

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146790.10 - phase: 0
(406 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
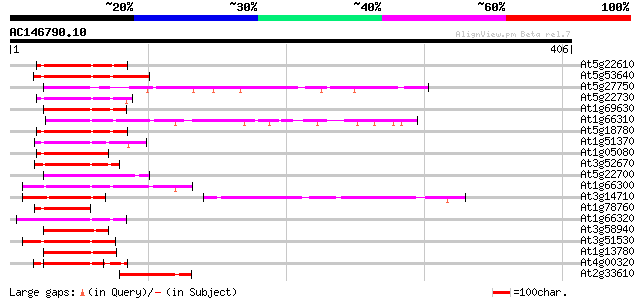
Score E
Sequences producing significant alignments: (bits) Value
At5g22610 unknown protein 58 8e-09
At5g53640 heat shock transcription factor HSF30-like protein 56 4e-08
At5g27750 unknown protein 55 7e-08
At5g22730 unknown protein 54 2e-07
At1g69630 hypothetical protein 54 2e-07
At1g66310 unknown protein 54 2e-07
At5g18780 putative protein 53 3e-07
At1g51370 unknown protein 52 6e-07
At1g05080 hypothetical protein 52 8e-07
At3g52670 putative protein 51 1e-06
At5g22700 unknown protein 50 2e-06
At1g66300 unknown protein 50 2e-06
At3g14710 unknown protein 50 2e-06
At1g78760 hypothetical protein 50 3e-06
At1g66320 hypothetical protein 50 3e-06
At3g58940 putative protein 49 4e-06
At3g51530 unknown protein 49 4e-06
At1g13780 hypothetical protein 49 4e-06
At4g00320 49 5e-06
At2g33610 putative SWI/SNF complex subunit SW13 49 5e-06
>At5g22610 unknown protein
Length = 502
Score = 58.2 bits (139), Expect = 8e-09
Identities = 33/66 (50%), Positives = 44/66 (66%), Gaps = 3/66 (4%)
Query: 20 EDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNPL 79
ED I SKLPE L++++LS+LPTKD+VRT VLSKRW + W I LDLD + + Y+
Sbjct: 17 EDLI-SKLPEVLLSQILSYLPTKDIVRTSVLSKRWKSVWL-LIPGLDLDSSEFPH-YDTF 73
Query: 80 CSYCEE 85
+ E
Sbjct: 74 VDFMNE 79
>At5g53640 heat shock transcription factor HSF30-like protein
Length = 454
Score = 55.8 bits (133), Expect = 4e-08
Identities = 34/85 (40%), Positives = 54/85 (63%), Gaps = 4/85 (4%)
Query: 18 IEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYN 77
++ED I S+ ESL+ ++L++LPTKDVV+T VLS RW + W + L+LD D S ++N
Sbjct: 19 LKEDRI-SQFYESLLCQILNYLPTKDVVKTSVLSTRWRSLWL-LVPSLELDSRDFS-DFN 75
Query: 78 PLCSYCEE-ICTDYCYDYDKYKLTV 101
S+C+ ++ +K KLT+
Sbjct: 76 TFVSFCDRYFDSNRVLCINKLKLTI 100
>At5g27750 unknown protein
Length = 459
Score = 55.1 bits (131), Expect = 7e-08
Identities = 79/299 (26%), Positives = 129/299 (42%), Gaps = 49/299 (16%)
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNPLCSYCE 84
S+LPESLI ++L LPTKD V+T VLS RW N W L++ +DL+
Sbjct: 8 SELPESLITQILLCLPTKDSVKTSVLSTRWKNLW------LNVPGLDLT----------- 50
Query: 85 EICTDYCYDYDKYK---LTVCPRCHVCGNHKANLRDAEV-YERRVEIDFES--GMKIESG 138
C+D+ ++ ++Y+ + R + N ++ L+ EV Y+RR F+ G I G
Sbjct: 51 --CSDFPFEDEEYEQVFINFIDR-FLEFNPESRLQTFEVDYKRREIRGFKDRIGTAINRG 107
Query: 139 KKEQQFVN----FVGRALLLTSISSMELERF---SLLINNKRDISLQNTWISSILNRRVK 191
+ V+ + ++ + M L F +L+ RD +L + + S+ +
Sbjct: 108 IRLLDAVSSTMYWEEECIMYPYLEFMPLNLFTSKTLVTLKLRDSALNDPGLLSMPCLKFM 167
Query: 192 ILRIHSSFYQLPFSALTSHYLFNCTSLEELELV----LHVSSTIKFPSISVHFGHLKLLK 247
ILR + L S C LE+L LV H+ + P V LK
Sbjct: 168 ILREVRWSGTMSLEKLVS----GCPVLEDLTLVRTLICHIDDD-ELPVTRVRSRSLKTFY 222
Query: 248 L---YGIFFKIDTSSDCLTLNLPLLRKFDIKNCNWSGGKDLIVEAPLLEIVSIEQDIEF 303
+ YG+ + L ++ P L +K ++ G IV L + IE +I+F
Sbjct: 223 VPLAYGVGCRSRVPDKVLEIDAPGLENMTLKEDHFDG----IVVKNLTSLFMIELNIKF 277
>At5g22730 unknown protein
Length = 457
Score = 53.9 bits (128), Expect = 2e-07
Identities = 34/79 (43%), Positives = 45/79 (56%), Gaps = 12/79 (15%)
Query: 20 EDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNPL 79
ED I SKLP+SLI ++L +LP KD+VRT LS RW + W I +LDLD + +YN
Sbjct: 18 EDLI-SKLPDSLITQILLYLPIKDIVRTSSLSSRWKSLWL-LIPRLDLDSEEFQ-DYNAF 74
Query: 80 CSYC---------EEICTD 89
+ E+IC D
Sbjct: 75 VGFMNKFIDFSGEEKICLD 93
>At1g69630 hypothetical protein
Length = 451
Score = 53.9 bits (128), Expect = 2e-07
Identities = 29/60 (48%), Positives = 38/60 (63%), Gaps = 2/60 (3%)
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNPLCSYCE 84
SKLP+ L+ VL LPTKDVV+T VLS+RW N W + LDLD+ D +N S+ +
Sbjct: 21 SKLPDCLLCEVLLNLPTKDVVKTSVLSRRWRNLW-KHVPGLDLDNTDFQ-EFNTFLSFVD 78
Score = 31.2 bits (69), Expect = 1.1
Identities = 29/107 (27%), Positives = 48/107 (44%), Gaps = 15/107 (14%)
Query: 174 DISLQNTWISSILNRRVKILRIHSSFY-----QLPFSALTSHYLFNCTSLEELELVLHVS 228
DI L WI++I+ R+V+ + + Y QLP S ++ C SL L+L
Sbjct: 105 DIFLIGRWINTIVTRKVQHIDVLDDSYGSWEVQLPSS------IYTCESLVSLKLCGLTL 158
Query: 229 STIKFPSISVHFGHLKLLKLYGIFFKIDTSSDCLTLNLPLLRKFDIK 275
++ +F S+ LK++ L F D + L P+L I+
Sbjct: 159 ASPEFVSLP----SLKVMDLIITKFADDMGLETLITKCPVLESLTIE 201
>At1g66310 unknown protein
Length = 442
Score = 53.9 bits (128), Expect = 2e-07
Identities = 88/317 (27%), Positives = 139/317 (43%), Gaps = 77/317 (24%)
Query: 27 LPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNPLCSYCEEI 86
LPESL+ +L LPTKDVV+T VLS +W N W + LDLD D + N N S+ +
Sbjct: 24 LPESLLCHILLNLPTKDVVKTSVLSSKWRNLW-RLVPGLDLDSSDFTEN-NTFVSFIDRF 81
Query: 87 CTDYCYDY-DKYKLTVCPRCHVCGNH-KANLRDA----EVYERRVEIDFESGMKIESGKK 140
+ + Y K+KL C++ G+ N A +V +R+V + + G
Sbjct: 82 MSFHSDLYLKKFKLRFF--CNLNGDEVSENAHIARWINDVVKRKVH-----NLDLTWGAV 134
Query: 141 EQQFVNFVGRALLLTSISSMELERFSLL------------INNKRDISLQNTWISSIL-- 186
E + ++ +L+ + + L L + D++L+ IS L
Sbjct: 135 EIPPILYLCNSLVSLKLCGVTLPNLELTSLPCVKVIVLEWVKFANDLALE-MLISGCLVL 193
Query: 187 ---------NRRVKILRIHSSFYQLPFSALTSHYLFNCTSLEEL--ELVLHVSSTIKFPS 235
N VKILR+ SS L FS +N +S + L +LVL +++
Sbjct: 194 ESLTLCRRPNDNVKILRV-SSQSLLRFS-------YNGSSYKGLHDDLVLEINAP----- 240
Query: 236 ISVHFGHLKLLKLYG----IFFKIDTSSDCL--TLNLPLLRKFDIKN-------CNWSGG 282
LK+LKL+ F +TSS + +N+ L +KFD K+ CN+ G
Sbjct: 241 ------KLKILKLFSHQLTTSFIRNTSSSIVEADINIGLGKKFDPKDLPKRNVICNFLAG 294
Query: 283 ----KDLIVEAPLLEIV 295
K+L + LE++
Sbjct: 295 ISSVKNLFIAPCTLEVI 311
>At5g18780 putative protein
Length = 441
Score = 52.8 bits (125), Expect = 3e-07
Identities = 33/66 (50%), Positives = 42/66 (63%), Gaps = 3/66 (4%)
Query: 20 EDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNPL 79
ED I S LPE L+ +LSFL TKD VRT VLS RW + W + +LDLD D S + NP
Sbjct: 10 EDRI-SILPEPLLCHILSFLRTKDSVRTSVLSSRWRDLWL-WVPRLDLDKSDFS-DDNPS 66
Query: 80 CSYCEE 85
S+ ++
Sbjct: 67 ASFVDK 72
>At1g51370 unknown protein
Length = 435
Score = 52.0 bits (123), Expect = 6e-07
Identities = 38/93 (40%), Positives = 48/93 (50%), Gaps = 15/93 (16%)
Query: 19 EEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNP 78
EED I S+LPE LI+ +L L TKD VRT LS +W W S+ LDLD S N N
Sbjct: 17 EEDRI-SQLPEPLISEILFHLSTKDSVRTSALSTKWRYLW-QSVPGLDLDPY-ASSNTNT 73
Query: 79 LCSYCE------------EICTDYCYDYDKYKL 99
+ S+ E ++ D Y +DKY L
Sbjct: 74 IVSFVESFFDSHRDSWIRKLRLDLGYHHDKYDL 106
>At1g05080 hypothetical protein
Length = 439
Score = 51.6 bits (122), Expect = 8e-07
Identities = 28/52 (53%), Positives = 34/52 (64%), Gaps = 1/52 (1%)
Query: 20 EDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDID 71
ED I S LPE L+ +L LPTKDVV T +LSKRW++ WT T DD+D
Sbjct: 12 EDRI-SVLPEDLLVVILDLLPTKDVVATMILSKRWLSIWTMVRTLEYTDDMD 62
>At3g52670 putative protein
Length = 384
Score = 51.2 bits (121), Expect = 1e-06
Identities = 29/61 (47%), Positives = 40/61 (65%), Gaps = 4/61 (6%)
Query: 19 EEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNP 78
+ED +N +LPE LI R+LSFLPT+ V+ T VLSK+W + W + L+ D D Y NP
Sbjct: 8 KEDRMN-QLPEDLILRILSFLPTELVIATSVLSKQWRSLW-KLVPNLEFDSDD--YEINP 63
Query: 79 L 79
+
Sbjct: 64 V 64
>At5g22700 unknown protein
Length = 452
Score = 50.4 bits (119), Expect = 2e-06
Identities = 29/78 (37%), Positives = 44/78 (56%), Gaps = 3/78 (3%)
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSS-ITKLDLDDIDLSYNYNPLCSYC 83
S LP+ L+ ++LS LPTK+ V T +LS RW + W S+ + +D+D D + + S
Sbjct: 24 SSLPDELLCQILSNLPTKNAVTTSILSTRWRSIWLSTPVLDIDIDAFDDATTFVSFASRF 83
Query: 84 EEICTDYCYDYDKYKLTV 101
E D C K+KL+V
Sbjct: 84 LEFSKDSC--LHKFKLSV 99
>At1g66300 unknown protein
Length = 456
Score = 50.4 bits (119), Expect = 2e-06
Identities = 45/130 (34%), Positives = 67/130 (50%), Gaps = 12/130 (9%)
Query: 10 KSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDD 69
+S+ S E D + LPESLI VL L TKDV++ CVLS +W W + LDLD
Sbjct: 18 ESSDKKSGYEVDWVRD-LPESLICHVLLNLSTKDVIKNCVLSTKWRYLW-RYVPGLDLDC 75
Query: 70 IDLSYNYNPLCSYCEEICTDYCYDY-DKYKLTVCPRCHVCGNHK-ANLRDA----EVYER 123
D + YN S+ + + Y +K+KL C + G+ + N + A +V +R
Sbjct: 76 SDFT-EYNTFVSFVDRFLSTNTESYLNKFKLGF--DCDLVGDEETGNAQMARWINDVVKR 132
Query: 124 RVE-IDFESG 132
+V+ +D E G
Sbjct: 133 KVQHLDLEWG 142
>At3g14710 unknown protein
Length = 442
Score = 50.1 bits (118), Expect = 2e-06
Identities = 53/196 (27%), Positives = 89/196 (45%), Gaps = 21/196 (10%)
Query: 141 EQQFVNFVGRALLLTSISSMELERFSLLINNKRDISLQNTWISSILNRRVKILRIHSSFY 200
+ +F +FV R ++ I S + F L N D +L +W+S +L R ++ L + +
Sbjct: 80 DSRFTDFVER--VVGDIGSPHISSFYLCSVNTFDETLLFSWLSKVLKRNLQRLVVTCNDL 137
Query: 201 QL-PFSALTSHYLFNCTSLEELELVLHVSSTIKFPSISVHFGHLKLLKLYGI-FFKIDTS 258
++ FS+L ++ SL EL L I P+I +LK L L F + +
Sbjct: 138 EIISFSSLFPSFV----SLVELRLRTKSILDISAPAI---LPNLKFLSLEDARIFNMSSV 190
Query: 259 SDCLTLNLPLLRKFDIKNCNWSGGKDLIVEAPLLEIVSIEQDIEFYNAASHDLHSQS--- 315
S L LN P+L F+ C +I+++PLL I E + S + + S
Sbjct: 191 SKNLVLNFPVLETFEASYCRCFRTDTVIIDSPLLRI------FEMFKCTSEHVPNPSQVC 244
Query: 316 -IKFNALHLKQFTYSG 330
I+ A L++ T+ G
Sbjct: 245 KIRVLASKLEKITFCG 260
Score = 48.5 bits (114), Expect = 6e-06
Identities = 29/60 (48%), Positives = 38/60 (63%), Gaps = 2/60 (3%)
Query: 10 KSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDD 69
K T++ S ED +S L ES+++ +LS LPT + V T VLSK W N WT +IT L DD
Sbjct: 15 KKTKLCSENFEDKFSSLL-ESVVSIILSQLPTAEAVSTSVLSKSWKNIWT-NITDLHFDD 72
>At1g78760 hypothetical protein
Length = 452
Score = 49.7 bits (117), Expect = 3e-06
Identities = 24/40 (60%), Positives = 31/40 (77%), Gaps = 1/40 (2%)
Query: 19 EEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRW 58
EED + S LPESLI +++ +PTKDVV++ VLSKRW N W
Sbjct: 14 EEDRL-SNLPESLICQIMLNIPTKDVVKSSVLSKRWRNLW 52
Score = 29.6 bits (65), Expect = 3.1
Identities = 33/141 (23%), Positives = 59/141 (41%), Gaps = 11/141 (7%)
Query: 144 FVNFVGRALLLTSISSMELERFSLLINNK-RDISLQNTWISSILNRRVKILRI-----HS 197
FV+FV R L L S + R + + R IS WI+ ++++++K+L + +
Sbjct: 71 FVSFVDRFLALDRESCFQSFRLRYDCDEEERTISNVKRWINIVVDQKLKVLDVLDYTWGN 130
Query: 198 SFYQLPFSALTSHYLFNCTSLEELELVLHVSSTIKFPSISVHFGHLKLLKLYGIFFKIDT 257
Q+P S T L SL+ ++L I P + V L ++K
Sbjct: 131 EEVQIPPSVYTCESL---VSLKLCNVILPNPKVISLPLVKVI--ELDIVKFSNALVLEKI 185
Query: 258 SSDCLTLNLPLLRKFDIKNCN 278
S C L ++ + + + N
Sbjct: 186 ISSCSALESLIISRSSVDDIN 206
>At1g66320 hypothetical protein
Length = 457
Score = 49.7 bits (117), Expect = 3e-06
Identities = 29/79 (36%), Positives = 46/79 (57%), Gaps = 2/79 (2%)
Query: 6 SKVHKSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKL 65
S +S+ S E D + LPESL+ +VL +PTKD+V+T V+S W + W + L
Sbjct: 14 SSFPESSDKNSGDEGDWVRRDLPESLLFQVLLNIPTKDLVKTSVVSPEWRHLW-RCVPGL 72
Query: 66 DLDDIDLSYNYNPLCSYCE 84
DLD+ D + ++ L S+ +
Sbjct: 73 DLDEADFT-QFDTLVSFID 90
>At3g58940 putative protein
Length = 618
Score = 49.3 bits (116), Expect = 4e-06
Identities = 25/47 (53%), Positives = 29/47 (61%), Gaps = 1/47 (2%)
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDID 71
S LPE + +LSFLPTK T VLSK W+N W T LD+DD D
Sbjct: 5 SNLPEEVRCHILSFLPTKHAALTSVLSKSWLNLWKFE-TNLDIDDSD 50
>At3g51530 unknown protein
Length = 455
Score = 49.3 bits (116), Expect = 4e-06
Identities = 27/67 (40%), Positives = 43/67 (63%), Gaps = 2/67 (2%)
Query: 10 KSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDD 69
KS++ + ED I S+LPE L+ ++LS LPTK V+ T VLSKRW++ W + +L+ +
Sbjct: 19 KSSRKPGFMSEDRI-SELPEVLLLQILSSLPTKLVISTSVLSKRWLSLW-KMVQRLEFES 76
Query: 70 IDLSYNY 76
Y++
Sbjct: 77 SRNIYDF 83
>At1g13780 hypothetical protein
Length = 451
Score = 49.3 bits (116), Expect = 4e-06
Identities = 26/53 (49%), Positives = 35/53 (65%), Gaps = 1/53 (1%)
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYN 77
S+LPESLI+++L LPTK V+T VLS RW N W ++ LDL+ D + N
Sbjct: 8 SELPESLISQILLHLPTKASVKTSVLSTRWKNLWL-NVPGLDLNCRDFPFQNN 59
>At4g00320
Length = 773
Score = 48.9 bits (115), Expect = 5e-06
Identities = 24/44 (54%), Positives = 32/44 (72%), Gaps = 1/44 (2%)
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLD 68
S+LP+ L+ +VLSFLPTKD VRT +LS RW + W + KL+ D
Sbjct: 5 SQLPDELLVKVLSFLPTKDAVRTSLLSMRWKSLW-MWLPKLEYD 47
Score = 42.4 bits (98), Expect = 5e-04
Identities = 28/68 (41%), Positives = 42/68 (61%), Gaps = 6/68 (8%)
Query: 18 IEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYN 77
I++D I S LP+++I +LSFLPTK T VL+KRW + + LD D+ S+ ++
Sbjct: 276 IKKDGI-SGLPDAMICHILSFLPTKVAASTTVLAKRW-KPLLAFMPNLDFDE---SFRFD 330
Query: 78 PLCSYCEE 85
P + CEE
Sbjct: 331 PRMT-CEE 337
>At2g33610 putative SWI/SNF complex subunit SW13
Length = 469
Score = 48.9 bits (115), Expect = 5e-06
Identities = 22/52 (42%), Positives = 33/52 (63%), Gaps = 2/52 (3%)
Query: 80 CSYCEEICTDYCYDYDKYKLTVCPRCHVCGNHKANLRDAEVYERRVEIDFES 131
C+ C+ IC+ C+ DKY LT+C RC+V N++ + +E +RVEI ES
Sbjct: 175 CNGCKAICSIACFACDKYDLTLCARCYVRSNYRVGINSSEF--KRVEISEES 224
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,040,661
Number of Sequences: 26719
Number of extensions: 388131
Number of successful extensions: 1744
Number of sequences better than 10.0: 211
Number of HSP's better than 10.0 without gapping: 188
Number of HSP's successfully gapped in prelim test: 23
Number of HSP's that attempted gapping in prelim test: 1467
Number of HSP's gapped (non-prelim): 293
length of query: 406
length of database: 11,318,596
effective HSP length: 102
effective length of query: 304
effective length of database: 8,593,258
effective search space: 2612350432
effective search space used: 2612350432
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146790.10


