

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146784.10 + phase: 0
(168 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
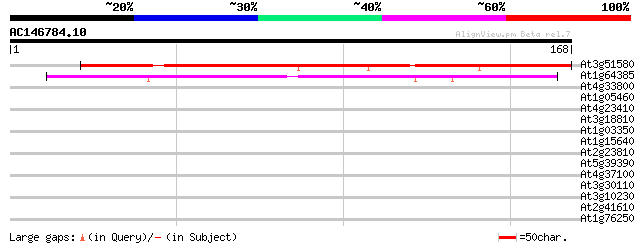
Score E
Sequences producing significant alignments: (bits) Value
At3g51580 unknown protein 141 2e-34
At1g64385 unknown protein 65 1e-11
At4g33800 unknown protein 31 0.28
At1g05460 unknown protein 30 0.61
At4g23410 unknown protein 29 1.0
At3g18810 protein kinase, putative 29 1.4
At1g03350 unknown protein 28 2.3
At1g15640 hypothetical protein 28 3.0
At2g23810 similar to senescence-associated protein 27 5.2
At5g39390 receptor protein kinase -like protein 27 6.8
At4g37100 putative protein 27 6.8
At3g30110 hypothetical protein 27 6.8
At3g10230 lycopene beta cyclase 27 6.8
At2g41610 hypothetical protein 27 6.8
At1g76250 unknown protein 26 8.9
>At3g51580 unknown protein
Length = 390
Score = 141 bits (355), Expect = 2e-34
Identities = 70/153 (45%), Positives = 101/153 (65%), Gaps = 10/153 (6%)
Query: 22 IKSESTELTLDAGKGDCVLHVTVVTPVPEASFFLRLPSFDKILTPVNGAYFLIFTVIVFA 81
I ++ ++ LD GKG C LH+ P E++ PS++K++TP+NGAYFLI +VI+F
Sbjct: 242 ISGDTNKIILDTGKGQCALHMY---PSEESTLPFHFPSYEKLVTPINGAYFLIVSVIIFG 298
Query: 82 VTWA-CCCIFKKKPRDEIPYQELEMA----LPESASATVVESAEGWDQGWDDDWDDNVAV 136
WA C C ++ +PY+ELE++ L + VE+A+ WD+GWDDDWD+N AV
Sbjct: 299 GIWAFCLCRKNRRAGSGVPYRELELSGGPGLENESGVHDVETAD-WDEGWDDDWDENNAV 357
Query: 137 KSP-VVRHAGSISANGLTSRSSNKDGWEDNWDD 168
KSP + SISANGLT+R+ N+DGW+ +WDD
Sbjct: 358 KSPGSAAKSVSISANGLTARAPNRDGWDHDWDD 390
>At1g64385 unknown protein
Length = 351
Score = 65.5 bits (158), Expect = 1e-11
Identities = 50/173 (28%), Positives = 77/173 (43%), Gaps = 23/173 (13%)
Query: 12 NIVFQVTIKQIKSESTELTLDAGKGDCVL---------HVTVVTPVPEASFFLRLPSFDK 62
+I +V+IK+ S + + L + KG C L H T S L +
Sbjct: 182 DIKVKVSIKKGGSNDSAIVLASSKGRCRLELKDLAAAAHETESDDTVSVSRPSILNISSR 241
Query: 63 ILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIPYQELEMALPESASATVVESAE-- 120
L + FL+ ++++ V ++K K R YQ L+M LP S A V +S +
Sbjct: 242 TLIVIIMISFLVLSLVIIPVI---IHVYKNKSRGNNKYQRLDMELPVSNPALVTKSDQES 298
Query: 121 ---GWDQGWDDDWD------DNVAVKSPVVRHAGSISANGLTSRSSNKDGWED 164
GW+ W DDWD D +PV+ S+S+ GL R +K+GW+D
Sbjct: 299 GDDGWNNNWGDDWDDENGGGDEEQPNTPVLPLTPSLSSRGLAPRRLSKEGWKD 351
>At4g33800 unknown protein
Length = 170
Score = 31.2 bits (69), Expect = 0.28
Identities = 16/49 (32%), Positives = 24/49 (48%), Gaps = 4/49 (8%)
Query: 118 SAEGWDQGWDDDWDDNVAVKSPVVRHAGSISANGLTSRSSNKDGWEDNW 166
S+ G + GW D +V+ SP +NG SR +KD W+ N+
Sbjct: 4 SSSGCESGWTLYLDQSVSSPSPSCFR----DSNGFDSRRRSKDSWDQNY 48
>At1g05460 unknown protein
Length = 1002
Score = 30.0 bits (66), Expect = 0.61
Identities = 15/43 (34%), Positives = 19/43 (43%), Gaps = 3/43 (6%)
Query: 126 WDDDWDDNVAVKSPVVRHAGSISANGLTSRSSNKDGWEDNWDD 168
W D W++N K G S G T + KD W D WD+
Sbjct: 868 WSDGWNNNGGTKEKNEWSDGWNSNGGGTKK---KDEWSDGWDN 907
>At4g23410 unknown protein
Length = 281
Score = 29.3 bits (64), Expect = 1.0
Identities = 17/59 (28%), Positives = 24/59 (39%), Gaps = 15/59 (25%)
Query: 77 VIVFAVTWAC--CCIFKKKPRDE---IPYQELEMALPESASATVVESAEGWDQGWDDDW 130
V++F + C CC FK R + PY M+ +S GW+Q W W
Sbjct: 227 VVIFLIAVYCVGCCAFKNAKRPQHYGFPYGRYGMS----------KSRPGWEQSWSRWW 275
>At3g18810 protein kinase, putative
Length = 700
Score = 28.9 bits (63), Expect = 1.4
Identities = 16/43 (37%), Positives = 23/43 (53%), Gaps = 4/43 (9%)
Query: 58 PSFDKILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIPY 100
P+ I+ V GA L +I+F V CC KKK + ++PY
Sbjct: 183 PNTAAIVGIVAGAGLLFLVMILFCV----CCCRKKKKKHQMPY 221
>At1g03350 unknown protein
Length = 470
Score = 28.1 bits (61), Expect = 2.3
Identities = 20/76 (26%), Positives = 33/76 (43%), Gaps = 1/76 (1%)
Query: 93 KPRDEIPYQELEMALPESASATVVESAEGWDQGWDDDWDDNVAVKSPVVRHAGSISANGL 152
KP+ + + A P+ A+++ + D GWD+ D + R GS + L
Sbjct: 381 KPKSDEAAPSQDSAKPDVAASSSTQQPSEEDLGWDEIEDMSSIDGKETSRSGGSPNRAEL 440
Query: 153 TSRSSNKDGWED-NWD 167
R S + ED +WD
Sbjct: 441 RKRLSAAEEDEDLSWD 456
>At1g15640 hypothetical protein
Length = 255
Score = 27.7 bits (60), Expect = 3.0
Identities = 20/61 (32%), Positives = 24/61 (38%), Gaps = 8/61 (13%)
Query: 38 CVLHVTVVTPVPEASFF----LRLPS----FDKILTPVNGAYFLIFTVIVFAVTWACCCI 89
C+LHV P A F LRL FD I + +LIF + CCI
Sbjct: 187 CILHVLPNLPCAAACFLYPMILRLTQSIDFFDDITEKIEDINWLIFVYFFGIFSCIICCI 246
Query: 90 F 90
F
Sbjct: 247 F 247
>At2g23810 similar to senescence-associated protein
Length = 195
Score = 26.9 bits (58), Expect = 5.2
Identities = 11/26 (42%), Positives = 17/26 (65%), Gaps = 3/26 (11%)
Query: 72 FLIFTVIVFAVTWACCCIFKKKPRDE 97
FL+F +IV++V CC F+ RD+
Sbjct: 163 FLVFLIIVYSVG---CCAFRNNKRDD 185
>At5g39390 receptor protein kinase -like protein
Length = 502
Score = 26.6 bits (57), Expect = 6.8
Identities = 19/58 (32%), Positives = 32/58 (54%), Gaps = 7/58 (12%)
Query: 61 DKILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIPYQELEMALPESASATVVES 118
+K+ V A +F +IV +++W FKKK D+I Y+EL A +S+ ++ S
Sbjct: 167 EKVAVGVGVALLFLF-IIVASLSW-----FKKK-NDKISYEELYNATSGFSSSNLIGS 217
>At4g37100 putative protein
Length = 896
Score = 26.6 bits (57), Expect = 6.8
Identities = 12/37 (32%), Positives = 18/37 (48%)
Query: 121 GWDQGWDDDWDDNVAVKSPVVRHAGSISANGLTSRSS 157
G D+G DD+WD V RH ++ GL ++
Sbjct: 694 GDDEGADDEWDRRDTETEIVCRHIDHVNMLGLNKTTT 730
>At3g30110 hypothetical protein
Length = 217
Score = 26.6 bits (57), Expect = 6.8
Identities = 16/76 (21%), Positives = 32/76 (42%)
Query: 93 KPRDEIPYQELEMALPESASATVVESAEGWDQGWDDDWDDNVAVKSPVVRHAGSISANGL 152
KP+D + SA+A ++S D+ A SP ++ ++ G
Sbjct: 7 KPKDPSIIPSTKTLTSTSAAAPALDSGSNLDKPIILKHKKKEATDSPAMKSGTKRASEGT 66
Query: 153 TSRSSNKDGWEDNWDD 168
TS+ + K ++ +W +
Sbjct: 67 TSKDNKKPYFQRSWTE 82
>At3g10230 lycopene beta cyclase
Length = 501
Score = 26.6 bits (57), Expect = 6.8
Identities = 11/31 (35%), Positives = 17/31 (54%)
Query: 136 VKSPVVRHAGSISANGLTSRSSNKDGWEDNW 166
V + +VR+ GS S+N L + + W D W
Sbjct: 378 VANAIVRYLGSPSSNSLRGDQLSAEVWRDLW 408
>At2g41610 hypothetical protein
Length = 304
Score = 26.6 bits (57), Expect = 6.8
Identities = 20/74 (27%), Positives = 36/74 (48%), Gaps = 12/74 (16%)
Query: 22 IKSESTELTLDAGKGDCVLHVTVVTPVPEASFFLRLPSFDKILTPVNGAYFLIFTVIVFA 81
+K+ LTL DCV ++ V++PV E F R+ F FL+ + V+
Sbjct: 238 VKNVVQGLTLLVLIRDCV-YLAVMSPVEEPILF-RVFVFG----------FLLLLICVYV 285
Query: 82 VTWACCCIFKKKPR 95
+T CC + +++ +
Sbjct: 286 ITKVCCLVSRRQSK 299
>At1g76250 unknown protein
Length = 434
Score = 26.2 bits (56), Expect = 8.9
Identities = 22/103 (21%), Positives = 39/103 (37%), Gaps = 9/103 (8%)
Query: 66 PVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIPYQELEMALPESASATVVESAEGWDQG 125
P+NGA TV A +C K E + E E + + + ++ W++G
Sbjct: 325 PLNGAKCRPCTV---ACNGSCVAEVMGKLNKEWSWTEWE-----NETVELCDAHGEWEKG 376
Query: 126 WDDDWDDNVAVKSPVVR-HAGSISANGLTSRSSNKDGWEDNWD 167
W+ +D+ K + R G + + N G W+
Sbjct: 377 WEKIFDETAGEKLALARKRVGGLDSRRCVEEFENMRGMAVKWE 419
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,763,386
Number of Sequences: 26719
Number of extensions: 153063
Number of successful extensions: 446
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 426
Number of HSP's gapped (non-prelim): 19
length of query: 168
length of database: 11,318,596
effective HSP length: 92
effective length of query: 76
effective length of database: 8,860,448
effective search space: 673394048
effective search space used: 673394048
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146784.10


