

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146784.13 + phase: 0 /pseudo
(36 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
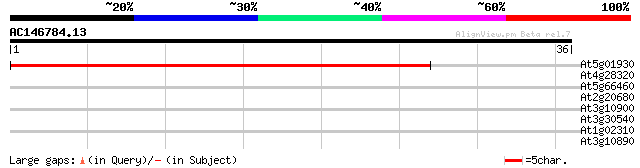
Score E
Sequences producing significant alignments: (bits) Value
At5g01930 (1-4)-beta-mannan endohydrolase-like protein 46 4e-06
At4g28320 putative (1-4)-beta-mannan endohydrolase 32 0.048
At5g66460 mannan endo-1,4-beta-mannosidase 32 0.063
At2g20680 (1-4)-beta-mannan endohydrolase 32 0.063
At3g10900 putative (1-4)-beta-mannan endohydrolase 30 0.24
At3g30540 putative beta-mannan endohydrolase 30 0.31
At1g02310 (1-4)-beta-mannan endohydrolase precursor like protein 29 0.41
At3g10890 putative (1-4)-beta-mannan endohydrolase 27 2.7
>At5g01930 (1-4)-beta-mannan endohydrolase-like protein
Length = 448
Score = 45.8 bits (107), Expect = 4e-06
Identities = 19/27 (70%), Positives = 23/27 (84%)
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSSKD 27
M AH+ED E YLGMPV+F+EFGVS+ D
Sbjct: 320 MQAHVEDAEMYLGMPVLFTEFGVSAHD 346
>At4g28320 putative (1-4)-beta-mannan endohydrolase
Length = 431
Score = 32.3 bits (72), Expect = 0.048
Identities = 14/27 (51%), Positives = 21/27 (76%)
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSSKD 27
M +HIED K L PV+F+EFG+S+++
Sbjct: 315 MQSHIEDGLKELKKPVLFTEFGLSNQN 341
>At5g66460 mannan endo-1,4-beta-mannosidase
Length = 431
Score = 32.0 bits (71), Expect = 0.063
Identities = 12/26 (46%), Positives = 18/26 (69%)
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSSK 26
+DAHI+D + L P++ +EFG S K
Sbjct: 301 LDAHIQDAQNVLHKPIILAEFGKSMK 326
>At2g20680 (1-4)-beta-mannan endohydrolase
Length = 403
Score = 32.0 bits (71), Expect = 0.063
Identities = 14/25 (56%), Positives = 20/25 (80%)
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSS 25
M +HIED +K L PV+F+EFG+S+
Sbjct: 286 MLSHIEDGDKELKKPVLFTEFGLSN 310
>At3g10900 putative (1-4)-beta-mannan endohydrolase
Length = 408
Score = 30.0 bits (66), Expect = 0.24
Identities = 11/25 (44%), Positives = 17/25 (68%)
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSS 25
++ HIED + L PV+ +EFG+ S
Sbjct: 303 LEGHIEDAQNILKKPVILAEFGLGS 327
>At3g30540 putative beta-mannan endohydrolase
Length = 351
Score = 29.6 bits (65), Expect = 0.31
Identities = 11/25 (44%), Positives = 17/25 (68%)
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSS 25
++ HIED + L PV+ +EFG+ S
Sbjct: 287 LEGHIEDAQNNLKKPVILAEFGLGS 311
>At1g02310 (1-4)-beta-mannan endohydrolase precursor like protein
Length = 411
Score = 29.3 bits (64), Expect = 0.41
Identities = 12/24 (50%), Positives = 17/24 (70%)
Query: 3 AHIEDTEKYLGMPVVFSEFGVSSK 26
AHIED + + P++ +EFG SSK
Sbjct: 302 AHIEDCDNIIKKPLLITEFGKSSK 325
>At3g10890 putative (1-4)-beta-mannan endohydrolase
Length = 414
Score = 26.6 bits (57), Expect = 2.7
Identities = 10/32 (31%), Positives = 17/32 (52%)
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSSKDLATTQ 32
++ H+ED + L P++ EFG + TQ
Sbjct: 304 LECHLEDAQNILKKPLILGEFGKPTNTPGYTQ 335
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.130 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 736,045
Number of Sequences: 26719
Number of extensions: 14377
Number of successful extensions: 21
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 13
Number of HSP's gapped (non-prelim): 8
length of query: 36
length of database: 11,318,596
effective HSP length: 12
effective length of query: 24
effective length of database: 10,997,968
effective search space: 263951232
effective search space used: 263951232
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC146784.13


