

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146789.1 + phase: 0
(431 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
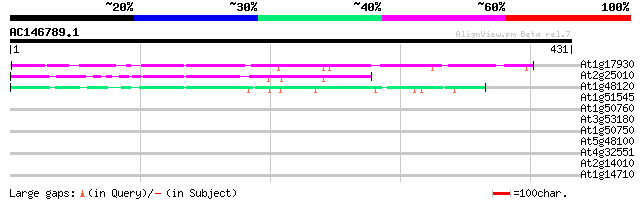
Score E
Sequences producing significant alignments: (bits) Value
At1g17930 unknown protein 100 1e-21
At2g25010 unknown protein 87 1e-17
At1g48120 serine/threonine phosphatase PP7, putative 77 2e-14
At1g51545 unknown protein 36 0.036
At1g50760 hypothetical protein 35 0.080
At3g53180 nodulin / glutamate-ammonia ligase - like protein 32 0.89
At1g50750 hypothetical protein 31 1.5
At5g48100 laccase (diphenol oxidase) 29 4.4
At4g32551 Leunig protein 29 4.4
At2g14010 hypothetical protein 28 7.5
At1g14710 unknown protein 28 9.8
>At1g17930 unknown protein
Length = 478
Score = 100 bits (250), Expect = 1e-21
Identities = 104/420 (24%), Positives = 180/420 (42%), Gaps = 61/420 (14%)
Query: 2 PMGEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEY- 60
P GEMT+TLD+V+ ++ L ++G+ + G K + + + + LG K+ E
Sbjct: 84 PCGEMTITLDEVSLILGLAVDGKPVV-GVKEKDEDPSQVCLRLLG-------KLPKGELS 135
Query: 61 GGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRS 120
G ++ L++ + A + +E+E Y R YL+Y+VG +F
Sbjct: 136 GNRVTAKWLKESFAECPKGATM-------KEIE-------YHTRAYLIYIVGSTIFATTD 181
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRALGGSVTLLTTWFLK 180
+I + YL D + Y+WGA LA+LY ++ +A + +GG +TLL W
Sbjct: 182 PSKISVDYLILFED-FEKAGEYAWGAAALAFLYRQIGNASQRSQSIIGGCLTLLQCWSYF 240
Query: 181 HFPGFFSVDLNTDYLEKYPVAARWK---LEKGHGEGITYRSLLDRIQLDDVCWRPYEEHR 237
H ++D ++P+A WK + + YR LD + +V W P+E
Sbjct: 241 H----LNIDRPKRTTRQFPLALLWKGRQQSRSKNDLFKYRKALDDLDPSNVSWCPFEGDL 296
Query: 238 EI--QDFEE--VFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPRHPTDVRDLPPPSIV 293
+I Q F++ + S + G + V H P+R ++Q+G Q IP ++PP
Sbjct: 297 DIVPQSFKDNLLLGRSRTKLIGPKVVEWHFPDRCMKQFGLCQVIP------GEVPPRKNE 350
Query: 294 QMF-IDFRTHTLKADARGVQAGEDTRRVADG------YVLWYTRVSHPQILPPIPGDLPR 346
+ D AD ++ E+ G Y+ W+ ++ +P + D
Sbjct: 351 KNHDEDLLEDMNTADEEWMRRRENIVENGGGNGDESEYMQWFNSIT----VPKLHRDTSL 406
Query: 347 PANEEQIIAEQWQRYEARSSPDTYDMVSGAVAYADAQLGQEEVMSMTPQ----QWYEAMT 402
A+ + A Q E S+ D+ + + +E VMSM+ WYEA T
Sbjct: 407 EADIMNVQAAILQFDEVASTLSLEDLHP-----EEREAIEEAVMSMSNALRVGDWYEAST 461
>At2g25010 unknown protein
Length = 509
Score = 87.4 bits (215), Expect = 1e-17
Identities = 77/286 (26%), Positives = 128/286 (43%), Gaps = 33/286 (11%)
Query: 1 MPMGEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEY 60
+P+GEMT+TLD+VA ++ L I+G + K G + M G + N+E
Sbjct: 92 LPLGEMTITLDEVALVLGLEIDGDPIVGSK-----VGDEVAMDMCGRLLGKLPSAANKE- 145
Query: 61 GGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRS 120
++ R++ ++L R +E PE+ + V+ + R YLLYL+G +F
Sbjct: 146 ---VNCSRVK---LNWLKR----TFSECPEDA-SFDVVKCH-TRAYLLYLIGSTIFATTD 193
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRALGGSVTLLTTWFLK 180
++ + YL D + Y+WGA LA LY L +A + G +TLL W
Sbjct: 194 GDKVSVKYLPLFED-FDQAGRYAWGAAALACLYRALGNASLKSQSNICGCLTLLQCW--- 249
Query: 181 HFPGFFSVDLNTDYLEK--YPVAARWKLE--KGHGEGITYRSLLDRIQLDDVCWRPYEEH 236
+F +D+ + +P+A WK + + + YR LD + + W PYE
Sbjct: 250 ---SYFHLDIGRPEKSEACFPLALLWKGKGSRSKTDLSEYRRELDDLDPSKITWCPYERF 306
Query: 237 REI----QDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
+ + + S + ++ H P+R LRQ+G Q IP
Sbjct: 307 ENLIPPHIKAKLILGRSKTTLVCFEKIELHFPDRCLRQFGKRQPIP 352
>At1g48120 serine/threonine phosphatase PP7, putative
Length = 1338
Score = 76.6 bits (187), Expect = 2e-14
Identities = 106/401 (26%), Positives = 161/401 (39%), Gaps = 65/401 (16%)
Query: 1 MPMGEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEY 60
+P GE+TVTL DV L+ L ++G + K + A L LG + +
Sbjct: 101 LPAGEITVTLQDVNILLGLRVDGPAVTGSTK---YNWADLCEDLLGHRPGPKDL-----H 152
Query: 61 GGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYC-VRCYLLYLVGCLLFGDR 119
G ++S LR+ + + DP+EV C R ++L L+ L+GD+
Sbjct: 153 GSHVSLAWLRENFRNL---------PADPDEVT------LKCHTRAFVLALMSGFLYGDK 197
Query: 120 SNKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRALGGSVTLLTTWFL 179
S + L +L + D + + SWG+ TLA LY EL A + + G + LL W
Sbjct: 198 SKHDVALTFLPLLRD-FDEVAKLSWGSATLALLYRELCRASKRTVSTICGPLVLLQLWAW 256
Query: 180 KHF----PGFFSVDLNTDYLEKY------PVAARWKL---EKGHGEGITYRSLLDRIQLD 226
+ PG D+ Y++ P+ R L E G YR D+ + +
Sbjct: 257 ERLHVGRPGRLK-DVGASYMDGIDGPLPDPLGCRASLSHKENPRGGLDFYRDQFDQQKDE 315
Query: 227 DVCWRPY-----EEHREIQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPR-- 279
V W+PY + I E W + + V H P+RVLRQ+G QTIP
Sbjct: 316 QVIWQPYTPDLLAKIPLICVSGENIWRTVAPLICFDVVEWHRPDRVLRQFGLHQTIPAPC 375
Query: 280 ------HPTDVRDLPPPSIVQMFIDFRTHTLKADAR--GVQAGE---DTRRVADGYVLWY 328
H D R S H +AR V +GE D Y+ WY
Sbjct: 376 DNEKALHAIDKRG---KSEYDWSARHSRHIGLWEARVSSVVSGEPECSPMDYNDPYMEWY 432
Query: 329 TRVSHPQILPPI----PGDLPRPANEEQIIAEQWQRYEARS 365
R++ +I+ P+ PG Q++ ++ ARS
Sbjct: 433 RRITR-RIISPMNERRPGQFLPTGFAFQVLVQRVAAIHARS 472
>At1g51545 unknown protein
Length = 629
Score = 36.2 bits (82), Expect = 0.036
Identities = 67/308 (21%), Positives = 113/308 (35%), Gaps = 65/308 (21%)
Query: 2 PMGEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEYG 61
P GE T+TL+DV L+ ++G + + + + V + E ++ N+
Sbjct: 49 PWGEATITLEDVLVLLGFSVQGSPVFAPLESSEMRDS--------VEKLEKARLENRGQD 100
Query: 62 GYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRSN 121
G + R + +S+LGR +++E +L + + +F D
Sbjct: 101 GLV---RQNLWVSSFLGRG------------DQMEH------EAFLAFWLSQFVFPDMCR 139
Query: 122 KRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRALGGSVTLLTTWFLKH 181
+ I L MA A ++ LA LY +L +VTL + + L
Sbjct: 140 RSISTKVLP-MAVRLARGERIAFAPAVLARLYRDLGQIQASAREKSTPNVTLKSLFKLVQ 198
Query: 182 ---FPGFFSVDLNTDYLEK-YPVAARWKLEKGHGEGITYRSLLDRIQLDDVCWRPYEEHR 237
+ F S + K P +RW + R+ L D WRPY +
Sbjct: 199 LWAWERFKSTSPKARVIPKGEPRISRWHSQTSKNV---------RLNLVDFDWRPYTKPL 249
Query: 238 EIQDF-----EEVFWYS-------GWI----------MCGVRRVYRHLPERVLRQYGYVQ 275
+I + EE W + G++ + GV V + P RV Q+G Q
Sbjct: 250 QIWNPPRFYPEEAMWMTVDDNLDDGFVSFARCMRVSQLVGVGIVEDYYPNRVAMQFGLAQ 309
Query: 276 TIPRHPTD 283
+P TD
Sbjct: 310 DLPGLVTD 317
>At1g50760 hypothetical protein
Length = 649
Score = 35.0 bits (79), Expect = 0.080
Identities = 27/113 (23%), Positives = 43/113 (37%), Gaps = 28/113 (24%)
Query: 199 PVAARWKLEKGHGEGITYRSLLDRIQLDDVCWRPYEEHREIQDFEEVFWYSGWIMC---- 254
P A W K + R +LD ++D WRPY R + +++ +Y MC
Sbjct: 280 PRVAIWNKLKQQTSNV--RQILDNSKIDSFEWRPYT--RSVTNWKFPQFYPEKAMCVHVS 335
Query: 255 --------------------GVRRVYRHLPERVLRQYGYVQTIPRHPTDVRDL 287
G+ +V + P RV Q+G +Q +P D +L
Sbjct: 336 PSLDEELISFARCIKDSELVGIEKVEHYFPNRVASQFGMLQDVPSPHVDRNNL 388
>At3g53180 nodulin / glutamate-ammonia ligase - like protein
Length = 845
Score = 31.6 bits (70), Expect = 0.89
Identities = 18/61 (29%), Positives = 31/61 (50%), Gaps = 1/61 (1%)
Query: 134 DGYAGMRNYSWGAMTLA-YLYGELADACRPGHRALGGSVTLLTTWFLKHFPGFFSVDLNT 192
DGYA Y GA ++ L+DAC G +L ++ F ++ GF+ ++++T
Sbjct: 328 DGYASPETYYLGAKKAREVIFLVLSDACASGDLSLMEAIDAAKDIFSRNSIGFYKLNIDT 387
Query: 193 D 193
D
Sbjct: 388 D 388
>At1g50750 hypothetical protein
Length = 816
Score = 30.8 bits (68), Expect = 1.5
Identities = 16/62 (25%), Positives = 30/62 (47%), Gaps = 1/62 (1%)
Query: 217 RSLLDRIQLDDVCWRPYEEHREIQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQT 276
R +LD +++D V W P + +++ + G+ V + P RV Q+G +Q
Sbjct: 258 RQILDSLKIDTV-WIPASPNLDVEFVSFARCIKVSQLVGIDNVEHYFPNRVASQFGMLQD 316
Query: 277 IP 278
+P
Sbjct: 317 VP 318
>At5g48100 laccase (diphenol oxidase)
Length = 565
Score = 29.3 bits (64), Expect = 4.4
Identities = 20/66 (30%), Positives = 29/66 (43%), Gaps = 7/66 (10%)
Query: 317 TRRVADGYVLWYTRVSHPQILPPIPGDLPRPANEEQIIAEQWQRYEARSSPDTYDMVSGA 376
TR G + Y R PQILP P+ +E II +W + + R + + GA
Sbjct: 128 TRATVHGLIFVYPRP--PQILP-----FPKADHEVPIILGEWWKRDVREVVEEFVRTGGA 180
Query: 377 VAYADA 382
+DA
Sbjct: 181 PNVSDA 186
>At4g32551 Leunig protein
Length = 931
Score = 29.3 bits (64), Expect = 4.4
Identities = 13/49 (26%), Positives = 24/49 (48%)
Query: 383 QLGQEEVMSMTPQQWYEAMTHMREQIAPILTRRRAQRPRRRHHHQDQDQ 431
Q Q +S QQ + M++ + +++ Q+ ++ HHHQ Q Q
Sbjct: 92 QQSQHPQVSQQQQQQQQQQIQMQQLLLQRAQQQQQQQQQQHHHHQQQQQ 140
>At2g14010 hypothetical protein
Length = 833
Score = 28.5 bits (62), Expect = 7.5
Identities = 24/86 (27%), Positives = 37/86 (42%), Gaps = 3/86 (3%)
Query: 118 DRSNKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRALGGSVTLLTTW 177
D+ +KR+E L +AD A + + ++Y E+ RPG GGSV L T
Sbjct: 360 DKKDKRMESRMLKAIADSEARLSSLLRATHGKHHMYEEVQ---RPGLGGKGGSVVDLDTS 416
Query: 178 FLKHFPGFFSVDLNTDYLEKYPVAAR 203
H + D+ L +Y + R
Sbjct: 417 MGVHTMKNSATDILDQVLGEYTLVQR 442
>At1g14710 unknown protein
Length = 598
Score = 28.1 bits (61), Expect = 9.8
Identities = 19/77 (24%), Positives = 38/77 (48%), Gaps = 5/77 (6%)
Query: 353 IIAEQWQRYEARSSPDTYDMVSGAVAYADAQLGQEEVMSMTPQQWYEAMTHMREQIAPIL 412
II Q +A + Y+ V G++ + +L +V++M Q + + + + I
Sbjct: 46 IIDSLCQHLQAVGDHNEYESVIGSIHHR--RLAWSQVLTM---QQFFPVADVSYNLQQIA 100
Query: 413 TRRRAQRPRRRHHHQDQ 429
+R+ Q P +RH++ DQ
Sbjct: 101 WKRQQQMPPQRHYNSDQ 117
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.139 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,348,283
Number of Sequences: 26719
Number of extensions: 457000
Number of successful extensions: 912
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 891
Number of HSP's gapped (non-prelim): 15
length of query: 431
length of database: 11,318,596
effective HSP length: 102
effective length of query: 329
effective length of database: 8,593,258
effective search space: 2827181882
effective search space used: 2827181882
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146789.1


