

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146784.12 - phase: 0 /pseudo
(60 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
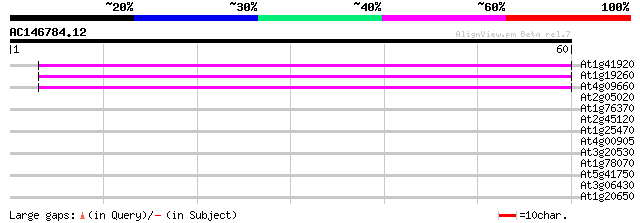
Score E
Sequences producing significant alignments: (bits) Value
At1g41920 hypothetical protein 42 6e-05
At1g19260 hypothetical protein 41 1e-04
At4g09660 putative protein 40 3e-04
At2g05020 hypothetical protein 36 0.004
At1g76370 putative protein kinase 32 0.078
At2g45120 putative C2H2-type zinc finger protein 28 1.1
At1g25470 unknown protein 27 1.9
At4g00905 putative protein 26 3.3
At3g20530 protein kinase, putative 25 7.3
At1g78070 unknown protein (At1g78070) 25 7.3
At5g41750 disease resistance protein-like 25 9.5
At3g06430 unknown protein 25 9.5
At1g20650 unknown protein 25 9.5
>At1g41920 hypothetical protein
Length = 496
Score = 42.0 bits (97), Expect = 6e-05
Identities = 21/57 (36%), Positives = 33/57 (57%)
Query: 4 RFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
RF E + +L C + L +SF +FD K+ RL++ Y DF+ DR ++ QL Y+
Sbjct: 429 RFDEVNTELLSCVASLSPIDSFHEFDQLKVLRLSEFYPQDFTHVDRRSLEHQLGLYI 485
>At1g19260 hypothetical protein
Length = 769
Score = 41.2 bits (95), Expect = 1e-04
Identities = 22/57 (38%), Positives = 33/57 (57%)
Query: 4 RFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
RF E ++ +L C S L +SF +FD L RL + Y DFS +R ++ QL+ Y+
Sbjct: 604 RFDEVNSELLICMSSLSPIDSFRQFDKSMLVRLTEFYPDDFSFVERRSLDHQLEIYL 660
>At4g09660 putative protein
Length = 664
Score = 39.7 bits (91), Expect = 3e-04
Identities = 21/57 (36%), Positives = 33/57 (57%)
Query: 4 RFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
RF E ++ +L C S L +SF +FD L RL + Y +FS +R ++ QL+ Y+
Sbjct: 548 RFDEVNSELLICMSSLSPIDSFCQFDKSMLVRLTEFYPDEFSFVERRSLDHQLEIYL 604
>At2g05020 hypothetical protein
Length = 370
Score = 35.8 bits (81), Expect = 0.004
Identities = 20/56 (35%), Positives = 32/56 (56%)
Query: 5 FRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
F E ++ +L + L +SFS+F L RL ++Y DFS +R ++ QLD Y+
Sbjct: 245 FDEVNSEILFWIASLSPMDSFSQFKKSMLVRLTELYLDDFSFVERISLDHQLDIYL 300
>At1g76370 putative protein kinase
Length = 381
Score = 31.6 bits (70), Expect = 0.078
Identities = 14/37 (37%), Positives = 22/37 (58%)
Query: 15 CFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGT 51
CFSCL+++ + + ++D L+ L D V D RGT
Sbjct: 3 CFSCLNTQTNDMRINIDTLSDLTDYASVATKIDPRGT 39
>At2g45120 putative C2H2-type zinc finger protein
Length = 314
Score = 27.7 bits (60), Expect = 1.1
Identities = 12/29 (41%), Positives = 18/29 (61%)
Query: 20 DSKNSFSKFDVDKLARLADIYHVDFSDDD 48
D+ +S +FD K+ RL D DF++DD
Sbjct: 62 DASSSSGEFDNQKMNRLDDELEFDFAEDD 90
>At1g25470 unknown protein
Length = 364
Score = 26.9 bits (58), Expect = 1.9
Identities = 16/46 (34%), Positives = 26/46 (55%), Gaps = 2/46 (4%)
Query: 5 FRERSNIVLDCFSCLDSKNSFSKFDVDKLARL--ADIYHVDFSDDD 48
F SNI+LD +S L++ +FS+F+ + L D ++F DD
Sbjct: 302 FLTDSNILLDDYSLLENDINFSRFENSLPSELPDCDFTEMEFQLDD 347
>At4g00905 putative protein
Length = 263
Score = 26.2 bits (56), Expect = 3.3
Identities = 10/24 (41%), Positives = 13/24 (53%)
Query: 6 RERSNIVLDCFSCLDSKNSFSKFD 29
R +S +VL C C K S +FD
Sbjct: 86 RPKSGVVLSCLDCFLKKGSLYRFD 109
>At3g20530 protein kinase, putative
Length = 386
Score = 25.0 bits (53), Expect = 7.3
Identities = 12/30 (40%), Positives = 17/30 (56%), Gaps = 1/30 (3%)
Query: 1 MDHR-FRERSNIVLDCFSCLDSKNSFSKFD 29
M HR F RS+ C+D+KN+ + FD
Sbjct: 10 MSHRRFNRRSSSRQSIKDCIDAKNNITTFD 39
>At1g78070 unknown protein (At1g78070)
Length = 229
Score = 25.0 bits (53), Expect = 7.3
Identities = 10/21 (47%), Positives = 12/21 (56%)
Query: 28 FDVDKLARLADIYHVDFSDDD 48
FD L L D +H DF DD+
Sbjct: 4 FDNSDLEYLVDEFHADFDDDE 24
>At5g41750 disease resistance protein-like
Length = 1068
Score = 24.6 bits (52), Expect = 9.5
Identities = 9/17 (52%), Positives = 12/17 (69%)
Query: 32 KLARLADIYHVDFSDDD 48
K R+ DIYHVDF ++
Sbjct: 331 KAHRIQDIYHVDFPSEE 347
>At3g06430 unknown protein
Length = 486
Score = 24.6 bits (52), Expect = 9.5
Identities = 11/24 (45%), Positives = 14/24 (57%)
Query: 8 RSNIVLDCFSCLDSKNSFSKFDVD 31
RSN++ D FS LD SF + D
Sbjct: 171 RSNLIDDAFSILDKMKSFPQCQPD 194
>At1g20650 unknown protein
Length = 381
Score = 24.6 bits (52), Expect = 9.5
Identities = 15/40 (37%), Positives = 19/40 (47%), Gaps = 4/40 (10%)
Query: 15 CFSCLDSKNSFSKFDVDKLARLADIYHVDFS---DDDRGT 51
CFSCL+ + + D+D AR Y D S D GT
Sbjct: 4 CFSCLNPRTKDIRVDIDN-ARCNSRYQTDSSVHGSDTTGT 42
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.141 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,330,227
Number of Sequences: 26719
Number of extensions: 41563
Number of successful extensions: 159
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 147
Number of HSP's gapped (non-prelim): 13
length of query: 60
length of database: 11,318,596
effective HSP length: 36
effective length of query: 24
effective length of database: 10,356,712
effective search space: 248561088
effective search space used: 248561088
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146784.12


