

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146789.8 + phase: 0 /pseudo
(2263 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
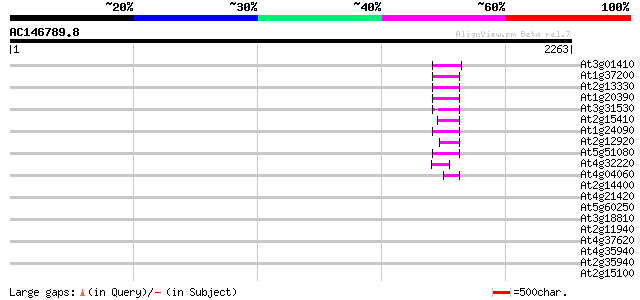
Score E
Sequences producing significant alignments: (bits) Value
At3g01410 putative RNase H 74 7e-13
At1g37200 hypothetical protein 74 1e-12
At2g13330 F14O4.9 72 3e-12
At1g20390 hypothetical protein 72 4e-12
At3g31530 hypothetical protein 67 2e-10
At2g15410 putative retroelement pol polyprotein 65 3e-10
At1g24090 unknown protein 62 5e-09
At2g12920 pseudogene 57 9e-08
At5g51080 unknown protein 56 3e-07
At4g32220 hypothetical protein 49 3e-05
At4g04060 putative transposon protein 46 3e-04
At2g14400 putative retroelement pol polyprotein 38 0.059
At4g21420 hypothetical protein 37 0.10
At5g60250 unknown protein 37 0.17
At3g18810 protein kinase, putative 37 0.17
At2g11940 putative retroelement gag/pol polyprotein 35 0.50
At4g37620 putative protein 35 0.66
At4g35940 putative protein 34 1.1
At2g35940 putative homeodomain transcription factor 33 1.5
At2g15100 putative retroelement pol polyprotein 33 1.5
>At3g01410 putative RNase H
Length = 290
Score = 74.3 bits (181), Expect = 7e-13
Identities = 39/118 (33%), Positives = 64/118 (54%), Gaps = 2/118 (1%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G GAVL + + + + TNN+AEY A ++G+ A+D KN+ + GDS LV
Sbjct: 167 GAGAVLRASDNSVLFYLREGVGNATNNVAEYRALLLGLRSALDKGFKNVHVLGDSMLVCM 226
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMAD--ALATVFNDQGES 1820
Q++G W+T H + A+ L+ F ++ HI R++N AD A + +F G++
Sbjct: 227 QVQGAWKTNHPKMAELCKQAKELMNSFKTFDIKHIAREKNSEADKQANSAIFLADGQT 284
>At1g37200 hypothetical protein
Length = 1564
Score = 73.6 bits (179), Expect = 1e-12
Identities = 43/110 (39%), Positives = 61/110 (55%)
Query: 1704 NGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVI 1763
+G G L +P G I + RL F+ +NN +EYEA I GI+ A +RI++I + DS LV
Sbjct: 1054 SGEGIQLTSPTGEVIEQSFRLGFNASNNESEYEALIDGIKLAQGMRIRDIHAHSDSQLVT 1113
Query: 1764 NQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
+Q G++E + Y + + L F EL IPR EN DALA +
Sbjct: 1114 SQFHGEYEAKDERMEAYLELVKTLTQQFESFELTRIPRGENTSTDALAAL 1163
>At2g13330 F14O4.9
Length = 889
Score = 72.4 bits (176), Expect = 3e-12
Identities = 41/110 (37%), Positives = 60/110 (54%)
Query: 1704 NGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVI 1763
+G+G L +P G I + +L F+ +NN +EYEA I GI+ A + I+ I Y DS LV
Sbjct: 656 SGVGIQLTSPTGEVIEQSLQLGFNASNNESEYEALIAGIKLAQEKGIREIHAYSDSQLVT 715
Query: 1764 NQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
+Q G++E + Y + + L F +L IPR EN AD LA +
Sbjct: 716 SQFHGEYEAKDERMEAYLELVKTLAQQFESFKLTRIPRGENTSADTLAAL 765
>At1g20390 hypothetical protein
Length = 1791
Score = 72.0 bits (175), Expect = 4e-12
Identities = 39/110 (35%), Positives = 60/110 (54%)
Query: 1704 NGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVI 1763
+GIG L++P + + RLRF TNN+AEYE I G+ A ++I I + DS L+
Sbjct: 1223 SGIGIRLVSPTAEVLEQSFRLRFVATNNVAEYEVLIAGLRLAAGMQITTIHAFTDSQLIA 1282
Query: 1764 NQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
Q+ G++E + + Y + + F +L IPR +N ADALA +
Sbjct: 1283 GQLSGEYEAKNEKMDAYLKIVQLMTKDFENFKLSKIPRGDNAPADALAAL 1332
>At3g31530 hypothetical protein
Length = 831
Score = 66.6 bits (161), Expect = 2e-10
Identities = 39/110 (35%), Positives = 57/110 (51%), Gaps = 11/110 (10%)
Query: 1704 NGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVI 1763
+G+G L +P G + TNN+AEYEA + G+ A L+I + DS L+
Sbjct: 427 SGVGIRLTSPTG-----------EATNNVAEYEALVAGLNLAWGLKIGKTRAFCDSQLIA 475
Query: 1764 NQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
NQ G++ T + Y + + L F++ EL IPR EN ADALA +
Sbjct: 476 NQFNGEYTTQDKKMEAYLIHVQNLAKNFDEFELTRIPRGENTSADALAAL 525
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 65.5 bits (158), Expect = 3e-10
Identities = 37/89 (41%), Positives = 48/89 (53%)
Query: 1725 RFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYA 1784
RF +NN AEYEA I G+ A + +K I+ Y DS LV +Q G +E + + Y
Sbjct: 1189 RFPASNNEAEYEALIAGLRLAHGIEVKKIQAYCDSQLVASQFSGNYEAKNERMDAYLKVV 1248
Query: 1785 RRLLTFFNKVELHHIPRDENQMADALATV 1813
R L F EL IPR +N ADALA +
Sbjct: 1249 RELSYNFEVFELTKIPRSDNAPADALAVL 1277
>At1g24090 unknown protein
Length = 535
Score = 61.6 bits (148), Expect = 5e-09
Identities = 42/113 (37%), Positives = 58/113 (51%), Gaps = 3/113 (2%)
Query: 1704 NGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVI 1763
+G AVL T G+ I + TNN AEY A I+G++ AI+ KNI++ GDS LV
Sbjct: 232 SGAAAVLKTEDGSLICRVRQGLGIATNNAAEYHALILGLKYAIEKGYKNIKVKGDSKLVC 291
Query: 1764 ---NQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
QIKG+W+ H L A+ L E+ H+ R+ N AD A +
Sbjct: 292 MQKQQIKGQWKVNHEVLAKLHKEAKLLCNKCVSFEISHVLRNLNADADEQANL 344
>At2g12920 pseudogene
Length = 863
Score = 57.4 bits (137), Expect = 9e-08
Identities = 31/81 (38%), Positives = 48/81 (58%)
Query: 1733 AEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFFN 1792
AEYEA + G++ A L+I I + DS+LV NQ G++ + Y +++ L F+
Sbjct: 559 AEYEALVAGLKLAQGLKIAKIRAFCDSSLVANQFNGEYTARDERMGAYLTHSQDLAKQFD 618
Query: 1793 KVELHHIPRDENQMADALATV 1813
+ EL IPR +N+ ADALA +
Sbjct: 619 EFELSRIPRGKNKSADALAAL 639
>At5g51080 unknown protein
Length = 322
Score = 55.8 bits (133), Expect = 3e-07
Identities = 37/110 (33%), Positives = 54/110 (48%)
Query: 1704 NGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVI 1763
+G AVL T G+ I + TNN AEY I+G++ AI+ I++ DS LV
Sbjct: 201 SGAAAVLKTEDGSLIFKMRQGLGIATNNAAEYHGLILGLKHAIEKGYTKIKVKTDSKLVC 260
Query: 1764 NQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
Q+KG+W+ H L A++L E+ H+ R N AD A +
Sbjct: 261 MQMKGQWKVNHEVLSKLHKEAKQLSDKCLSFEISHVLRSLNSDADEQANM 310
>At4g32220 hypothetical protein
Length = 452
Score = 49.3 bits (116), Expect = 3e-05
Identities = 25/72 (34%), Positives = 43/72 (59%)
Query: 1700 SMYTNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDS 1759
S+ + +G +L +P + + RL+F +NN AEYEA + G+ A L K I+ + DS
Sbjct: 218 SLQGSSLGILLQSPTREILEQSLRLQFKASNNEAEYEALLAGLRLAKGLGAKQIKAFSDS 277
Query: 1760 ALVINQIKGKWE 1771
LV+++ G++E
Sbjct: 278 QLVVSRFSGEFE 289
>At4g04060 putative transposon protein
Length = 375
Score = 45.8 bits (107), Expect = 3e-04
Identities = 25/64 (39%), Positives = 35/64 (54%)
Query: 1750 IKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADA 1809
I++I + DS LVI+Q G++E + Y + + L F EL IPR EN ADA
Sbjct: 3 IRDIHAHSDSQLVISQFHGEYEAKDERMEAYLELVKTLTQQFESFELTMIPRGENTSADA 62
Query: 1810 LATV 1813
LA +
Sbjct: 63 LAAL 66
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 38.1 bits (87), Expect = 0.059
Identities = 19/53 (35%), Positives = 28/53 (51%)
Query: 1761 LVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
+++NQ G +E + + Y + L F K EL IPR +N ADALA +
Sbjct: 790 IIVNQFNGDYEAKDSRMEAYLHVVKDLAKNFRKFELIRIPRGQNTTADALAAL 842
>At4g21420 hypothetical protein
Length = 229
Score = 37.4 bits (85), Expect = 0.10
Identities = 20/85 (23%), Positives = 46/85 (53%)
Query: 1727 DCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARR 1786
+ T+ +AE+ A G+E A++ + ++ + GD+ ++++ I + + +Y +
Sbjct: 108 EATSTMAEFAALKRGLELALENGLTDLWLEGDAKIIMDIISRRGRLRCEKTNKHVNYIKV 167
Query: 1787 LLTFFNKVELHHIPRDENQMADALA 1811
++ N L H+ R+ N++AD LA
Sbjct: 168 VMPELNNCVLSHVYREGNRVADKLA 192
>At5g60250 unknown protein
Length = 655
Score = 36.6 bits (83), Expect = 0.17
Identities = 22/79 (27%), Positives = 39/79 (48%)
Query: 1733 AEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFFN 1792
AE +A I G+ EA+ L IK+I + DS + + GKW + D + ++ F+
Sbjct: 198 AELKALIRGLTEALKLGIKHIVFFCDSYPIFQYVTGKWMAKQKKISLLLDDLQSIMQHFS 257
Query: 1793 KVELHHIPRDENQMADALA 1811
+ + R++ + A LA
Sbjct: 258 SYQHVLVARNDVKFAYKLA 276
>At3g18810 protein kinase, putative
Length = 700
Score = 36.6 bits (83), Expect = 0.17
Identities = 17/33 (51%), Positives = 21/33 (63%)
Query: 438 NNHTNNNHTNNSHTNNTLNKISNNNLINNAHNN 470
+N+ NNN NN+ NN NK +NNN NN NN
Sbjct: 83 DNNNNNNGNNNNDNNNGNNKDNNNNGNNNNGNN 115
Score = 34.3 bits (77), Expect = 0.86
Identities = 14/33 (42%), Positives = 21/33 (63%)
Query: 438 NNHTNNNHTNNSHTNNTLNKISNNNLINNAHNN 470
NN+ +NN+ NN + NN N +N + NN +NN
Sbjct: 79 NNNNDNNNNNNGNNNNDNNNGNNKDNNNNGNNN 111
Score = 34.3 bits (77), Expect = 0.86
Identities = 15/33 (45%), Positives = 21/33 (63%)
Query: 438 NNHTNNNHTNNSHTNNTLNKISNNNLINNAHNN 470
N++ NNN+ NN++ NN N NNN NN + N
Sbjct: 82 NDNNNNNNGNNNNDNNNGNNKDNNNNGNNNNGN 114
Score = 33.1 bits (74), Expect = 1.9
Identities = 14/32 (43%), Positives = 21/32 (64%)
Query: 438 NNHTNNNHTNNSHTNNTLNKISNNNLINNAHN 469
NN+ + N+ NN++ NN N +NNN NN +N
Sbjct: 70 NNNNDGNNGNNNNDNNNNNNGNNNNDNNNGNN 101
Score = 32.7 bits (73), Expect = 2.5
Identities = 15/33 (45%), Positives = 19/33 (57%)
Query: 438 NNHTNNNHTNNSHTNNTLNKISNNNLINNAHNN 470
NN+ NNN+ NN+ N N NNN NN + N
Sbjct: 87 NNNGNNNNDNNNGNNKDNNNNGNNNNGNNNNGN 119
Score = 32.3 bits (72), Expect = 3.3
Identities = 15/33 (45%), Positives = 18/33 (54%)
Query: 438 NNHTNNNHTNNSHTNNTLNKISNNNLINNAHNN 470
NN NNN NN+ +N N + NN NN NN
Sbjct: 106 NNGNNNNGNNNNGNDNNGNNNNGNNNDNNNQNN 138
Score = 32.3 bits (72), Expect = 3.3
Identities = 17/38 (44%), Positives = 26/38 (67%), Gaps = 2/38 (5%)
Query: 433 HHKILNNHTNNNHTNNSHTNNTLNKISNNNLINNAHNN 470
++K NN+ NNN+ NN++ N+ N +NNN NN +NN
Sbjct: 100 NNKDNNNNGNNNNGNNNNGND--NNGNNNNGNNNDNNN 135
Score = 31.2 bits (69), Expect = 7.2
Identities = 16/35 (45%), Positives = 19/35 (53%), Gaps = 2/35 (5%)
Query: 438 NNHTNNNHTNNSHTNNTLNKISNNN--LINNAHNN 470
NN+ NNN N + NN N NNN NN +NN
Sbjct: 92 NNNDNNNGNNKDNNNNGNNNNGNNNNGNDNNGNNN 126
Score = 30.8 bits (68), Expect = 9.5
Identities = 14/33 (42%), Positives = 21/33 (63%), Gaps = 2/33 (6%)
Query: 438 NNHTNNNHTNNSHTNNTLNKISNNNLINNAHNN 470
NN+ NN+ +N++ NN N + NN NN +NN
Sbjct: 86 NNNNGNNNNDNNNGNNKDNNNNGNN--NNGNNN 116
>At2g11940 putative retroelement gag/pol polyprotein
Length = 1212
Score = 35.0 bits (79), Expect = 0.50
Identities = 20/54 (37%), Positives = 25/54 (46%)
Query: 1760 ALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
A + G +E + Y D R L F K EL +PR EN ADALA +
Sbjct: 740 ATTASDYSGDYEAKDNRMEAYLDLLRELAEKFEKFELIKVPRAENSAADALAAL 793
>At4g37620 putative protein
Length = 132
Score = 34.7 bits (78), Expect = 0.66
Identities = 25/82 (30%), Positives = 40/82 (48%), Gaps = 8/82 (9%)
Query: 1733 AEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFFN 1792
AE A ++ +++A DL ++ I DS +I I T L +L +
Sbjct: 37 AEALAMLLALQQAKDLGFTSLSIASDSQQLIKAI-----TSRTPLTELHGILHDILLLAS 91
Query: 1793 K---VELHHIPRDENQMADALA 1811
+ V H IPR+EN++ADAL+
Sbjct: 92 EIGFVRFHSIPRNENRLADALS 113
>At4g35940 putative protein
Length = 451
Score = 33.9 bits (76), Expect = 1.1
Identities = 19/62 (30%), Positives = 26/62 (41%), Gaps = 9/62 (14%)
Query: 423 HNILNSHKILHHKILNNHTNNNHT---------NNSHTNNTLNKISNNNLINNAHNNLDH 473
HN N +I + LN NNN+ N H NN +I +N HNN +
Sbjct: 188 HNNNNEKRIEKQQPLNGRHNNNNEKLMEKQQPLNGRHNNNNEKRIEKQQPLNGRHNNKEK 247
Query: 474 QE 475
Q+
Sbjct: 248 QK 249
>At2g35940 putative homeodomain transcription factor
Length = 680
Score = 33.5 bits (75), Expect = 1.5
Identities = 13/25 (52%), Positives = 18/25 (72%)
Query: 438 NNHTNNNHTNNSHTNNTLNKISNNN 462
N+ NNN++NNS+ NNT +NNN
Sbjct: 39 NDSNNNNNSNNSNNNNTNTNTNNNN 63
>At2g15100 putative retroelement pol polyprotein
Length = 1329
Score = 33.5 bits (75), Expect = 1.5
Identities = 18/50 (36%), Positives = 27/50 (54%)
Query: 1764 NQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
N+ G++E A + Y + R + F + EL IPR EN A+ALA +
Sbjct: 790 NRYSGEYEAKDACMEAYLNLVREVSGRFEQFELTRIPRAENSAANALAAL 839
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.357 0.157 0.557
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,834,434
Number of Sequences: 26719
Number of extensions: 1545237
Number of successful extensions: 7645
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 7443
Number of HSP's gapped (non-prelim): 102
length of query: 2263
length of database: 11,318,596
effective HSP length: 115
effective length of query: 2148
effective length of database: 8,245,911
effective search space: 17712216828
effective search space used: 17712216828
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC146789.8


