + FL+G + A ++
Sbjct: 72 CKYDNQFLQLFTSSLYLAALVASFVASATCSKLGRRPTMQFASIFFLIGVGLTAGAVNLV 131
Query: 135 MLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAK 194
MLI+GR+ LGFG+GF NQAVPL+LSE+AP + RG LNI FQL +TIGIL+AN++NYF A
Sbjct: 132 MLIIGRLFLGFGVGFGNQAVPLFLSEIAPAQLRGGLNIVFQLMVTIGILIANIVNYFTAT 191
Query: 195 IKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRGIEDVDEE 254
+ +GWR++LGGA +PA+I+ GSL++ +TP S+IER + K L++IRG++D+++E
Sbjct: 192 VHP-YGWRIALGGAGIPAVILLFGSLLIIETPTSLIERNKNEEGKEALRKIRGVDDINDE 250
Query: 255 FNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIG 314
+ +V A + + QV++P+R LL+ RP + +L+ FQQFTGIN IMFYAPVLF ++G
Sbjct: 251 YESIVHACDIASQVKDPYRKLLKPASRPPFIIGMLLQLFQQFTGINAIMFYAPVLFQTVG 310
Query: 315 FKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAIGAKFG 374
F DA+L+SAVITG +NV+AT V IY VD+ GRR L L+ MLICQ+ + + G
Sbjct: 311 FGSDAALLSAVITGSINVLATFVGIYLVDRTGRRFLLLQSSVHMLICQLIIGIILAKDLG 370
Query: 375 TSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNM 434
+G G P+ A+VVV+F+C+YV GFAWSWGPLGWL+PSE FPLE RSA +V VS NM
Sbjct: 371 VTGTLGR-PQ--ALVVVIFVCVYVMGFAWSWGPLGWLIPSETFPLETRSAGFAVAVSCNM 427
Query: 435 LFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGIPIEEM-DRVWKSHP 493
FTF++AQ FL MLC M+ G+F FF+ +++VM ++ FF +PETKGI I++M + VWK H
Sbjct: 428 FFTFVIAQAFLSMLCGMRSGIFFFFSGWIIVMGLFAFFFIPETKGIAIDDMRESVWKPHW 487
Query: 494 FWSRFVEHGDHGNGVE 509
FW R++ D + +E
Sbjct: 488 FWKRYMLPEDDHHDIE 503
>At1g77210 unknown protein
Length = 504
Score = 530 bits (1364), Expect = e-150
Identities = 266/507 (52%), Positives = 356/507 (69%), Gaps = 8/507 (1%)
Query: 1 MAGGGIPIGGGNKE---YPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKF 57
MAGG + GG K Y +T + CIV +MGG +FGYD+G+SGGVTSMD FLK+F
Sbjct: 1 MAGGALTDEGGLKRAHLYEHRITSYFIFACIVGSMGGSLFGYDLGVSGGVTSMDDFLKEF 60
Query: 58 FPAVYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGG 117
FP +Y++K + YC+YD+Q LT+FTSSLY A L+S+ AS +TR +GR+ S+L G
Sbjct: 61 FPGIYKRKQMHLNETDYCKYDNQILTLFTSSLYFAGLISTFGASYVTRIYGRRGSILVGS 120
Query: 118 LLFLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLS 177
+ F +G +IN A ++ MLI+GRI LG GIGF NQAVPLYLSEMAP K RG +N FQL+
Sbjct: 121 VSFFLGGVINAAAKNILMLILGRIFLGIGIGFGNQAVPLYLSEMAPAKIRGTVNQLFQLT 180
Query: 178 ITIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDG 237
IGILVAN++NY +I WGWRLSLG A VPA+++ +G LVLP+TPNS++E+G +
Sbjct: 181 TCIGILVANLINYKTEQIH-PWGWRLSLGLATVPAILMFLGGLVLPETPNSLVEQGKLEK 239
Query: 238 AKAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTM-AVLIPFFQQF 296
AKA L ++RG +++ EF DLV AS+A+ V+NP+RNLL R+ RPQL + A+ +P FQQ
Sbjct: 240 AKAVLIKVRGTNNIEAEFQDLVEASDAARAVKNPFRNLLARRNRPQLVIGAIGLPAFQQL 299
Query: 297 TGINVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGA 356
TG+N I+FYAPV+F S+GF ASL+S+ IT VVA +S+Y DK+GRR L LE
Sbjct: 300 TGMNSILFYAPVMFQSLGFGGSASLISSTITNAALVVAAIMSMYSADKFGRRFLLLEASV 359
Query: 357 QMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEI 416
+M V V + KFG LP+ +++V+ IC++V + SWGP+GWLVPSE+
Sbjct: 360 EMFCYMVVVGVTLALKFGEG---KELPKSLGLILVVLICLFVLAYGRSWGPMGWLVPSEL 416
Query: 417 FPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPE 476
FPLE RSA QSV V VN+ FT L+AQ FL+ LCH+K+G+FL FA +L M +V+FLLPE
Sbjct: 417 FPLETRSAGQSVVVCVNLFFTALIAQCFLVSLCHLKYGIFLLFAGLILGMGSFVYFLLPE 476
Query: 477 TKGIPIEEMDRVWKSHPFWSRFVEHGD 503
TK +PIEE+ +W+ H W ++VE D
Sbjct: 477 TKQVPIEEVYLLWRQHWLWKKYVEDVD 503
>At5g61520 monosaccharide transporter STP3
Length = 514
Score = 525 bits (1351), Expect = e-149
Identities = 265/493 (53%), Positives = 350/493 (70%), Gaps = 11/493 (2%)
Query: 17 GNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDK-----ST 71
G +T FV +C++AAMGG+IFGYDIG+SGGV SM PFLK+FFP VY+ + +D+ S
Sbjct: 18 GKITYFVVASCVMAAMGGVIFGYDIGVSGGVMSMGPFLKRFFPKVYKLQEEDRRRRGNSN 77
Query: 72 NQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFAN 131
N YC ++SQ LT FTSSLY++ L+++L+AS++TR +GRK S+ GG+ FL GA + G A
Sbjct: 78 NHYCLFNSQLLTSFTSSLYVSGLIATLLASSVTRSWGRKPSIFLGGVSFLAGAALGGSAQ 137
Query: 132 HVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYF 191
+V MLI+ R+LLG G+GFANQ+VPLYLSEMAP KYRGA++ GFQL I IG L ANV+NY
Sbjct: 138 NVAMLIIARLLLGVGVGFANQSVPLYLSEMAPAKYRGAISNGFQLCIGIGFLSANVINYE 197
Query: 192 FAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIE-RGDRDGAKAQLKRIRGIED 250
IK GWR+SL A +PA I+T+GSL LP+TPNS+I+ GD + L+R+RG D
Sbjct: 198 TQNIK--HGWRISLATAAIPASILTLGSLFLPETPNSIIQTTGDVHKTELMLRRVRGTND 255
Query: 251 VDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLF 310
V +E DLV AS S N + LLQRKYRP+L MA++IPFFQQ TGINV+ FYAPVL+
Sbjct: 256 VQDELTDLVEASSGSDTDSNAFLKLLQRKYRPELVMALVIPFFQQVTGINVVAFYAPVLY 315
Query: 311 NSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAIG 370
++GF + SLMS ++TG+V +T +S+ VD+ GR+ LFL GG QML+ QV + +
Sbjct: 316 RTVGFGESGSLMSTLVTGIVGTSSTLLSMLVVDRIGRKTLFLIGGLQMLVSQVTIGVIV- 374
Query: 371 AKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNV 430
+ G + E Y VV+ +C+YVAGF WSWGPLGWLVPSEIFPLEIRS AQSV V
Sbjct: 375 --MVADVHDGVIKEGYGYAVVVLVCVYVAGFGWSWGPLGWLVPSEIFPLEIRSVAQSVTV 432
Query: 431 SVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGIPIEEMDRVWK 490
+V+ +FTF VAQ MLC + G+F F+ +++VM++ V LPETK +PIE++ +W+
Sbjct: 433 AVSFVFTFAVAQSAPPMLCKFRAGIFFFYGGWLVVMTVAVQLFLPETKNVPIEKVVGLWE 492
Query: 491 SHPFWSRFVEHGD 503
H FW R D
Sbjct: 493 KHWFWRRMTSKRD 505
>At1g34580 monosaccharide transporter like protein
Length = 506
Score = 516 bits (1328), Expect = e-146
Identities = 263/508 (51%), Positives = 358/508 (69%), Gaps = 14/508 (2%)
Query: 1 MAGGGIPI---GGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKF 57
MAGGG+ + GN + +T V ++CIVAA GLIFGYDIGISGGVT+M PFL+KF
Sbjct: 1 MAGGGLALDVSSAGNID--AKITAAVVMSCIVAASCGLIFGYDIGISGGVTTMKPFLEKF 58
Query: 58 FPAVYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGG 117
FP+V +K ++ K TN YC YDSQ LT FTSSLY+A L++SLVAS +T +GR+ +M+ GG
Sbjct: 59 FPSVLKKASEAK-TNVYCVYDSQLLTAFTSSLYVAGLVASLVASRLTAAYGRRTTMILGG 117
Query: 118 LLFLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLS 177
FL GALING A ++ MLI GRILLGFG+GF NQA P+YLSE+AP ++RGA NIGF
Sbjct: 118 FTFLFGALINGLAANIAMLISGRILLGFGVGFTNQAAPVYLSEVAPPRWRGAFNIGFSCF 177
Query: 178 ITIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDG 237
I++G++ AN++NY + GWR+SLG A VPA I+T+G L + DTP+S++ RG D
Sbjct: 178 ISMGVVAANLINYGTDSHRN--GWRISLGLAAVPAAIMTVGCLFISDTPSSLLARGKHDE 235
Query: 238 AKAQLKRIRGIE---DVDEEFNDLVAASEASMQ--VENPWRNLLQRKYRPQLTMAVLIPF 292
A L ++RG+E DV+ E +LV +S+ +++ E + +LQR+YRP L +AV+IP
Sbjct: 236 AHTSLLKLRGVENIADVETELAELVRSSQLAIEARAELFMKTILQRRYRPHLVVAVVIPC 295
Query: 293 FQQFTGINVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFL 352
FQQ TGI V FYAPVLF S+GF +L++ I G VN+ + +S +D++GRR LF+
Sbjct: 296 FQQLTGITVNAFYAPVLFRSVGFGSGPALIATFILGFVNLGSLLLSTMVIDRFGRRFLFI 355
Query: 353 EGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLV 412
GG ML+CQ+AVA + G +G+ G + + YA+ VV+ +CIY AGF WSWGPL WLV
Sbjct: 356 AGGILMLLCQIAVAVLLAVTVGATGD-GEMKKGYAVTVVVLLCIYAAGFGWSWGPLSWLV 414
Query: 413 PSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFF 472
PSEIFPL+IR A QS++V+VN TF ++Q FL LC K+G FLF+ ++ M+I+V
Sbjct: 415 PSEIFPLKIRPAGQSLSVAVNFAATFALSQTFLATLCDFKYGAFLFYGGWIFTMTIFVIM 474
Query: 473 LLPETKGIPIEEMDRVWKSHPFWSRFVE 500
LPETKGIP++ M +VW+ H +W RF +
Sbjct: 475 FLPETKGIPVDSMYQVWEKHWYWQRFTK 502
>At1g07340 hexose transporter like protein
Length = 383
Score = 376 bits (965), Expect = e-104
Identities = 192/370 (51%), Positives = 261/370 (69%), Gaps = 4/370 (1%)
Query: 1 MAGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPA 60
MA G + + G K +P LT V + C++AA+GGL+FGYDIGISGGVTSMD FL FFP
Sbjct: 1 MAVGSMNVEEGTKAFPAKLTGQVFLCCVIAAVGGLMFGYDIGISGGVTSMDTFLLDFFPH 60
Query: 61 VYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLF 120
VY KK++ N YC++D Q L +FTSSLYLA + +S ++S ++R FGRK +++ + F
Sbjct: 61 VYEKKHRVHENN-YCKFDDQLLQLFTSSLYLAGIFASFISSYVSRAFGRKPTIMLASIFF 119
Query: 121 LVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITI 180
LVGA++N A + MLI GRILLGFGIGF NQ VPL++SE+AP +YRG LN+ FQ ITI
Sbjct: 120 LVGAILNLSAQELGMLIGGRILLGFGIGFGNQTVPLFISEIAPARYRGGLNVMFQFLITI 179
Query: 181 GILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKA 240
GIL A+ +NY + +K GWR SLGGA VPALI+ IGS + +TP S+IERG + K
Sbjct: 180 GILAASYVNYLTSTLKN--GWRYSLGGAAVPALILLIGSFFIHETPASLIERGKDEKGKQ 237
Query: 241 QLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQR-KYRPQLTMAVLIPFFQQFTGI 299
L++IRGIED++ EFN++ A+E + +V++P++ L + + RP L L+ FFQQFTGI
Sbjct: 238 VLRKIRGIEDIELEFNEIKYATEVATKVKSPFKELFTKSENRPPLVCGTLLQFFQQFTGI 297
Query: 300 NVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQML 359
NV+MFYAPVLF ++G D+ASL+S V+T VN +AT +S+ VD GRR L +EG QM
Sbjct: 298 NVVMFYAPVLFQTMGSGDNASLISTVVTNGVNAIATVISLLVVDFAGRRCLLMEGALQMT 357
Query: 360 ICQVAVAAAI 369
Q+ + +
Sbjct: 358 ATQMTIGGIL 367
>At2g16130 putative sugar transporter
Length = 511
Score = 226 bits (577), Expect = 2e-59
Identities = 149/496 (30%), Positives = 247/496 (49%), Gaps = 43/496 (8%)
Query: 17 GNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQ 76
GN + F I+A+M +I GYDIG+ G A++ K + S Q
Sbjct: 20 GNRSRFAFACAILASMTSIILGYDIGVMSGA------------AIFIKDDLKLSDVQ--- 64
Query: 77 YDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWML 136
L + L + +L+ S A + GR+ +++ G F GAL+ GFA + +
Sbjct: 65 -----LEILMGILNIYSLIGSGAAGRTSDWIGRRYTIVLAGFFFFCGALLMGFATNYPFI 119
Query: 137 IVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIK 196
+VGR + G G+G+A P+Y +E+AP RG L+ ++ I IGIL+ V NYFFAK+
Sbjct: 120 MVGRFVAGIGVGYAMMIAPVYTTEVAPASSRGFLSSFPEIFINIGILLGYVSNYFFAKLP 179
Query: 197 GGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERG--------------DRDGAKAQL 242
GWR LG VP++ + IG L +P++P ++ +G ++ A ++L
Sbjct: 180 EHIGWRFMLGIGAVPSVFLAIGVLAMPESPRWLVMQGRLGDAFKVLDKTSNTKEEAISRL 239
Query: 243 KRI-RGIEDVDEEFNDLVAASEASMQVENPWRNLLQR---KYRPQLTMAVLIPFFQQFTG 298
I R + D+ +D++ + W++LL R R L + I F QQ +G
Sbjct: 240 NDIKRAVGIPDDMTDDVIVVPNKKSAGKGVWKDLLVRPTPSVRHILIACLGIHFSQQASG 299
Query: 299 INVIMFYAPVLFNSIGFKD-DASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQ 357
I+ ++ Y+P +F+ G K + L++ V GVV + V VD++GRRAL L
Sbjct: 300 IDAVVLYSPTIFSRAGLKSKNDQLLATVAVGVVKTLFIVVGTCLVDRFGRRALLLTSMGG 359
Query: 358 MLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIF 417
M A+ ++ NPG +W + V + +VA F+ GP+ W+ SEIF
Sbjct: 360 MFFSLTALGTSLTV---IDRNPGQTLKWAIGLAVTTVMTFVATFSLGAGPVTWVYASEIF 416
Query: 418 PLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKF-GLFLFFAFFVLVMSIYVFFLLPE 476
P+ +R+ S+ V +N L + ++ FL + + G FL FA + ++ F LPE
Sbjct: 417 PVRLRAQGASLGVMLNRLMSGIIGMTFLSLSKGLTIGGAFLLFAGVAVAAWVFFFTFLPE 476
Query: 477 TKGIPIEEMDRVWKSH 492
T+G+P+EE++ ++ S+
Sbjct: 477 TRGVPLEEIESLFGSY 492
>At2g16120 putative sugar transporter
Length = 511
Score = 224 bits (572), Expect = 7e-59
Identities = 152/500 (30%), Positives = 249/500 (49%), Gaps = 51/500 (10%)
Query: 17 GNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQ 76
GN + + I+A+M +I GYDIG+ G + ++ K + S Q
Sbjct: 20 GNRSRYAFACAILASMTSIILGYDIGVMSGAS------------IFIKDDLKLSDVQ--- 64
Query: 77 YDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWML 136
L + L + +L+ S A + GR+ +++ G F GAL+ GFA + +
Sbjct: 65 -----LEILMGILNIYSLVGSGAAGRTSDWLGRRYTIVLAGAFFFCGALLMGFATNYPFI 119
Query: 137 IVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIK 196
+VGR + G G+G+A P+Y +E+AP RG L ++ I IGIL+ V NYFF+K+
Sbjct: 120 MVGRFVAGIGVGYAMMIAPVYTAEVAPASSRGFLTSFPEIFINIGILLGYVSNYFFSKLP 179
Query: 197 GGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKR--------IRGI 248
GWR LG VP++ + IG L +P++P ++ +G A L + I +
Sbjct: 180 EHLGWRFMLGVGAVPSVFLAIGVLAMPESPRWLVLQGRLGDAFKVLDKTSNTKEEAISRL 239
Query: 249 EDV-------DEEFNDLVAASEASMQVENPWRNLLQR---KYRPQLTMAVLIPFFQQFTG 298
+D+ D+ +D++ + W++LL R R L + I F QQ +G
Sbjct: 240 DDIKRAVGIPDDMTDDVIVVPNKKSAGKGVWKDLLVRPTPSVRHILIACLGIHFAQQASG 299
Query: 299 INVIMFYAPVLFNSIGFKD-DASLMSAVITGVVN----VVATCVSIYGVDKWGRRALFLE 353
I+ ++ Y+P +F+ G K + L++ V GVV VV TCV VD++GRRAL L
Sbjct: 300 IDAVVLYSPTIFSKAGLKSKNDQLLATVAVGVVKTLFIVVGTCV----VDRFGRRALLLT 355
Query: 354 GGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVP 413
M + A+ ++ + NPG +W + V + +VA F+ GP+ W+
Sbjct: 356 SMGGMFLSLTALGTSLTV---INRNPGQTLKWAIGLAVTTVMTFVATFSIGAGPVTWVYC 412
Query: 414 SEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKF-GLFLFFAFFVLVMSIYVFF 472
SEIFP+ +R+ S+ V +N L + ++ FL + + G FL FA ++ F
Sbjct: 413 SEIFPVRLRAQGASLGVMLNRLMSGIIGMTFLSLSKGLTIGGAFLLFAGVAAAAWVFFFT 472
Query: 473 LLPETKGIPIEEMDRVWKSH 492
LPET+GIP+EEM+ ++ S+
Sbjct: 473 FLPETRGIPLEEMETLFGSY 492
>At2g18480 putative sugar transporter
Length = 508
Score = 214 bits (544), Expect = 1e-55
Identities = 151/495 (30%), Positives = 249/495 (49%), Gaps = 51/495 (10%)
Query: 18 NLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQY 77
++ F IVA++ +IFGYD G+ G F++ D N
Sbjct: 17 HMNKFAFGCAIVASIISIIFGYDTGVMSGAQI---FIRD-----------DLKIN----- 57
Query: 78 DSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLI 137
D+Q + + L L AL+ SL A + GR+ ++ ++FLVG+++ G+ + +L+
Sbjct: 58 DTQ-IEVLAGILNLCALVGSLTAGKTSDVIGRRYTIALSAVIFLVGSVLMGYGPNYPVLM 116
Query: 138 VGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKG 197
VGR + G G+GFA P+Y +E++ +RG L +L I++GIL+ V NY F K+
Sbjct: 117 VGRCIAGVGVGFALMIAPVYSAEISSASHRGFLTSLPELCISLGILLGYVSNYCFGKLTL 176
Query: 198 GWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRGI-EDVDEEFN 256
GWRL LG A P+LI+ G +P++P ++ +G + AK + + E+ +E F
Sbjct: 177 KLGWRLMLGIAAFPSLILAFGITRMPESPRWLVMQGRLEEAKKIMVLVSNTEEEAEERFR 236
Query: 257 DLVAASEASM--------------QVENPWRNLLQRKYRPQ----LTMAVLIPFFQQFTG 298
D++ A+E + ++ WR L+ K RP L AV I FF+ TG
Sbjct: 237 DILTAAEVDVTEIKEVGGGVKKKNHGKSVWRELV-IKPRPAVRLILIAAVGIHFFEHATG 295
Query: 299 INVIMFYAPVLFNSIG-FKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFL--EGG 355
I ++ Y+P +F G D L++ V G+ ++ + +DK GRR L L GG
Sbjct: 296 IEAVVLYSPRIFKKAGVVSKDKLLLATVGVGLTKAFFIIIATFLLDKVGRRKLLLTSTGG 355
Query: 356 AQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSE 415
+ +AV+ + +FG W + ++ +VA F+ GP+ W+ SE
Sbjct: 356 MVFALTSLAVSLTMVQRFGRLA-------WALSLSIVSTYAFVAFFSIGLGPITWVYSSE 408
Query: 416 IFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKF-GLFLFFAFFVLVMSIYVFFLL 474
IFPL +R+ S+ V+VN + V+ FL M + G+F FA + + FF+L
Sbjct: 409 IFPLRLRAQGASIGVAVNRIMNATVSMSFLSMTKAITTGGVFFVFAGIAVAAWWFFFFML 468
Query: 475 PETKGIPIEEMDRVW 489
PETKG+P+EEM++++
Sbjct: 469 PETKGLPLEEMEKLF 483
Score = 31.6 bits (70), Expect = 1.1
Identities = 54/250 (21%), Positives = 96/250 (37%), Gaps = 60/250 (24%)
Query: 28 IVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKK---NKDKSTNQYCQYDSQTLTM 84
++AA+G F + GI V + P +++K +KDK L +
Sbjct: 281 LIAAVGIHFFEHATGIEAVVL--------YSPRIFKKAGVVSKDK------------LLL 320
Query: 85 FTSSLYLAALLSSLVASTITRRFGRKLSMLF--GGLLFLVGAL------INGFANHVWML 136
T + L ++A+ + + GR+ +L GG++F + +L + F W L
Sbjct: 321 ATVGVGLTKAFFIIIATFLLDKVGRRKLLLTSTGGMVFALTSLAVSLTMVQRFGRLAWAL 380
Query: 137 IVGRI-----LLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYF 191
+ + + F IG +Y SE+ P + R +IG+ V ++N
Sbjct: 381 SLSIVSTYAFVAFFSIGLG-PITWVYSSEIFPLRLRAQ-------GASIGVAVNRIMNAT 432
Query: 192 FAKIKGGWGWRLSLGGAMVPALIITIGS-----LVLPDTPNSMIE-----------RGDR 235
+ ++ GG I + + +LP+T +E RGDR
Sbjct: 433 VSMSFLSMTKAITTGGVFFVFAGIAVAAWWFFFFMLPETKGLPLEEMEKLFGGGGPRGDR 492
Query: 236 DGAKAQLKRI 245
DG + Q K I
Sbjct: 493 DGLEIQTKTI 502
>At4g36670 sugar transporter like protein
Length = 493
Score = 210 bits (535), Expect = 1e-54
Identities = 148/497 (29%), Positives = 240/497 (47%), Gaps = 44/497 (8%)
Query: 14 EYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQ 73
E P + F IVA++ +IFGYD G+ G F+++ D TN
Sbjct: 8 EKPAGVNRFALQCAIVASIVSIIFGYDTGVMSGAMV---FIEE-----------DLKTND 53
Query: 74 YCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHV 133
+ + T L L AL+ SL+A + GR+ +++ +LF++G+++ G+ +
Sbjct: 54 V------QIEVLTGILNLCALVGSLLAGRTSDIIGRRYTIVLASILFMLGSILMGWGPNY 107
Query: 134 WMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFA 193
+L+ GR G G+GFA P+Y +E+A +RG L L I+IGIL+ ++NYFF+
Sbjct: 108 PVLLSGRCTAGLGVGFALMVAPVYSAEIATASHRGLLASLPHLCISIGILLGYIVNYFFS 167
Query: 194 KIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRI-RGIEDVD 252
K+ GWRL LG A VP+L++ G L +P++P +I +G K L+ + E+ +
Sbjct: 168 KLPMHIGWRLMLGIAAVPSLVLAFGILKMPESPRWLIMQGRLKEGKEILELVSNSPEEAE 227
Query: 253 EEFNDLVAASEASMQV--------------ENPWRNLLQR---KYRPQLTMAVLIPFFQQ 295
F D+ AA+ + E W+ L+ R R L A+ I FFQ
Sbjct: 228 LRFQDIKAAAGIDPKCVDDVVKMEGKKTHGEGVWKELILRPTPAVRRVLLTALGIHFFQH 287
Query: 296 FTGINVIMFYAPVLFNSIGFKDDASLMSAVI-TGVVNVVATCVSIYGVDKWGRRALFLEG 354
+GI ++ Y P +F G L I G++ + +DK GRR L L
Sbjct: 288 ASGIEAVLLYGPRIFKKAGITTKDKLFLVTIGVGIMKTTFIFTATLLLDKVGRRKLLLTS 347
Query: 355 GAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPS 414
M+I +G + N G W ++ ++ +VA F+ GP+ W+ S
Sbjct: 348 VGGMVI----ALTMLGFGLTMAQNAGGKLAWALVLSIVAAYSFVAFFSIGLGPITWVYSS 403
Query: 415 EIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKF-GLFLFFAFFVLVMSIYVFFL 473
E+FPL++R+ S+ V+VN + V+ FL + + G F FA V + FFL
Sbjct: 404 EVFPLKLRAQGASLGVAVNRVMNATVSMSFLSLTSAITTGGAFFMFAGVAAVAWNFFFFL 463
Query: 474 LPETKGIPIEEMDRVWK 490
LPETKG +EE++ +++
Sbjct: 464 LPETKGKSLEEIEALFQ 480
>At2g43330 membrane transporter like protein
Length = 509
Score = 209 bits (532), Expect = 3e-54
Identities = 140/475 (29%), Positives = 239/475 (49%), Gaps = 34/475 (7%)
Query: 28 IVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQYDSQTLTMFTS 87
+ A +GGL+FGYD G+ G ++K F V + ++ S
Sbjct: 36 VTAGIGGLLFGYDTGVISGALL---YIKDDFEVVKQSSFLQET--------------IVS 78
Query: 88 SLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRILLGFGI 147
+ A++ + I +GRK + LF ++F GA++ A ++LI GR+L+G G+
Sbjct: 79 MALVGAMIGAAAGGWINDYYGRKKATLFADVVFAAGAIVMAAAPDPYVLISGRLLVGLGV 138
Query: 148 GFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWRLSLGG 207
G A+ P+Y++E +P + RG L L IT G ++ ++N F ++ G W W L + G
Sbjct: 139 GVASVTAPVYIAEASPSEVRGGLVSTNVLMITGGQFLSYLVNSAFTQVPGTWRWMLGVSG 198
Query: 208 AMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRGIEDVDEEFNDLVAASEASMQ 267
VPA+I I L +P++P + + + A L R I +++E + L AA E Q
Sbjct: 199 --VPAVIQFILMLFMPESPRWLFMKNRKAEAIQVLARTYDISRLEDEIDHLSAAEEEEKQ 256
Query: 268 VENP--WRNLLQRKYRPQLTMAVL----IPFFQQFTGINVIMFYAPVLFNSIGF-KDDAS 320
+ + ++ + K +L +A L + FQQFTGIN +M+Y+P + GF + +
Sbjct: 257 RKRTVGYLDVFRSK---ELRLAFLAGAGLQAFQQFTGINTVMYYSPTIVQMAGFHSNQLA 313
Query: 321 LMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPG 380
L ++I +N T V IY +D GR+ L L ++I + ++ + + TS + G
Sbjct: 314 LFLSLIVAAMNAAGTVVGIYFIDHCGRKKLALSSLFGVIISLLILSVSFFKQSETSSD-G 372
Query: 381 NLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLV 440
L W A VL + +Y+ FA GP+ W V SEI+P + R ++ +VN + +V
Sbjct: 373 GLYGWLA---VLGLALYIVFFAPGMGPVPWTVNSEIYPQQYRGICGGMSATVNWISNLIV 429
Query: 441 AQVFLIMLCHMKFGL-FLFFAFFVLVMSIYVFFLLPETKGIPIEEMDRVWKSHPF 494
AQ FL + G+ FL A ++ I+V +PET+G+ E++++WK +
Sbjct: 430 AQTFLTIAEAAGTGMTFLILAGIAVLAVIFVIVFVPETQGLTFSEVEQIWKERAY 484
>At3g18830 sugar transporter like protein
Length = 539
Score = 206 bits (525), Expect = 2e-53
Identities = 140/482 (29%), Positives = 234/482 (48%), Gaps = 44/482 (9%)
Query: 28 IVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQYDSQTLTMFTS 87
I+A+M ++ GYDIG+ G +Y K++ + + + +
Sbjct: 41 ILASMTSILLGYDIGVMSGAM------------IYIKRD--------LKINDLQIGILAG 80
Query: 88 SLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRILLGFGI 147
SL + +L+ S A + GR+ +++ G +F GA++ G + + L+ GR + G G+
Sbjct: 81 SLNIYSLIGSCAAGRTSDWIGRRYTIVLAGAIFFAGAILMGLSPNYAFLMFGRFIAGIGV 140
Query: 148 GFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWRLSLGG 207
G+A P+Y +E++P RG LN ++ I GI++ V N F+ + GWRL LG
Sbjct: 141 GYALMIAPVYTAEVSPASSRGFLNSFPEVFINAGIMLGYVSNLAFSNLPLKVGWRLMLGI 200
Query: 208 AMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRG--------IEDVDEE----- 254
VP++I+ IG L +P++P ++ +G AK L + +ED+
Sbjct: 201 GAVPSVILAIGVLAMPESPRWLVMQGRLGDAKRVLDKTSDSPTEATLRLEDIKHAAGIPA 260
Query: 255 --FNDLVAASEASMQVENPWRNLLQR---KYRPQLTMAVLIPFFQQFTGINVIMFYAPVL 309
+D+V S + E WR LL R R + A+ I FFQQ +GI+ ++ ++P +
Sbjct: 261 DCHDDVVQVSRRNSHGEGVWRELLIRPTPAVRRVMIAAIGIHFFQQASGIDAVVLFSPRI 320
Query: 310 FNSIGFK-DDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAA 368
F + G K D L++ V GVV V+ + +D+ GRR L L M++ AA
Sbjct: 321 FKTAGLKTDHQQLLATVAVGVVKTSFILVATFLLDRIGRRPLLLTSVGGMVLS----LAA 376
Query: 369 IGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSV 428
+G W +V + + YVA F+ GP+ W+ SEIFPL +RS S+
Sbjct: 377 LGTSLTIIDQSEKKVMWAVVVAIATVMTYVATFSIGAGPITWVYSSEIFPLRLRSQGSSM 436
Query: 429 NVSVNMLFTFLVAQVFLIMLCHMKF-GLFLFFAFFVLVMSIYVFFLLPETKGIPIEEMDR 487
V VN + + +++ FL M M G F F V ++ + LPET+G +E+MD
Sbjct: 437 GVVVNRVTSGVISISFLPMSKAMTTGGAFYLFGGIATVAWVFFYTFLPETQGRMLEDMDE 496
Query: 488 VW 489
++
Sbjct: 497 LF 498
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.143 0.444
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,397,894
Number of Sequences: 26719
Number of extensions: 485026
Number of successful extensions: 1923
Number of sequences better than 10.0: 103
Number of HSP's better than 10.0 without gapping: 68
Number of HSP's successfully gapped in prelim test: 35
Number of HSP's that attempted gapping in prelim test: 1588
Number of HSP's gapped (non-prelim): 133
length of query: 518
length of database: 11,318,596
effective HSP length: 104
effective length of query: 414
effective length of database: 8,539,820
effective search space: 3535485480
effective search space used: 3535485480
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146792.3


