

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146776.6 - phase: 0
(388 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
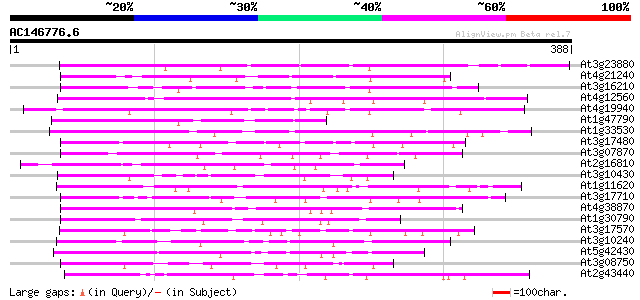
Sequences producing significant alignments: (bits) Value
At3g23880 unknown protein 151 5e-37
At4g21240 putative protein 92 4e-19
At3g16210 hypothetical protein 92 5e-19
At4g12560 putative protein 91 1e-18
At4g19940 putative protein 84 1e-16
At1g47790 hypothetical protein 81 1e-15
At1g33530 hypothetical protein 80 1e-15
At3g17480 hypothetical protein 80 2e-15
At3g07870 unknown protein 80 2e-15
At2g16810 unknown protein 79 6e-15
At3g10430 hypothetical protein 77 1e-14
At1g11620 hypothetical protein 77 1e-14
At3g17710 unknown protein 77 2e-14
At4g38870 putative protein 77 2e-14
At1g30790 hypothetical protein 75 6e-14
At3g17570 hypothetical protein 74 1e-13
At3g10240 hypothetical protein 73 2e-13
At5g42430 putative protein 72 5e-13
At3g08750 hypothetical protein 71 9e-13
At2g43440 hypothetical protein 71 9e-13
>At3g23880 unknown protein
Length = 364
Score = 151 bits (382), Expect = 5e-37
Identities = 114/364 (31%), Positives = 185/364 (50%), Gaps = 25/364 (6%)
Query: 35 DMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIA 94
++P E++ EILLRLPV+SL +F+CVC W++LIS+ FA KH I S
Sbjct: 13 NLPLEMMEEILLRLPVKSLTRFKCVCSSWRSLISETLFALKHALILETSKATTSTKSPYG 72
Query: 95 KCNLVSYPLKPL----LDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCLSDFY--QFTL 148
Y LK L N S V D E++ ++G+C+GL+C Y L
Sbjct: 73 VITTSRYHLKSCCIHSLYNASTVYVSEHDGELLGRDYYQVVGTCHGLVCFHVDYDKSLYL 132
Query: 149 WNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKTLIY 208
WNP+IKL+ + S + + ++ D + + GFGYD+ D YKV+A++Q + + + +T IY
Sbjct: 133 WNPTIKLQQRLSSSDL--ETSDDECVVTYGFGYDESEDDYKVVALLQQRHQV-KIETKIY 189
Query: 209 TFGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIV----SKKVIVFFDIEKETYGE 264
+ K W + FP D R G+ +++G LNW S I+ +D+ ++ + E
Sbjct: 190 STRQKLWRSNTSFPSGVVVADKSRSGI--YINGTLNWAATSSSSSWTIISYDMSRDEFKE 247
Query: 265 MSLPQDYGDKNTVLYVSSNRIYVSF-DHSNKTHWVVWMMKEYGVVESWTKLMIIPQDKLT 323
+ P G + + R +S + + VW+MKE+G V SW+KL+ I
Sbjct: 248 LPGPVCCGRGCFTMTLGDLRGCLSMVCYCKGANADVWVMKEFGEVYSWSKLLSI------ 301
Query: 324 SPGPYCLSDALFISEHGVLLMRPQHSKLSVYNLNNDGGLDYCTTISGQFARYLHIYNESL 383
PG L+IS+ G++++ S L++YN +N G Y + S R +Y +++
Sbjct: 302 -PGLTDFVRPLWISD-GLVVLLEFRSGLALYNCSN-GRFHYPVSNSISGCRDAKVYLKTM 358
Query: 384 VSPH 387
VSP+
Sbjct: 359 VSPN 362
>At4g21240 putative protein
Length = 417
Score = 92.4 bits (228), Expect = 4e-19
Identities = 76/303 (25%), Positives = 132/303 (43%), Gaps = 55/303 (18%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+P ++I +ILLRLP +S ++FR V KLW ++ + P F + S A+P S
Sbjct: 36 IPLDMIPDILLRLPAKSAVRFRIVSKLWLSITTRPYFIR-----SFAFPS------STRL 84
Query: 96 CNLVSYPLKPLLDNPSAHRVEPADFEMI---------HTTSMTIIGSCNGLLCLSDFYQF 146
C + + + S H+ + + + H S NGL+C DFY
Sbjct: 85 CLMACVKARDMRLFISLHQHDDGSYAHVDRCEIKSPKHDYYNPSSESVNGLVCFGDFYNI 144
Query: 147 TLWNPSIKL-----KSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLD 201
+WNPS++ + KP T+ + F+ GYD V D+YKVL++ + Y+
Sbjct: 145 VVWNPSMRQHVTLPEPKPHSTV--------RYFIRSCLGYDPVEDKYKVLSI--SGYHNG 194
Query: 202 ETKTLIYTFGGKD-WTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIVS-----------K 249
L++T G ++ W IQ P D G VG ++G++ + +
Sbjct: 195 NHDPLVFTLGPQESWRVIQNSPLD-IPLPTGGSRVGTCINGHVYYEAQIRFKVDDIFNFE 253
Query: 250 KVIVFFDIEKETYGEMSLPQDYGDKNTVLYVSSNRIYVSFDHSNKTHWVV-------WMM 302
+++ FD+ E + + P D +N +L + D+S+ WV+ W +
Sbjct: 254 NILMSFDVRYEKFNTIKKPADPTLRNFMLNYQGKLAWFCSDYSSIRFWVLDDGDKQEWSL 313
Query: 303 KEY 305
K +
Sbjct: 314 KNF 316
>At3g16210 hypothetical protein
Length = 360
Score = 92.0 bits (227), Expect = 5e-19
Identities = 85/302 (28%), Positives = 139/302 (45%), Gaps = 43/302 (14%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+PEE+ +EIL+RL ++ L +FRCVCK W+ LI+DP F + + +S A FVS
Sbjct: 5 LPEELAIEILVRLSMKDLARFRCVCKTWRDLINDPGFTETYRDMSPA------KFVSFYD 58
Query: 96 CNLVSYPLKPLLDNPSAHRV------EPADFEMIHTTSMTIIGSCNGLLCLS-DFYQFTL 148
N +LD H V P D MI + T + C+G LC++ + +
Sbjct: 59 KNFY------MLDVEGKHPVITNKLDFPLDQSMIDES--TCVLHCDGTLCVTLKNHTLMV 110
Query: 149 WNP-SIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKTLI 207
WNP S + K P+P I + L GFGYD V+D YKV+ + LD + +
Sbjct: 111 WNPFSKQFKIVPNPGI-----YQDSNIL--GFGYDPVHDDYKVVTFID---RLDVSTAHV 160
Query: 208 YTFGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIV----SKKVIVFFDIEKETYG 263
+ F W + R G F+ L WI + + I+ F++ Y
Sbjct: 161 FEFRTGSWGESLRISYPDWHYRDRR---GTFLDQYLYWIAYRSSADRFILCFNLSTHEYR 217
Query: 264 EMSLP-QDYGDKNTVLYVSSNRIYVSFDHSNKTHWVVWMMKEYGVVESWTKLMIIPQDKL 322
++ LP + G ++ L V+S ++ ++ K + +M++ G SW+K++ +
Sbjct: 218 KLPLPVYNQGVTSSWLGVTSQKLCITEYEMCKKEIRISVMEKTG---SWSKIISLSMSSF 274
Query: 323 TS 324
S
Sbjct: 275 IS 276
>At4g12560 putative protein
Length = 408
Score = 90.5 bits (223), Expect = 1e-18
Identities = 83/352 (23%), Positives = 157/352 (44%), Gaps = 37/352 (10%)
Query: 34 ADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSI 93
A +P +I+ +I LRLP ++L++ R + K LI+DP F + H+ + + +
Sbjct: 2 ATIPMDIVNDIFLRLPAKTLVRCRALSKPCYHLINDPDFIESHLHRVLQTGDHLMILLRG 61
Query: 94 AKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCLSDF-YQFTLWNPS 152
A L S +D S V + M + GS NGL+ LS+ ++NPS
Sbjct: 62 A-LRLYS------VDLDSLDSVSDVEHPMKRGGPTEVFGSSNGLIGLSNSPTDLAVFNPS 114
Query: 153 IK-LKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKTL----- 206
+ + P +I D ++ +++ G GYD V+D YKV+ +VQ + +D L
Sbjct: 115 TRQIHRLPPSSIDLPDGSSTRGYVFYGLGYDSVSDDYKVVRMVQ--FKIDSEDELGCSFP 172
Query: 207 ----IYTFGGKDWTTIQKFPCDPSRCD------LGRLGVGKFVSGNLNWIVSKK------ 250
+++ W I+ L R G G +L+W++ ++
Sbjct: 173 YEVKVFSLKKNSWKRIESVASSIQLLFYFYYHLLYRRGYGVLAGNSLHWVLPRRPGLIAF 232
Query: 251 -VIVFFDIEKETYGEMSLPQDYGDKNTVLYVSSNRI---YVSFDHSNKTHWVVWMMKEYG 306
+IV FD+ E + + P+ + N + + + + ++++ VWMMKEY
Sbjct: 233 NLIVRFDLALEEFEIVRFPEAVANGNVDIQMDIGVLDGCLCLMCNYDQSYVDVWMMKEYN 292
Query: 307 VVESWTKLMIIPQDKLTSPGPYCLSDALFISEHGVLLMRPQHSKLSVYNLNN 358
V +SWTK+ + + K Y + ++ + +L+ ++KL ++L +
Sbjct: 293 VRDSWTKVFTVQKPKSVKSFSY-MRPLVYSKDKKKVLLELNNTKLVWFDLES 343
>At4g19940 putative protein
Length = 411
Score = 84.3 bits (207), Expect = 1e-16
Identities = 94/375 (25%), Positives = 163/375 (43%), Gaps = 45/375 (12%)
Query: 10 KTRRRLRQHSQNVTVFTQSVSEITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISD 69
+ RRR ++ +++ Q + EI P ++++EIL RLP +SL++F+ V KLW +LI
Sbjct: 13 RERRRTKRRRRDLCKSRQPIPEI----PFDLVIEILTRLPAKSLMRFKSVSKLWSSLICS 68
Query: 70 PQFAKKHVSIST---AYPQLVSVFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTT 126
F + + +S+ + L S S K L+S P D + V D M
Sbjct: 69 RNFTNRLLKLSSPPRLFMCLSSSDNSHLKTVLLSLSSPPDSDITMSSSVIDQDLTMPGMK 128
Query: 127 SMTIIGSCNGLLCLSDFYQFTLWNPS----IKLKSKPSPTIIAFDSFDSKRFLYRGFGYD 182
I GL+CL ++N + + L TI+A + SK+ +Y G+D
Sbjct: 129 GYQISHVFRGLMCLVKKSSAQIYNTTTRQLVVLPDIEESTILA-EEHKSKKIMYH-IGHD 186
Query: 183 QVNDRYKVLAVVQNCYNLDETKTLIYTFGGKDWTTI---------QKFPCD-PSRCDLGR 232
V D+YKV+ +V DE + YTF + W + +K C P LG+
Sbjct: 187 PVYDQYKVVCIVSRA--SDEVEE--YTFLSEHWVLLLEGEGSRRWRKISCKYPPHVPLGQ 242
Query: 233 LGVGKFVSGNLNWIV-----SKKVIVFFDIEKETYGEMSLPQDYGDKNTVLYVSSNRIYV 287
G +SG ++++ +V+V FD E + + +P D K L +I +
Sbjct: 243 ---GLTLSGRMHYLAWVRVSDNRVLVIFDTHSEEFSMLQVPGDIFWKYNGLLEYGGKIAI 299
Query: 288 SFDHSNKTHWVVWMMKEYGVVES-----W-TKLMIIPQDKLTSPGPYCLSDALFISEHGV 341
N T + + E VVE W +K++++ +L L + +G
Sbjct: 300 ----LNYTKVDIEGVMELWVVEDEEKNLWSSKILVVNPLQLQMVNSIISLTVLGTTRNGE 355
Query: 342 LLMRPQHSKLSVYNL 356
+++ P +V+N+
Sbjct: 356 VILVPGPEDKTVFNI 370
>At1g47790 hypothetical protein
Length = 389
Score = 80.9 bits (198), Expect = 1e-15
Identities = 59/198 (29%), Positives = 93/198 (46%), Gaps = 21/198 (10%)
Query: 30 SEITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSV 89
S+ T+ P ++ EILLRLPV+S+++FRCV KLW ++I+DP F K + + S+ L+
Sbjct: 19 SKPTSSFPLDLASEILLRLPVKSVVRFRCVSKLWSSIITDPYFIKTYETQSSTRQSLLFC 78
Query: 90 FVSIAKCNLVSYPLKPLLDNPSAHRV-------EPADFEMIHTTSMTIIGSCNGLLCLSD 142
F K + S P N S+ P +F T S +GL+C
Sbjct: 79 FKQSDKLFVFSIPKHHYDSNSSSQAAIDRFQVKLPQEFSYPSPTE-----SVHGLICFHV 133
Query: 143 FYQFTLWNPSIK-LKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLD 201
+WNPS++ + P P S + L GYD + ++KV+ + +N D
Sbjct: 134 LATVIVWNPSMRQFLTLPKPR-------KSWKELTVFLGYDPIEGKHKVVCLPRN-RTCD 185
Query: 202 ETKTLIYTFGGKDWTTIQ 219
E + L K W T++
Sbjct: 186 ECQVLTLGSAQKSWRTVK 203
>At1g33530 hypothetical protein
Length = 441
Score = 80.5 bits (197), Expect = 1e-15
Identities = 82/357 (22%), Positives = 147/357 (40%), Gaps = 46/357 (12%)
Query: 28 SVSEITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLV 87
S + + ++P+ ++ EIL RLPV+ L++ + + K WK+LI A+KH+ + L
Sbjct: 89 SETTLAVELPDVLVEEILQRLPVKYLVRLKSISKGWKSLIESDHLAEKHLRLLEKKYGLK 148
Query: 88 SVFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCL--SDFYQ 145
+ +++ + S +K + + +++ + GSCNGL+C+ D
Sbjct: 149 EIKITVERSTSKSICIKFFSRRSGMNAINSDSDDLLR-----VPGSCNGLVCVYELDSVY 203
Query: 146 FTLWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKT 205
L NP + +P L GFG D V YKV+ + Y D T
Sbjct: 204 IYLLNPMTGVTRTLTP--------PRGTKLSVGFGIDVVTGTYKVMVL----YGFDRVGT 251
Query: 206 LIYTFGGKDWTTIQKFPCD-PSRCDLGRLGVGKFVSGNLNWIVSK--KVIVFFDIEKETY 262
+++ W K P C FV+G+L W+++ I+ D+ E +
Sbjct: 252 VVFDLDTNKWRQRYKTAGPMPLSCIPTPERNPVFVNGSLFWLLASDFSEILVMDLHTEKF 311
Query: 263 GEMSLPQDYGDKNT-----VLYVSSNRIYVSFDHSNKTHWVVWMMKEYGVVESWTKLM-- 315
+S P D D + ++ +R+ VS + H VW++ + + E W +
Sbjct: 312 RTLSQPNDMDDVDVSSGYIYMWSLEDRLCVS-NVRQGLHSYVWVLVQDELSEKWERTRFN 370
Query: 316 ----IIPQDKLTSP-------GPYCLSDALFISEHGVLLMRPQHSKLSVYNLNNDGG 361
+ P L S PY LS + I + Q+S ++++ N D G
Sbjct: 371 LLGHVFPPLSLNSAWFSQTLVSPYQLSSSTCIGSR-----QRQNSTSALFSRNGDTG 422
>At3g17480 hypothetical protein
Length = 373
Score = 79.7 bits (195), Expect = 2e-15
Identities = 75/304 (24%), Positives = 128/304 (41%), Gaps = 38/304 (12%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+ E+++ +IL R+P SL++ R CK W +++D +F KKH TA + + + + +
Sbjct: 11 LTEDLVEDILSRVPATSLVRLRSTCKQWNAILNDRRFIKKH--FDTAEKEYLDMLLRSLR 68
Query: 96 CNLVSYPLKPLLDN--PSAHRVEPADFEMIHTTSMTI----IGSCNGLLCLSDFYQFTLW 149
+ +S L L DN PS + + + S + + CNGLL ++ +W
Sbjct: 69 VSSMSVNLHGLHDNIDPSIELKRELNLKGLLCNSGKVESFEVFHCNGLLLFTNTSTIVVW 128
Query: 150 NPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVN--DRYKVLAVVQNCYNLDETKTLI 207
NP I +S +++ Y GY+ N YK+L + + N + I
Sbjct: 129 NP-----CTGQTKWIQTESANTRYHKY-ALGYENKNLCRDYKILRFLDDGTNFE---LEI 179
Query: 208 YTFGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIVSKK----------VIVFFDI 257
Y F W + D D+G G+ V GN WIV + FD
Sbjct: 180 YEFNSSSWRVLDSVEID-FELDIGSQGMS--VKGNTYWIVIDDHEVDGELVNCFLTSFDF 236
Query: 258 EKETYG-EMSLPQD---YGDKNTVLYVSSNRIYVSFDHSNKTHWVVWMMKEY--GVVESW 311
+E +G + LP Y D ++ V R+ V F + +W+ + +V SW
Sbjct: 237 TRERFGPHVCLPFQTSWYSDVISLSSVREERLSVLFHEEDSLTMEIWVTNDITDTIVTSW 296
Query: 312 TKLM 315
+K +
Sbjct: 297 SKFL 300
>At3g07870 unknown protein
Length = 417
Score = 79.7 bits (195), Expect = 2e-15
Identities = 77/313 (24%), Positives = 137/313 (43%), Gaps = 51/313 (16%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+PE+II +I RLP+ S+ + VC+ W+++++ +H +S++ + +
Sbjct: 28 LPEDIIADIFSRLPISSIARLMFVCRSWRSVLT------QHGRLSSSSSSPTKPCLLLHC 81
Query: 96 CNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSM---TIIGSCNGLLCLSDFYQFTLWNPS 152
+ + L L + R++ F + +SM ++GSCNGLLCLSD +L+N S
Sbjct: 82 DSPIRNGLHFLDLSEEEKRIKTKKFTLRFASSMPEFDVVGSCNGLLCLSD----SLYNDS 137
Query: 153 IKLKSKPSPTIIAFDSFDSK---RFLYRGFGYDQVNDRYKVLAVV--------------- 194
+ L + + + +K + L GFG+ ++ YKVL +V
Sbjct: 138 LYLYNPFTTNSLELPECSNKYHDQELVFGFGFHEMTKEYKVLKIVYFRGSSSNNNGIYRG 197
Query: 195 QNCYNLDETKTLIYTFGGK------DWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWI-- 246
+ +++ I T K W ++ K P + V+G L+++
Sbjct: 198 RGRIQYKQSEVQILTLSSKTTDQSLSWRSLGKAP-----YKFVKRSSEALVNGRLHFVTR 252
Query: 247 ----VSKKVIVFFDIEKETYGEMSLPQDYGDKNTVLY--VSSNRIYVSFDHSNKTHWVVW 300
V + V FD+E E + E+ P D G N + V+ + + N +W
Sbjct: 253 PRRHVPDRKFVSFDLEDEEFKEIPKP-DCGGLNRTNHRLVNLKGCLCAVVYGNYGKLDIW 311
Query: 301 MMKEYGVVESWTK 313
+MK YGV ESW K
Sbjct: 312 VMKTYGVKESWGK 324
>At2g16810 unknown protein
Length = 295
Score = 78.6 bits (192), Expect = 6e-15
Identities = 76/283 (26%), Positives = 129/283 (44%), Gaps = 40/283 (14%)
Query: 8 RSKTRRRLRQHSQNVTVFTQSVSEITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLI 67
RS++RRR +Q + + + +P+++++EIL RLP +S+++F+CV K W +L+
Sbjct: 22 RSRSRRRAKQDPK---------TWVARYIPQDLLIEILTRLPPKSVMRFKCVSKFWSSLL 72
Query: 68 SDPQFAKKHVSISTAYPQLVSVFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTS 127
S F + + I + PQ S+ C L Y L SA P F +
Sbjct: 73 SSRYFCNRFL-IVPSQPQ-----PSLYMCLLDRYNYSKSLILSSAPSTSPYSFVFDQDLT 126
Query: 128 MTIIGS-----CNGLLCLSDFYQFTLWNPSIK----LKSKPSPTIIAFDSFDSKRFLYRG 178
+ +G G + + + ++NP+ + L + IIA ++ F+
Sbjct: 127 IRKMGGFFLRILRGFIFFTRNLKARIYNPTTRQLVILPTIKESDIIAGPPYNILYFIC-- 184
Query: 179 FGYDQVNDRYKVLAVVQ--NCYNLDETKTLIYTF---GGKDWTTI-QKFPCD-PSRCDLG 231
+D VNDRYK+L V + +L K+ ++ F G W + +FP PS DL
Sbjct: 185 --HDPVNDRYKLLCTVSYASDNDLQNLKSELWIFVLEAGGSWKRVANEFPHHVPSHLDLN 242
Query: 232 RLGVGKFVSGNLNWIVSKK-VIVFFDIEKETYGEMSLPQDYGD 273
GV F L W ++V FD+ E + M +P++ GD
Sbjct: 243 MNGVLYF----LAWTDPHTCMLVSFDVRSEEFNTMQVPRNAGD 281
>At3g10430 hypothetical protein
Length = 370
Score = 77.4 bits (189), Expect = 1e-14
Identities = 77/252 (30%), Positives = 115/252 (45%), Gaps = 45/252 (17%)
Query: 34 ADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLIS-DPQFAKKHVSIST------AYPQL 86
+ +P ++I+EIL R P SLL+F+ CK W LIS D +F KH+ ST +
Sbjct: 3 SSLPFDLILEILQRTPAESLLRFKSTCKKWYELISNDKRFMYKHLDKSTKRFLRIENRER 62
Query: 87 VSVFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCLSDFYQF 146
V + + + VS + N H+ F +IH + ++G C L
Sbjct: 63 VQILDPVTEILAVS-----TIPNELRHKY----FTLIHCDGL-MLGMCYEE--LGSDPNL 110
Query: 147 TLWNPSI-KLK-SKPSPTIIAFDSFDSKRFLYRGFGYDQV-NDRYKVLAVVQNCYNLDE- 202
+WNP + K+K KPSP ++ + D Y GFGYD+ D YK+L + D+
Sbjct: 111 AVWNPVMRKIKWIKPSPPLVCYWGSD-----YLGFGYDKTFRDNYKILRFTYLGDDDDDE 165
Query: 203 --TKTLIYTFGGKDWTTIQ-KFPCDPSRCDLGRLGV---GKFVSGNLNWI---VSKKVIV 253
K IY F W +I+ KF G + V G V+G++ WI K I+
Sbjct: 166 SYPKCQIYEFNSGSWRSIEAKFD--------GEIDVEVDGVSVNGSMYWIELQEKKNFIL 217
Query: 254 FFDIEKETYGEM 265
FD KET+ +
Sbjct: 218 SFDFSKETFNRI 229
>At1g11620 hypothetical protein
Length = 363
Score = 77.4 bits (189), Expect = 1e-14
Identities = 91/350 (26%), Positives = 149/350 (42%), Gaps = 55/350 (15%)
Query: 33 TADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVS 92
T D+ +++ EIL R+P RSL++ R CK W+ LI++P+F KH+S Q +VF +
Sbjct: 3 TMDLSSDLVEEILSRVPARSLVRLRSTCKQWEALIAEPRFVNKHLSHMRYREQQFTVFNN 62
Query: 93 IAKCNLVSYPLKPLLDNPSAH------RVEPADFEM---IHTTSMTIIGSCNGLLCLSDF 143
+ + PL + +++ + E ++ I + I C+GLL
Sbjct: 63 -------EHIVSPLFGSTTSYVGIDFNKPENCGVKLPFPIALSPAINISHCDGLLLYVTK 115
Query: 144 YQFTLWNPSIKLKSKPSPTIIAFDSFDSKRFLY-RGFGYDQVNDRYKVLAVVQNCYNLDE 202
+ NP + K I + FD Y G+ ++Q + Y V C +
Sbjct: 116 SMLLVANPLLSQKR----WIKCSEGFDHSMDAYGLGYLFNQSSGFYDYKVVRFRCGIKNS 171
Query: 203 TKTLIYTFGGKDW-----TTIQKFPCDP--SRCDLGR---LGVGKFVSGNLNWIVSKKVI 252
++ +Y F W T F P S C G LG K SGN I
Sbjct: 172 SRVEVYAFKSDSWKVVVDTNFGGFDGLPLSSVCLRGTPYWLGYNK--SGN-----ELMSI 224
Query: 253 VFFDIEKETYGEMSL-PQDYGDKNTVLYVS-----SNRIYVSFDHSNKTHWVVWMMKEYG 306
FD KE + + L PQ G +N V Y+S +++ + + +W+MK+
Sbjct: 225 QSFDFSKERFEPLFLPPQSIGSRNLVKYISLGIFRGDQLSLLLECHETCKLHLWVMKK-- 282
Query: 307 VVESWTKLMI--IPQDKLTSPGPYCLSDALFISEHGVLLMRPQHSKLSVY 354
+ W++LM +PQD + G Y S FI +G L + + +S+Y
Sbjct: 283 --QHWSRLMTVDVPQDAIY--GKYFSS---FIERNGRLALLIKSRNISIY 325
>At3g17710 unknown protein
Length = 368
Score = 77.0 bits (188), Expect = 2e-14
Identities = 90/333 (27%), Positives = 144/333 (43%), Gaps = 46/333 (13%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+P ++ EIL RLP RSL++FR VCK W L SD +F KKH + A PQ +F++ +K
Sbjct: 6 LPWDLEEEILSRLPPRSLVRFRTVCKHWNGLFSDKRFVKKH--LVRARPQF--IFLTESK 61
Query: 96 CNLVSYPLKPLLDNPSAHRVEPADFE-MIHTTSMTIIGSCNGLLCLSDFYQ--FTLWNPS 152
Y ++ L R P DF + T I +C+GLL DF++ +WNP
Sbjct: 62 ---KMYSIEIDLGGTIEVREVPYDFHCQPMKKNFTTIMACDGLL-FRDFWKQGVAVWNPW 117
Query: 153 IKLKSKPSPTIIAFDSFDSKRFLYRGFGYD--QVNDRYKVLAVVQNCYNLDET------K 204
++ + + ++ K F + G GYD + + YK+L L ++ +
Sbjct: 118 LRQ--------VGWIEYEDKGFRFCGVGYDSCKPDKCYKILGYFNCTRRLSDSLQEGYYQ 169
Query: 205 TLIYTFGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIVS------KKVIVFFDIE 258
IY + + KF P+ +L V+GNL W+ + I FD
Sbjct: 170 AAIYECASQAF----KFIDTPNPFNLWPNKDPLSVNGNLYWLAHNHPETLEYFIETFDFS 225
Query: 259 KETYGEMSL---PQDYGDKNTVLYVSS----NRIYVSFDHSNKTHWVVWMMKEYGVVESW 311
E + L +D+G VL V + + F+ + WV M + W
Sbjct: 226 MEIFKPFCLLPCRKDFGSNELVLAVFKEDRFSLLKQCFETTKIEIWVTKMKIDREEEVVW 285
Query: 312 TKLMIIPQDKLTS-PGPYCLSDALFISEHGVLL 343
K M +P L + YC S + FI + +++
Sbjct: 286 IKFMTLPTTNLPNLDDTYCCS-SYFIFDKTIIM 317
>At4g38870 putative protein
Length = 426
Score = 76.6 bits (187), Expect = 2e-14
Identities = 74/297 (24%), Positives = 131/297 (43%), Gaps = 30/297 (10%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQ-LVSVFVSIA 94
+P ++I+EIL +L ++ L++F CV KLW ++I DP F K ++ S P+ LV VF + +
Sbjct: 53 LPVDLIMEILKKLSLKPLIRFLCVSKLWASIIRDPYFMKLFLNESLKRPKSLVFVFRAQS 112
Query: 95 KCNLV-SYPLKPLLDNPSAHRVEPADFEMIHTT-----SMTIIGSCNGLLCLSDFYQFTL 148
++ S LK + S+ A H T MTI S +GL+C +
Sbjct: 113 LGSIFSSVHLKSTREISSSSSSSSASSITYHVTCYTQQRMTISPSVHGLICYGPPSSLVI 172
Query: 149 WNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKTL-- 206
+NP + +S P I A +R + + GYD ++ YKV+ + + L + L
Sbjct: 173 YNPCTR-RSITLPKIKA-----GRRAINQYIGYDPLDGNYKVVCITRGMPMLRNRRGLAE 226
Query: 207 ---IYTFGGKD--WTTIQKF-----PCDPSRCDLGRLGVGKFVSGNLNWIVSKKVIVFFD 256
+ T G +D W I P C G L F+ LN + I+ FD
Sbjct: 227 EIQVLTLGTRDSSWRMIHDIIPPHSPVSEELCINGVLYYRAFIGTKLN----ESAIMSFD 282
Query: 257 IEKETYGEMSLPQDYGDKNTVLYVSSNRIYVSFDHSNKTHWVVWMMKEYGVVESWTK 313
+ E + + +P ++ + + + ++ +W++++ E W+K
Sbjct: 283 VRSEKFDLIKVPCNFRSFSKLAKYEGKLAVIFYEKKTSGIIGLWILEDASNGE-WSK 338
>At1g30790 hypothetical protein
Length = 399
Score = 75.1 bits (183), Expect = 6e-14
Identities = 73/263 (27%), Positives = 111/263 (41%), Gaps = 42/263 (15%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+P ++ VEIL RLP +SL++F+CV KLW ++I + F SIS+ P+ + V+ +
Sbjct: 9 IPFDLTVEILTRLPAKSLMKFKCVSKLWSSIIHNQSFIDSFYSISSTRPRFI---VAFSN 65
Query: 96 CNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIG------------SCNGLLCLSDF 143
+ S K L S+H + +I TI S NG + S +
Sbjct: 66 GSFPSDKEKRLFIFSSSHEGHESSSSVITNLDTTIPSLTVSNNLASRCISVNGFIACSLY 125
Query: 144 YQFTLWNPSIKLKSKPSPTIIAFDSFDSKR---FLYRGFGYDQVNDRYKVLAVVQNCY-N 199
+FT+ NPS + +I S R GYD V+D++K LA++ +C N
Sbjct: 126 TRFTICNPSTR-------QVIVLPILPSGRAPDMRSTCIGYDPVDDQFKALALISSCIPN 178
Query: 200 LDET-KTLIYTFGGK----DWTTIQ-------KFPCDPSRCDLGRLGVGKFVSGNLNWIV 247
D T + L+ T G W IQ P C G + G +
Sbjct: 179 KDSTVEHLVLTLKGDKKNYSWRQIQGNNNIPPYSPVTMRVCINGVVYYGAWTPRQ----S 234
Query: 248 SKKVIVFFDIEKETYGEMSLPQD 270
VIV FD+ E + P+D
Sbjct: 235 MNAVIVCFDVRSEKITFIKTPKD 257
>At3g17570 hypothetical protein
Length = 381
Score = 73.9 bits (180), Expect = 1e-13
Identities = 85/320 (26%), Positives = 138/320 (42%), Gaps = 59/320 (18%)
Query: 35 DMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVS------ISTAYPQLVS 88
D+P ++ EIL R+P SL + + CK W TL DP+F KKHV IS ++ S
Sbjct: 4 DLPRDLETEILSRVPATSLQKLKPTCKRWYTLFKDPEFLKKHVGRAEREVISLMSLRVYS 63
Query: 89 VFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNG-LLCLSDFYQFT 147
+ V+++ + S + +L++ D E + + +T CNG LLC +D +
Sbjct: 64 LSVNLSGIH-SSVEMTGMLNSLK-------DSEDVKISDIT---ECNGLLLCTTDDSRLV 112
Query: 148 LWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGF--GYDQVND---RYKVLAVVQNCYN--L 200
+WNP I + S +S +Y+ F GYD N YK+L CY+ +
Sbjct: 113 VWNP-----YTGETRWIPYKS-NSPYEMYQKFVLGYDNTNKSRYSYKIL----RCYHGLI 162
Query: 201 D-ETKTLIYTFGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIVS----KKVIVFF 255
D + IY F W ++F + C GV + GN W S + +I+ F
Sbjct: 163 DFGYEFEIYEFNSHSW---RRFYDNSPNCSFESKGV--TLKGNTYWFASDTEGRHIILRF 217
Query: 256 DIEKETYGEMSLPQDYG-------DKNTVLYVSSNR---IYVSFDHSNKTHWVVWMMKEY 305
D E +G +SL G + + V + +Y FD N + + +
Sbjct: 218 DFATERFGRLSLLYQSGGYVDNVVETGVLSAVREEKLALLYERFDELNDSSEMKIWVTNT 277
Query: 306 GVVE----SWTKLMIIPQDK 321
+VE SW+ +++ K
Sbjct: 278 KIVEAKDLSWSDFLVVDSSK 297
>At3g10240 hypothetical protein
Length = 389
Score = 73.2 bits (178), Expect = 2e-13
Identities = 72/296 (24%), Positives = 125/296 (41%), Gaps = 48/296 (16%)
Query: 33 TADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVS 92
++ +P +++ EILLRLP +S+ +FRCV K W ++ ++P F ++T P+L+ F +
Sbjct: 24 SSSIPLDLVSEILLRLPEKSVARFRCVSKPWSSITTEPYFIN---LLTTRSPRLLLCFKA 80
Query: 93 IAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTI----------------IGSCNG 136
K + S P HR + H+ S I S NG
Sbjct: 81 NEKFFVSSIP---------QHRQTFETWNKSHSYSQLIDRYHMEFSEEMNYFPPTESVNG 131
Query: 137 LLCLSDFYQFTLWNPSIK-LKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQ 195
L+C + + +WNPS + L P P +S D FL GYD V ++KV+ ++
Sbjct: 132 LICFQESARLIVWNPSTRQLLILPKPN---GNSNDLTIFL----GYDPVEGKHKVMC-ME 183
Query: 196 NCYNLDETKTLIYTFGGKDWTTIQKFP------CDPSRCDLGRLGVGKFVSGNLNWIVSK 249
D + L K W T++ D RC G + +V W
Sbjct: 184 FSATYDTCRVLTLGSAQKLWRTVKTHNKHRSDYYDSGRCINGVVYHIAYVKDMCVW---- 239
Query: 250 KVIVFFDIEKETYGEMSLPQDYGDKNTVLYVSSNRIYVSFDHSNKTHWVVWMMKEY 305
V++ FD+ E + + LP K+ ++ + V + K +W+++++
Sbjct: 240 -VLMSFDVRSEIFDMIELPSSDVHKDVLIDYNGRLACVGREIIEKNGIRLWILEKH 294
>At5g42430 putative protein
Length = 395
Score = 72.0 bits (175), Expect = 5e-13
Identities = 72/269 (26%), Positives = 130/269 (47%), Gaps = 26/269 (9%)
Query: 31 EITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVF 90
E + +P ++I+EIL RLP +S+ +F CV KLW +++ P F + ++IS+A P+L+ F
Sbjct: 5 ENSDSIPIDLILEILSRLPAKSITRFHCVSKLWGSMLCRPYFNELFLTISSARPRLLFAF 64
Query: 91 VSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCLSDFY---QFT 147
+ S P +P + V ADF + ++ I +GL+ S +
Sbjct: 65 SKHGEWRFFSSP-QPQNPYGKSSFVATADFHTKFSQNLNICNYTSGLVYFSAMWITKADV 123
Query: 148 LWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKTL- 206
+ NPS + ++ + +++ F F +D V ++KVL + N N +ETK +
Sbjct: 124 ICNPSTGHYAMLPKLLLTYG--ETRSF----FLFDPVGKQFKVL--LMNKINNNETKDIH 175
Query: 207 IYTFGGKD--WTTIQKFP-----CDPSRCDLGRLGVGKFVSGNLNWIVSKKVIVFFDIEK 259
I T G + W IQ+ P C G L +++ N++ + IV FD+
Sbjct: 176 ILTLGTRKVRWRKIQQCPLIHIVSHEWICINGAL---YYIAYNIDDFLG--YIVCFDVRS 230
Query: 260 ETYGEMSLPQD-YGDKNTVLYVSSNRIYV 287
E + ++L QD + +++T L ++ V
Sbjct: 231 EKFKCLNLNQDCFSERSTKLIYYKGKLGV 259
>At3g08750 hypothetical protein
Length = 369
Score = 71.2 bits (173), Expect = 9e-13
Identities = 71/250 (28%), Positives = 105/250 (41%), Gaps = 51/250 (20%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVS-------ISTAYPQLVS 88
+P E+I EIL ++P SL++F+ CK W LI++ +F H+ I T Q++
Sbjct: 12 LPFELIEEILYKIPAESLIRFKSTCKKWYNLITEKRFMYNHLDHYSPERFIRTYDQQIID 71
Query: 89 VFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLL---CLSDFYQ 145
I L+ P R + M+H C+GL+ C
Sbjct: 72 PVTEILSDALI----------PDEFRDLYPIYSMVH---------CDGLMLCTCRKWDNS 112
Query: 146 FTLWNPSIK-LK-SKPSPTIIAFDSFDSKRFLYRGFGYDQ--VNDRYKVLAVVQNCYNLD 201
+WNP ++ +K KPS + D Y G GYD D YK+L ++ D
Sbjct: 113 LAVWNPVLREIKWIKPSVCYLHTD--------YVGIGYDDNVSRDNYKILKLLGRLPKDD 164
Query: 202 ET--KTLIYTFGGKDW-TTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIVSKK---VIVFF 255
++ IY F W T + KF D D+ R G V G + WI KK I+ F
Sbjct: 165 DSDPNCEIYEFKSDSWKTLVAKFDWD---IDI-RCNNGVSVKGKMYWIAKKKEDFTIIRF 220
Query: 256 DIEKETYGEM 265
D ET+ E+
Sbjct: 221 DFSTETFKEI 230
>At2g43440 hypothetical protein
Length = 779
Score = 71.2 bits (173), Expect = 9e-13
Identities = 78/339 (23%), Positives = 150/339 (44%), Gaps = 43/339 (12%)
Query: 39 EIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAKCNL 98
E++ +I LRLP++S+L+F+ V + W++++ F ++ ++ + ++++ + CN
Sbjct: 16 ELLEDIFLRLPLKSILKFKTVSRQWRSILESKLFVERRGNLQKHHRKILAAY----NCN- 70
Query: 99 VSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCLSDFYQFTLWNPSI----K 154
Y ++P + P + + +H + +C+GLLC+++ F + NPS +
Sbjct: 71 --YFMRPSI-FPESRFEGDEEIVYLHCDAAQPSMTCDGLLCITEPGWFNVLNPSAGQLRR 127
Query: 155 LKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKTLIYTFGGKD 214
P P +++ GFG D V RYK +V+ C++ Y FG D
Sbjct: 128 FPPGPGPVKGPQENW------LLGFGRDNVTGRYK---IVRMCFH------DCYEFGILD 172
Query: 215 WTT--IQKFPCDPSRCDLGRLGVGKFVSGNLNW--IVSKKVIVFFDIEKETY-GEMSLPQ 269
T K P G V V+G++ W I + +I+ D+ +E+Y G LP
Sbjct: 173 IETGVWSKLRSPPHNMLPGSKSV--CVNGSIYWLQISAGYIILAMDLHEESYHGVHHLPA 230
Query: 270 DYGDKNTVLYVSSNRIYVSFDHSNKTHWV--VWMM----KEYGVVESWTK---LMIIPQD 320
+ + T L +R+ ++ + W+ +W M K + SW+K + +
Sbjct: 231 TWVTQETQLVNLEDRLAMAMTTNVGPEWILEIWSMDIEGKGWSKGYSWSKSYSISLAHGV 290
Query: 321 KLTSPGPYCLSDALFISEHGVLLMRPQHSKLSVYNLNND 359
+ P +++S+ G LL H +L Y D
Sbjct: 291 GFSWPWQKRWFTPVWVSKQGNLLFYDNHKRLFKYYSGTD 329
Score = 62.8 bits (151), Expect = 3e-10
Identities = 71/334 (21%), Positives = 147/334 (43%), Gaps = 47/334 (14%)
Query: 39 EIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAKCNL 98
E++ EI L LP++S+L+F+ V K W++++ F ++ ++ +P++++ + C+
Sbjct: 402 ELLEEIFLGLPLKSILKFKTVSKQWRSILESNLFVERRRTLQKNHPKILAAY----NCDY 457
Query: 99 VSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCLSDFYQFTLWNPSIKLKSK 158
+ P +L P + + +HT + +C+GL+C+++ F + N S
Sbjct: 458 CTRP--GIL--PKSQFEGDEEIVYLHTDATQPSMTCDGLVCITEPGWFNVLNVS------ 507
Query: 159 PSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAV-VQNCYNLDETKTLIYTFGGKDWTT 217
+ +RFL G D+V +YK++ + +CY I +W+
Sbjct: 508 ---------TGQLRRFL-PGPDPDKVTGKYKIVRMCFHDCYEFG-----ILDIESGEWS- 551
Query: 218 IQKFPCDPSRCDLGRLGVGKFVSGNLNW--IVSKKVIVFFDIEKETY-GEMSLPQDYGDK 274
K P +G V V+G++ W I +I+ D+ +ET+ G LP + +
Sbjct: 552 --KLMSPPHIMRVGSKSV--CVNGSIYWLQISVSYIILALDLHQETFNGVYHLPATWVTQ 607
Query: 275 NTVLYVSSNRIYVSFDHSNKTHWV--VWMM----KEYGVVESWTK---LMIIPQDKLTSP 325
+T L +R+ ++ W+ +W M K + +W+K + + + ++ P
Sbjct: 608 DTQLVNLEDRLAMAMTTKVGPEWILEIWSMDIEEKGWSKRYTWSKAYSISLAHRVVVSWP 667
Query: 326 GPYCLSDALFISEHGVLLMRPQHSKLSVYNLNND 359
+ +S+ G L+ H +L Y D
Sbjct: 668 WQKRWFTPVSVSKQGNLVFYDNHKRLFKYYSGTD 701
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.138 0.430
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,222,274
Number of Sequences: 26719
Number of extensions: 403549
Number of successful extensions: 1338
Number of sequences better than 10.0: 296
Number of HSP's better than 10.0 without gapping: 253
Number of HSP's successfully gapped in prelim test: 43
Number of HSP's that attempted gapping in prelim test: 933
Number of HSP's gapped (non-prelim): 335
length of query: 388
length of database: 11,318,596
effective HSP length: 101
effective length of query: 287
effective length of database: 8,619,977
effective search space: 2473933399
effective search space used: 2473933399
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146776.6


