

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146760.13 + phase: 0 /pseudo/partial
(612 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
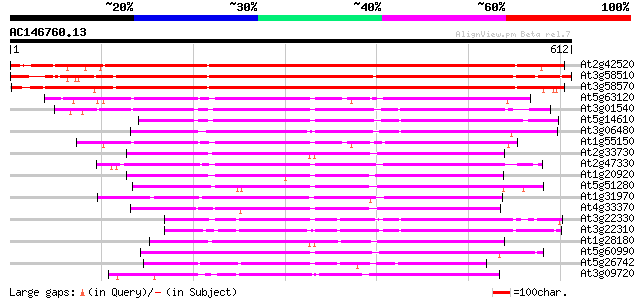
Score E
Sequences producing significant alignments: (bits) Value
At2g42520 putative ATP-dependent RNA helicase 902 0.0
At3g58510 ATP-dependent RNA helicase-like protein 871 0.0
At3g58570 ATP-dependent RNA helicase-like protein 858 0.0
At5g63120 ATP-dependent RNA helicase-like protein 383 e-106
At3g01540 AT3g01540/F4P13_9 343 2e-94
At5g14610 DRH1 DEAD box protein - like 338 4e-93
At3g06480 putative RNA helicase 334 1e-91
At1g55150 unknown protein 330 1e-90
At2g33730 putative U5 small nuclear ribonucleoprotein, an RNA he... 313 2e-85
At2g47330 putative ATP-dependent RNA helicase 305 4e-83
At1g20920 putative RNA helicase 295 7e-80
At5g51280 DEAD-box protein abstrakt 287 1e-77
At1g31970 unknown protein 280 2e-75
At4g33370 putative protein 278 5e-75
At3g22330 DEAD-Box RNA helicase like protein 275 7e-74
At3g22310 putative RNA helicase 272 5e-73
At1g28180 hypothetical protein 246 2e-65
At5g60990 replication protein A1 - like 234 1e-61
At5g26742 unknown protein 233 3e-61
At3g09720 RNA helicase like protein 223 2e-58
>At2g42520 putative ATP-dependent RNA helicase
Length = 633
Score = 902 bits (2331), Expect = 0.0
Identities = 473/635 (74%), Positives = 516/635 (80%), Gaps = 43/635 (6%)
Query: 1 MRSSWADLAANSAAENAPTPPPSAAAAVANNAANTRPVYVPPHLRNRGPSAPAPAAPAFD 60
M +SWAD+A + EN + + + N+ +RP YVPPHLRNR P+A P AP
Sbjct: 1 MSASWADVADS---EN------TGSGSSNQNSHPSRPAYVPPHLRNR-PAASEPVAPLPA 50
Query: 61 N---------SGSRWAPPPRNDYRGGGGGR--------GGGGYGNRGGGGWDRR------ 97
N SGSRWAP GGGGG G GYG RGGGGW+ R
Sbjct: 51 NDRVGYGGPPSGSRWAPGGSGVGVGGGGGYRADAGRPGSGSGYGGRGGGGWNNRSGGWDR 110
Query: 98 ---EANPFADQDDSEEPVTQEEQENTGINFDAYEDIPVETSGGNVPPPVNTFAEIDLGEA 154
E NPF + DDSE EQ+NT INFDAYEDIP+ETSG NVPPPVNTFAEIDLGEA
Sbjct: 111 REREVNPF-ENDDSEPEPAFTEQDNTVINFDAYEDIPIETSGDNVPPPVNTFAEIDLGEA 169
Query: 155 LNQNIRRCKYVKPTPVQRHAIPISLGGRDLMACAQTGSGKTAAFCFPIISGIMTGQPAQR 214
LN NIRRCKYVKPTPVQRHAIPI L GRDLMACAQTGSGKTAAFCFPIISGIM Q QR
Sbjct: 170 LNLNIRRCKYVKPTPVQRHAIPILLEGRDLMACAQTGSGKTAAFCFPIISGIMKDQHVQR 229
Query: 215 PPRGVRTVCPLALVLSPTRELSMQIHEEARKFSYQTGVRVVVAYGGAPINQQLRELERGV 274
P RG RTV PLA++LSPTREL+ QIH+EA+KFSYQTGV+VVVAYGG PINQQLRELERGV
Sbjct: 230 P-RGSRTVYPLAVILSPTRELASQIHDEAKKFSYQTGVKVVVAYGGTPINQQLRELERGV 288
Query: 275 DILVATPGRLVDLLERARVSLSMIRYLALDEADRMLDMGFEPQIRKIVEQMDMPPAGVRQ 334
DILVATPGRL DLLERARVS+ MIR+LALDEADRMLDMGFEPQIRKIVEQMDMPP GVRQ
Sbjct: 289 DILVATPGRLNDLLERARVSMQMIRFLALDEADRMLDMGFEPQIRKIVEQMDMPPRGVRQ 348
Query: 335 TMLFSATFPKEIQRLASDFLSNYIFLAVGRVGSSTDLIDQRVEYVQESDKRSHLMDLLHA 394
T+LFSATFP+EIQRLA+DFL+NYIFLAVGRVGSSTDLI QRVE+V +SDKRSHLMDLLHA
Sbjct: 349 TLLFSATFPREIQRLAADFLANYIFLAVGRVGSSTDLIVQRVEFVLDSDKRSHLMDLLHA 408
Query: 395 QRANGVQGKQALTLVFVETKKGADALEHWLCLNNFPATTIHGDRSQQEREAALRSFKSGN 454
QR NG+QGKQALTLVFVETK+GAD+LE+WLC+N FPAT+IHGDR+QQERE AL++FKSG
Sbjct: 409 QRENGIQGKQALTLVFVETKRGADSLENWLCINGFPATSIHGDRTQQEREVALKAFKSGR 468
Query: 455 TPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRAGKKGLATAFFNENNTS 514
TPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRAGK GLATAFFN+ NTS
Sbjct: 469 TPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRAGKSGLATAFFNDGNTS 528
Query: 515 MARSLQDLMQEANQEVPAWLSRFAARSSFGGGKNRRSGGGRFGGRDFRREGSFSRGGSDY 574
+AR L +LMQEANQEVP WL+R+A+RSSFGGGKNRRS GGRFGGRDFRREGSF G Y
Sbjct: 529 LARPLAELMQEANQEVPEWLTRYASRSSFGGGKNRRS-GGRFGGRDFRREGSFGSGRGGY 587
Query: 575 HGAG---NSGGGYGASGGY-GGGGYGGGYAGNSAG 605
G G GGGYG GGY GGGGYGGGY G S+G
Sbjct: 588 GGGGGGYGGGGGYGGGGGYGGGGGYGGGYGGASSG 622
Score = 28.9 bits (63), Expect = 8.8
Identities = 11/22 (50%), Positives = 11/22 (50%)
Query: 74 YRGGGGGRGGGGYGNRGGGGWD 95
Y GG GG GGYG WD
Sbjct: 612 YGGGYGGASSGGYGGEPPSAWD 633
>At3g58510 ATP-dependent RNA helicase-like protein
Length = 612
Score = 871 bits (2250), Expect = 0.0
Identities = 463/641 (72%), Positives = 505/641 (78%), Gaps = 59/641 (9%)
Query: 1 MRSSWADLAANSAAENAPTPPPSAAAAVANNAANTRPVYVPPHLRNRGPSAPAPAAPAFD 60
M +SWAD+A + A + PP YVPPHLRNR PS P AAP
Sbjct: 1 MSASWADVADSEKAVSQSKPP-----------------YVPPHLRNR-PSEPV-AAPLPQ 41
Query: 61 N---------SGSRWAPPP-----------RND-------YRGGGGGRGGGGYGNRGGGG 93
N +GSRWAPP RND Y GGGG GGGG+ NR GG
Sbjct: 42 NDHAGYGGQPAGSRWAPPSSGGGGASGGGYRNDGGRTGYGYGAGGGGGGGGGWNNRSGG- 100
Query: 94 WDRRE--ANPFADQDDSEEPVTQEEQENTGINFDAYEDIPVETSGGNVPPPVNTFAEIDL 151
WDRRE NPF D + E T EQENTGINFDAYEDIPVETSGG+VPPPVNTFA+IDL
Sbjct: 101 WDRREREVNPFGDDAELEPVFT--EQENTGINFDAYEDIPVETSGGDVPPPVNTFADIDL 158
Query: 152 GEALNQNIRRCKYVKPTPVQRHAIPISLGGRDLMACAQTGSGKTAAFCFPIISGIMTGQP 211
G+ALN NIRRCKYV+PTPVQRHAIPI L RDLMACAQTGSGKTAAFCFPIISGIM Q
Sbjct: 159 GDALNLNIRRCKYVRPTPVQRHAIPILLAERDLMACAQTGSGKTAAFCFPIISGIMKDQH 218
Query: 212 AQRPPRGVRTVCPLALVLSPTRELSMQIHEEARKFSYQTGVRVVVAYGGAPINQQLRELE 271
+RP RG R V P A++LSPTREL+ QIH+EA+KFSYQTGV+VVVAYGG PI+QQLRELE
Sbjct: 219 VERP-RGSRAVYPFAVILSPTRELACQIHDEAKKFSYQTGVKVVVAYGGTPIHQQLRELE 277
Query: 272 RGVDILVATPGRLVDLLERARVSLSMIRYLALDEADRMLDMGFEPQIRKIVEQMDMPPAG 331
RG DILVATPGRL DLLERARVS+ MIR+LALDEADRMLDMGFEPQIRKIVEQMDMPP G
Sbjct: 278 RGCDILVATPGRLNDLLERARVSMQMIRFLALDEADRMLDMGFEPQIRKIVEQMDMPPRG 337
Query: 332 VRQTMLFSATFPKEIQRLASDFLSNYIFLAVGRVGSSTDLIDQRVEYVQESDKRSHLMDL 391
VRQTMLFSATFP +IQRLA+DF+SNYIFLAVGRVGSSTDLI QRVE+VQESDKRSHLMDL
Sbjct: 338 VRQTMLFSATFPSQIQRLAADFMSNYIFLAVGRVGSSTDLITQRVEFVQESDKRSHLMDL 397
Query: 392 LHAQRANGVQGKQALTLVFVETKKGADALEHWLCLNNFPATTIHGDRSQQEREAALRSFK 451
LHAQR Q KQ+LTLVFVETK+GAD LE+WLC+N FPAT+IHGDR+QQERE ALRSFK
Sbjct: 398 LHAQRE--TQDKQSLTLVFVETKRGADTLENWLCMNEFPATSIHGDRTQQEREVALRSFK 455
Query: 452 SGNTPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRAGKKGLATAFFNEN 511
+G TPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRAGK G+ATAFFNEN
Sbjct: 456 TGRTPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRAGKSGIATAFFNEN 515
Query: 512 NTSMARSLQDLMQEANQEVPAWLSRFAARSSFGGGKNRRSGGGRFGGRDFRREGSFSRGG 571
N +ARSL +LMQEANQEVP WL+R+A+R+SFGGGK R GGRFGGRDFRREGS+SRGG
Sbjct: 516 NAQLARSLAELMQEANQEVPEWLTRYASRASFGGGKKR--SGGRFGGRDFRREGSYSRGG 573
Query: 572 SDYHGAGNSGGGYGASGGYGGGGYGGGYAGNSAGPXVTTAW 612
G G G Y GGYGGGGYGG +G G VT+AW
Sbjct: 574 GG--GGGGGGSDYYGGGGYGGGGYGGAPSG-GYGAGVTSAW 611
>At3g58570 ATP-dependent RNA helicase-like protein
Length = 646
Score = 858 bits (2216), Expect = 0.0
Identities = 460/630 (73%), Positives = 507/630 (80%), Gaps = 41/630 (6%)
Query: 3 SSWADLAANSAAENAPTPPPSAAAAVANNAANTRPVYVPPHLRNRGPSAPAPA-APAFDN 61
+SWAD++ + A PS + + T YVPPHLR+R PS+ A +P ++
Sbjct: 4 NSWADVSESERA-------PSGGGWGYSRPSRTN--YVPPHLRSRTPSSEFVAPSPGNND 54
Query: 62 SGSRWAPPPRNDYRGGGGGRGGGGYGNRGG---------GGWDRR--EANPFADQDDSEE 110
G RG G G G GYG RGG GGWDRR E NPF + +++
Sbjct: 55 RGGYGGANSGYGGRGQGYGGRGSGYGGRGGPVGGWNARSGGWDRRDTETNPFGNDGNADP 114
Query: 111 PVTQEEQENTGINFDAYEDIPVETSGGNVPPPVNTFAEIDLGEALNQNIRRCKYVKPTPV 170
V EQENT INF+AYEDIP+ETSG NVPPPVNTFAEIDLGEALN NI+RCKYVKPTPV
Sbjct: 115 AVN--EQENTVINFEAYEDIPIETSGDNVPPPVNTFAEIDLGEALNLNIQRCKYVKPTPV 172
Query: 171 QRHAIPISLGGRDLMACAQTGSGKTAAFCFPIISGIMTGQPAQRPPRGVRTVCPLALVLS 230
QR+AIPI GRDLMACAQTGSGKTAAFCFPIISGIM Q +RP RGVR V PLA++LS
Sbjct: 173 QRNAIPILAAGRDLMACAQTGSGKTAAFCFPIISGIMKDQHIERP-RGVRGVYPLAVILS 231
Query: 231 PTRELSMQIHEEARKFSYQTGVRVVVAYGGAPINQQLRELERGVDILVATPGRLVDLLER 290
PTREL+ QIH+EARKFSYQTGV+VVVAYGG P+NQQ+RELERGVDILVATPGRL DLLER
Sbjct: 232 PTRELACQIHDEARKFSYQTGVKVVVAYGGTPVNQQIRELERGVDILVATPGRLNDLLER 291
Query: 291 ARVSLSMIRYLALDEADRMLDMGFEPQIRKIVEQMDMPPAGVRQTMLFSATFPKEIQRLA 350
RVSL M+R+LALDEADRMLDMGFEPQIRKIV+QMDMPP GVRQTMLFSATFP+EIQRLA
Sbjct: 292 GRVSLQMVRFLALDEADRMLDMGFEPQIRKIVQQMDMPPPGVRQTMLFSATFPREIQRLA 351
Query: 351 SDFLSNYIFLAVGRVGSSTDLIDQRVEYVQESDKRSHLMDLLHAQRANGVQGKQALTLVF 410
SDFLSNYIFLAVGRVGSSTDLI QRVE+V +SDKRSHLMDLLHAQR NG QGKQALTLVF
Sbjct: 352 SDFLSNYIFLAVGRVGSSTDLIVQRVEFVHDSDKRSHLMDLLHAQRENGNQGKQALTLVF 411
Query: 411 VETKKGADALEHWLCLNNFPATTIHGDRSQQEREAALRSFKSGNTPILVATDVAARGLDI 470
VETKKGAD+LE+WLC+N FPATTIHGDRSQQERE ALRSFK+G TPILVATDVAARGLDI
Sbjct: 412 VETKKGADSLENWLCINGFPATTIHGDRSQQEREVALRSFKTGRTPILVATDVAARGLDI 471
Query: 471 PHVAHVVNFDLPNDIDDYVHRIGRTGRAGKKGLATAFFNENNTSMARSLQDLMQEANQEV 530
PHVAHVVNFDLPNDIDDYVHRIGRTGRAG GLATAFFN+NNT+MA+ L +LMQEANQEV
Sbjct: 472 PHVAHVVNFDLPNDIDDYVHRIGRTGRAGNSGLATAFFNDNNTTMAKPLAELMQEANQEV 531
Query: 531 PAWLSRFAARSSFGGGKNRRSGGGRFGGRDFRREGSFSR--GGSDYHGAGNS-----GGG 583
P WL+R+A+R+SFGGGKNRRS GGRFGGRDFRRE SFSR GG+DY+G G GGG
Sbjct: 532 PDWLTRYASRASFGGGKNRRS-GGRFGGRDFRRE-SFSRGGGGADYYGGGGGYGGVPGGG 589
Query: 584 YGA-SGGYG---GGGY----GGGYAGNSAG 605
YGA GGYG GGGY GGGYA G
Sbjct: 590 YGAMPGGYGPVPGGGYGNVPGGGYAPYGRG 619
>At5g63120 ATP-dependent RNA helicase-like protein
Length = 591
Score = 383 bits (983), Expect = e-106
Identities = 243/569 (42%), Positives = 325/569 (56%), Gaps = 62/569 (10%)
Query: 39 YVPPHLRNRGPSAPAPAAPAFDNSGSRWAP---PPRNDYR-GGGGGRGG-GGYGNRGGGG 93
Y P + G P+P A DNSG P PP + G GGGRGG G YG+R GGG
Sbjct: 35 YNPRYTGGGGGYGPSPVM-AGDNSGYNRYPSFQPPSGGFSVGRGGGRGGYGQYGDRNGGG 93
Query: 94 -W---------DRREANP-------------FADQDDSEEPVTQEEQENTGINFDAYEDI 130
W +RE + F E P Q E + DI
Sbjct: 94 NWGGGGGRGGSSKRELDSVSLPKQNFGNLVHFEKNFYVESPTVQAMTEQDVAMYRTERDI 153
Query: 131 PVETSGGNVPPPVNTFAEIDLGEALNQNIRRCKYVKPTPVQRHAIPISLGGRDLMACAQT 190
VE G +VP P+ F + + + + + I + + +PTP+Q P++L GRDL+ A+T
Sbjct: 154 SVE--GRDVPKPMKMFQDANFPDNILEAIAKLGFTEPTPIQAQGWPMALKGRDLIGIAET 211
Query: 191 GSGKTAAFCFPIISGIMTGQPAQRPPRGVRTVCPLALVLSPTRELSMQIHEEARKFSYQT 250
GSGKT A+ P + + + QP G P+ L+L+PTREL++QI EE+RKF ++
Sbjct: 212 GSGKTLAYLLPALVHV-SAQPRLGQDDG-----PIVLILAPTRELAVQIQEESRKFGLRS 265
Query: 251 GVRVVVAYGGAPINQQLRELERGVDILVATPGRLVDLLERARVSLSMIRYLALDEADRML 310
GVR YGGAP Q+R+L RGV+I++ATPGRL+D+LE +L + YL LDEADRML
Sbjct: 266 GVRSTCIYGGAPKGPQIRDLRRGVEIVIATPGRLIDMLECQHTNLKRVTYLVLDEADRML 325
Query: 311 DMGFEPQIRKIVEQMDMPPAGVRQTMLFSATFPKEIQRLASDFLSNYIFLAVGRVGSSTD 370
DMGFEPQIRKIV Q+ RQT+L+SAT+P+E++ LA FL + +G STD
Sbjct: 326 DMGFEPQIRKIVSQIRPD----RQTLLWSATWPREVETLARQFLRDPYKAIIG----STD 377
Query: 371 L-----IDQRVEYVQESDKRSHLMDLLHAQRANGVQGKQALTLVFVETKKGADALEHWLC 425
L I+Q +E V +K + L+ LL Q +G + L+FVETK+G D + L
Sbjct: 378 LKANQSINQVIEIVPTPEKYNRLLTLLK-QLMDGSK-----ILIFVETKRGCDQVTRQLR 431
Query: 426 LNNFPATTIHGDRSQQEREAALRSFKSGNTPILVATDVAARGLDIPHVAHVVNFDLPNDI 485
++ +PA IHGD++Q ER+ L FKSG +PI+ ATDVAARGLD+ + VVN+D PN +
Sbjct: 432 MDGWPALAIHGDKTQSERDRVLAEFKSGRSPIMTATDVAARGLDVKDIKCVVNYDFPNTL 491
Query: 486 DDYVHRIGRTGRAGKKGLATAFFNENNTSMARSLQDLMQEANQEVPAWLSRFAARS---- 541
+DY+HRIGRTGRAG KG+A FF +N AR L ++QEA Q VP LS S
Sbjct: 492 EDYIHRIGRTGRAGAKGMAFTFFTHDNAKFARELVKILQEAGQVVPPTLSALVRSSGSGY 551
Query: 542 -SFGGGKN-RRSGGGRFGGRDFRREGSFS 568
GGG+N R GGGR GG +R S S
Sbjct: 552 GGSGGGRNFRPRGGGRGGGFGDKRSRSTS 580
Score = 30.4 bits (67), Expect = 3.0
Identities = 16/36 (44%), Positives = 18/36 (49%), Gaps = 2/36 (5%)
Query: 557 GGRDFRREGSFSRGGSDYHGAGNSGGGYGASGGYGG 592
GG R G RGG +G N GG +G GG GG
Sbjct: 70 GGFSVGRGGG--RGGYGQYGDRNGGGNWGGGGGRGG 103
Score = 30.0 bits (66), Expect = 3.9
Identities = 22/64 (34%), Positives = 26/64 (40%), Gaps = 12/64 (18%)
Query: 552 GGGRFG--------GRDFRREGSFSRGGSDYH-GAGNSGGGYGASGGYGGGGY---GGGY 599
GGG +G + R SF + G G GGYG G GGG GGG
Sbjct: 42 GGGGYGPSPVMAGDNSGYNRYPSFQPPSGGFSVGRGGGRGGYGQYGDRNGGGNWGGGGGR 101
Query: 600 AGNS 603
G+S
Sbjct: 102 GGSS 105
>At3g01540 AT3g01540/F4P13_9
Length = 619
Score = 343 bits (880), Expect = 2e-94
Identities = 223/561 (39%), Positives = 298/561 (52%), Gaps = 60/561 (10%)
Query: 49 PSAPAPAAPAFDNSGS-------RWAPPPRNDYRGGG---------GGRGGGGYGNRGGG 92
PS P +PA S S +APP +D G GGR G Y N
Sbjct: 51 PSLPPKFSPAVSVSSSVQVQQTDAYAPPKDDDKYSRGSERVSRFSEGGRSGPPYSNGAAN 110
Query: 93 GWDRREANPFADQDDSEEPVTQEEQENTGINFDAYEDIPVETSGGNVPPPVNTFAEIDLG 152
G ++ A P + E + + +I V SGG VPPP+ +F
Sbjct: 111 GVG--DSAYGAASTRVPLPSSAPASELSPEAYSRRHEITV--SGGQVPPPLMSFEATGFP 166
Query: 153 EALNQNIRRCKYVKPTPVQRHAIPISLGGRDLMACAQTGSGKTAAFCFPIISGIMTGQPA 212
L + + + PTP+Q + PI++ GRD++A A+TGSGKT + P G + Q
Sbjct: 167 PELLREVLSAGFSAPTPIQAQSWPIAMQGRDIVAIAKTGSGKTLGYLIP---GFLHLQRI 223
Query: 213 QRPPRGVRTVCPLALVLSPTRELSMQIHEEARKFSYQTGVRVVVAYGGAPINQQLRELER 272
+ R + P LVLSPTREL+ QI EEA KF + + YGGAP QLR+LER
Sbjct: 224 RNDSR----MGPTILVLSPTRELATQIQEEAVKFGRSSRISCTCLYGGAPKGPQLRDLER 279
Query: 273 GVDILVATPGRLVDLLERARVSLSMIRYLALDEADRMLDMGFEPQIRKIVEQMDMPPAGV 332
G DI+VATPGRL D+LE R+SL I YL LDEADRMLDMGFEPQIRKIV+++
Sbjct: 280 GADIVVATPGRLNDILEMRRISLRQISYLVLDEADRMLDMGFEPQIRKIVKEIPTK---- 335
Query: 333 RQTMLFSATFPKEIQRLASDFLSNYIFLAVGRVGS--STDLIDQRVEYVQESDKRSHLMD 390
RQT++++AT+PK ++++A+D L N + +G V + I Q +E V +K+ L
Sbjct: 336 RQTLMYTATWPKGVRKIAADLLVNPAQVNIGNVDELVANKSITQHIEVVAPMEKQRRLEQ 395
Query: 391 LLHAQRANGVQGKQALTLVFVETKKGADALEHWLCLNNFPATTIHGDRSQQEREAALRSF 450
+L +Q + ++F TK+ D L L F A IHGD+SQ ER+ L F
Sbjct: 396 ILRSQEPG------SKVIIFCSTKRMCDQLTRNL-TRQFGAAAIHGDKSQPERDNVLNQF 448
Query: 451 KSGNTPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRAGKKGLATAFFNE 510
+SG TP+LVATDVAARGLD+ + VVN+D PN ++DYVHRIGRTGRAG G A FF +
Sbjct: 449 RSGRTPVLVATDVAARGLDVKDIRAVVNYDFPNGVEDYVHRIGRTGRAGATGQAFTFFGD 508
Query: 511 NNTSMARSLQDLMQEANQEVPAWLSRFAARSSFGGGKNRRSGGGRFGGRDFRREGSFSRG 570
++ A L +++ ANQ VP + A R GGG N+ FSR
Sbjct: 509 QDSKHASDLIKILEGANQRVPPQIREMATRG--GGGMNK-----------------FSRW 549
Query: 571 GSDYHGAGNSG-GGYGASGGY 590
G G G G GYG G +
Sbjct: 550 GPPSGGRGRGGDSGYGGRGSF 570
>At5g14610 DRH1 DEAD box protein - like
Length = 713
Score = 338 bits (868), Expect = 4e-93
Identities = 201/460 (43%), Positives = 264/460 (56%), Gaps = 24/460 (5%)
Query: 141 PPVNTFAEIDLGEALNQNIRRCKYVKPTPVQRHAIPISLGGRDLMACAQTGSGKTAAFCF 200
PP F + A + + + P+P+Q + PI++ RD++A A+TGSGKT +
Sbjct: 226 PPAAGFNSYLVLPANGRMVYSAGFSAPSPIQAQSWPIAMQNRDIVAIAKTGSGKTLGYLI 285
Query: 201 PIISGIMTGQPAQRPPRGVRTVCPLALVLSPTRELSMQIHEEARKFSYQTGVRVVVAYGG 260
P G M Q R + P LVLSPTREL+ QI EA KF + + YGG
Sbjct: 286 P---GFMHLQRIHNDSR----MGPTILVLSPTRELATQIQVEALKFGKSSKISCACLYGG 338
Query: 261 APINQQLRELERGVDILVATPGRLVDLLERARVSLSMIRYLALDEADRMLDMGFEPQIRK 320
AP QL+E+ERGVDI+VATPGRL D+LE R+SL + YL LDEADRMLDMGFEPQIRK
Sbjct: 339 APKGPQLKEIERGVDIVVATPGRLNDILEMKRISLHQVSYLVLDEADRMLDMGFEPQIRK 398
Query: 321 IVEQMDMPPAGVRQTMLFSATFPKEIQRLASDFLSNYIFLAVGRVGS--STDLIDQRVEY 378
IV ++ RQT++++AT+PKE++++A+D L N + +G V + I Q +E
Sbjct: 399 IVNEVPTK----RQTLMYTATWPKEVRKIAADLLVNPAQVNIGNVDELVANKSITQTIEV 454
Query: 379 VQESDKRSHLMDLLHAQRANGVQGKQALTLVFVETKKGADALEHWLCLNNFPATTIHGDR 438
+ +K S L +L +Q + ++F TK+ D L L F A IHGD+
Sbjct: 455 LAPMEKHSRLEQILRSQEPG------SKIIIFCSTKRMCDQLARNLT-RTFGAAAIHGDK 507
Query: 439 SQQEREAALRSFKSGNTPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRA 498
SQ ER+ L F+SG TP+LVATDVAARGLD+ + VVN+D PN ++DYVHRIGRTGRA
Sbjct: 508 SQAERDDVLNQFRSGRTPVLVATDVAARGLDVKDIRVVVNYDFPNGVEDYVHRIGRTGRA 567
Query: 499 GKKGLATAFFNENNTSMARSLQDLMQEANQEVPAWLSRFAARSSFGGGKNRRSGGGRFGG 558
G GLA FF + + A L +++ ANQ+VP + A R G K RR G GG
Sbjct: 568 GATGLAYTFFGDQDAKHASDLIKILEGANQKVPPQVREMATRGGGGMNKFRRWGTPSSGG 627
Query: 559 RDFRREGSFSRGGSDYHGAGNSGGGYGASGGYGGGGYGGG 598
G G S Y G G SG G GYGG G GG
Sbjct: 628 GG----GRGGYGDSGYGGRGESGYGSRGDSGYGGRGDSGG 663
Score = 38.1 bits (87), Expect = 0.014
Identities = 26/72 (36%), Positives = 31/72 (42%), Gaps = 12/72 (16%)
Query: 40 VPPHLRNRGPSAPAPAAPAFDNSGSRWAPPPRNDYRGGGGGRGG---GGYGNRGGGGWDR 96
VPP +R A N RW P GGGGGRGG GYG RG G+
Sbjct: 599 VPPQVREM-----ATRGGGGMNKFRRWGTPSS----GGGGGRGGYGDSGYGGRGESGYGS 649
Query: 97 REANPFADQDDS 108
R + + + DS
Sbjct: 650 RGDSGYGGRGDS 661
Score = 36.2 bits (82), Expect = 0.055
Identities = 36/145 (24%), Positives = 54/145 (36%), Gaps = 29/145 (20%)
Query: 47 RGPSAPAPAAPAFDNSGSRWAPPPRNDYRGGGGGRGGGGYGNRGGGGWDRREANPFADQD 106
RG P + + N R P ND G G GG RG + A +
Sbjct: 85 RGSDGPKSDSGSRFNEAGRTGPISSNDAASGLGNASSGGSSARG-------PPSSAAGNE 137
Query: 107 DSEEPVTQEEQENTGINFDAYEDIPVETSGGNVPPPVNTFAEIDLGEALNQNIRRCKYVK 166
S E ++ + + SGG VPPP+ +F L N+ +R C
Sbjct: 138 LSPEAYCRKHE--------------ITVSGGQVPPPLMSFEATGLP---NELLRECPNFM 180
Query: 167 PTPVQRHAIP---ISLGGRDLMACA 188
P + +P +LGG +++ CA
Sbjct: 181 PYALS--FVPGAIFALGGANVVRCA 203
>At3g06480 putative RNA helicase
Length = 1088
Score = 334 bits (856), Expect = 1e-91
Identities = 199/476 (41%), Positives = 276/476 (57%), Gaps = 28/476 (5%)
Query: 132 VETSGGNVPPPVNTFAEIDLGEALNQNIRRCKYVKPTPVQRHAIPISLGGRDLMACAQTG 191
V T+G N+P P TF L + + + + PTP+Q PI+L RD++A A+TG
Sbjct: 423 VTTTGENIPAPYITFESSGLPPEILRELLSAGFPSPTPIQAQTWPIALQSRDIVAIAKTG 482
Query: 192 SGKTAAFCFPIISGIMTGQPAQRPPRGVRTVCPLALVLSPTRELSMQIHEEARKFSYQTG 251
SGKT + P + R R P L+L+PTREL+ QI +EA +F +
Sbjct: 483 SGKTLGYLIPAFILL-------RHCRNDSRNGPTVLILAPTRELATQIQDEALRFGRSSR 535
Query: 252 VRVVVAYGGAPINQQLRELERGVDILVATPGRLVDLLERARVSLSMIRYLALDEADRMLD 311
+ YGGAP QL+ELERG DI+VATPGRL D+LE + + L LDEADRMLD
Sbjct: 536 ISCTCLYGGAPKGPQLKELERGADIVVATPGRLNDILEMKMIDFQQVSLLVLDEADRMLD 595
Query: 312 MGFEPQIRKIVEQMDMPPAGVRQTMLFSATFPKEIQRLASDFLSNYIFLAVGRVG--SST 369
MGFEPQIRKIV ++ PP RQT++++AT+PKE++++ASD L N + + +GRV ++
Sbjct: 596 MGFEPQIRKIVNEI--PPR--RQTLMYTATWPKEVRKIASDLLVNPVQVNIGRVDELAAN 651
Query: 370 DLIDQRVEYVQESDKRSHLMDLLHAQRANGVQGKQALTLVFVETKKGADALEHWLCLNNF 429
I Q VE V + +K L +L +Q + + ++F TK+ D L + +F
Sbjct: 652 KAITQYVEVVPQMEKERRLEQILRSQE------RGSKVIIFCSTKRLCDHLARSVG-RHF 704
Query: 430 PATTIHGDRSQQEREAALRSFKSGNTPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYV 489
A IHGD++Q ER+ L F+SG + +L+ATDVAARGLDI + V+N+D P ++DYV
Sbjct: 705 GAVVIHGDKTQGERDWVLNQFRSGKSCVLIATDVAARGLDIKDIRVVINYDFPTGVEDYV 764
Query: 490 HRIGRTGRAGKKGLATAFFNENNTSMARSLQDLMQEANQEVPAWLSRFAARSSFGGG--- 546
HRIGRTGRAG G+A FF E + A L +++ ANQ+VP + A R GGG
Sbjct: 765 HRIGRTGRAGATGVAFTFFTEQDWKYAPDLIKVLEGANQQVPPQVRDIAMRGGGGGGPGY 824
Query: 547 -KNRRSGGGRF--GGRDFRREGSFSRGGSDYHGAGNSGGGYGASGGYGG--GGYGG 597
++RR RF GG R + GG +G GG G GG+GG GG+GG
Sbjct: 825 SQDRRGMVNRFDSGGGGTRWDSGGGFGGRGGGFSGREGGFGGREGGFGGREGGFGG 880
Score = 32.7 bits (73), Expect = 0.61
Identities = 32/93 (34%), Positives = 38/93 (40%), Gaps = 30/93 (32%)
Query: 40 VPPHLRN---RGPSAPAPA--------APAFDNSG--SRWAPPPRNDYRGGGGGRGGGGY 86
VPP +R+ RG P FD+ G +RW D GG GGRGGG
Sbjct: 805 VPPQVRDIAMRGGGGGGPGYSQDRRGMVNRFDSGGGGTRW------DSGGGFGGRGGGFS 858
Query: 87 GNRGG-----GGWDRRE------ANPFADQDDS 108
G GG GG+ RE F +DDS
Sbjct: 859 GREGGFGGREGGFGGREGGFGGRGGRFGMRDDS 891
>At1g55150 unknown protein
Length = 501
Score = 330 bits (847), Expect = 1e-90
Identities = 194/506 (38%), Positives = 291/506 (57%), Gaps = 46/506 (9%)
Query: 73 DYRGGGGGRGGGGYGNRGGGGWDRREAN--------------PFADQDDSEEPVTQEEQE 118
D R G G YG+ G +++ + PF E P +
Sbjct: 16 DRRSDSGFGGTSSYGSSGSHTSSKKDNDGNESPRKLDLDGLTPFEKNFYVESPAVAAMTD 75
Query: 119 NTGINFDAYEDIPVETSGGNVPPPVNTFAEIDLGEALNQNIRRCKYVKPTPVQRHAIPIS 178
+ +I VE G ++P PV +F ++ + + + +++ + +PTP+Q P++
Sbjct: 76 TEVEEYRKLREITVE--GKDIPKPVKSFRDVGFPDYVLEEVKKAGFTEPTPIQSQGWPMA 133
Query: 179 LGGRDLMACAQTGSGKTAAFCFPIISGIMTGQPAQRPPRGVRTVCPLALVLSPTRELSMQ 238
+ GRDL+ A+TGSGKT ++ P I + QP G P+ LVL+PTREL++Q
Sbjct: 134 MKGRDLIGIAETGSGKTLSYLLPAIVHV-NAQPMLAHGDG-----PIVLVLAPTRELAVQ 187
Query: 239 IHEEARKFSYQTGVRVVVAYGGAPINQQLRELERGVDILVATPGRLVDLLERARVSLSMI 298
I +EA KF + ++ YGG P Q+R+L++GV+I++ATPGRL+D++E +L +
Sbjct: 188 IQQEASKFGSSSKIKTTCIYGGVPKGPQVRDLQKGVEIVIATPGRLIDMMESNNTNLRRV 247
Query: 299 RYLALDEADRMLDMGFEPQIRKIVEQMDMPPAGVRQTMLFSATFPKEIQRLASDFLSNYI 358
YL LDEADRMLDMGF+PQIRKIV + RQT+ +SAT+PKE+++L+ FL N
Sbjct: 248 TYLVLDEADRMLDMGFDPQIRKIVSHIRPD----RQTLYWSATWPKEVEQLSKKFLYNPY 303
Query: 359 FLAVGRVGSSTDL-----IDQRVEYVQESDKRSHLMDLLHAQRANGVQGKQALTLVFVET 413
+ +G S+DL I Q V+ + ES K + L+ LL + + G + L VF++T
Sbjct: 304 KVIIG----SSDLKANRAIRQIVDVISESQKYNKLVKLLE----DIMDGSRIL--VFLDT 353
Query: 414 KKGADALEHWLCLNNFPATTIHGDRSQQEREAALRSFKSGNTPILVATDVAARGLDIPHV 473
KKG D + L ++ +PA +IHGD+SQ ER+ L F+SG +PI+ ATDVAARGLD+ V
Sbjct: 354 KKGCDQITRQLRMDGWPALSIHGDKSQAERDWVLSEFRSGKSPIMTATDVAARGLDVKDV 413
Query: 474 AHVVNFDLPNDIDDYVHRIGRTGRAGKKGLATAFFNENNTSMARSLQDLMQEANQEVPAW 533
+V+N+D P ++DYVHRIGRTGRAG KG A FF N A+ L +++QEA Q+V
Sbjct: 414 KYVINYDFPGSLEDYVHRIGRTGRAGAKGTAYTFFTVANARFAKELTNILQEAGQKVSPE 473
Query: 534 LSRFAARSS-----FGGGKNRRSGGG 554
L+ ++ GG ++R S G
Sbjct: 474 LASMGRSTAPPPPGLGGFRDRGSRRG 499
>At2g33730 putative U5 small nuclear ribonucleoprotein, an RNA
helicase
Length = 733
Score = 313 bits (802), Expect = 2e-85
Identities = 172/425 (40%), Positives = 252/425 (58%), Gaps = 23/425 (5%)
Query: 128 EDIPVETSGGNVPPPVNTFAEIDLGEALNQNIRRCKYVKPTPVQRHAIPISLGGRDLMAC 187
ED + G +P P+ ++ E L L + + R Y KP+P+Q AIP+ L RD++
Sbjct: 297 EDFNISYKGSRIPRPMRSWEESKLTSELLKAVERAGYKKPSPIQMAAIPLGLQQRDVIGI 356
Query: 188 AQTGSGKTAAFCFPIISGIMTGQPAQRPPRGVRTVCPLALVLSPTRELSMQIHEEARKFS 247
A+TGSGKTAAF P+++ I P T P A+V++PTREL+ QI EE KF+
Sbjct: 357 AETGSGKTAAFVLPMLAYISRLPPMSEENE---TEGPYAVVMAPTRELAQQIEEETVKFA 413
Query: 248 YQTGVRVVVAYGGAPINQQLRELERGVDILVATPGRLVDLLERARVSLSMIRYLALDEAD 307
+ G RV GG I +Q ++ +G +I++ATPGRL+D LER L+ Y+ LDEAD
Sbjct: 414 HYLGFRVTSIVGGQSIEEQGLKITQGCEIVIATPGRLIDCLERRYAVLNQCNYVVLDEAD 473
Query: 308 RMLDMGFEPQIRKIVEQM---DMPPAG----------VRQTMLFSATFPKEIQRLASDFL 354
RM+DMGFEPQ+ +++ M ++ P R T +FSAT P ++RLA +L
Sbjct: 474 RMIDMGFEPQVAGVLDAMPSSNLKPENEEEELDEKKIYRTTYMFSATMPPGVERLARKYL 533
Query: 355 SNYIFLAVGRVGSSTDLIDQRVEYVQESDKRSHLMDLLHAQRANGVQGKQALTLVFVETK 414
N + + +G G +TDLI Q V ++ES+K L LL + + +VFV TK
Sbjct: 534 RNPVVVTIGTAGKTTDLISQHVIMMKESEKFFRLQKLLD-------ELGEKTAIVFVNTK 586
Query: 415 KGADALEHWLCLNNFPATTIHGDRSQQEREAALRSFKSGNTPILVATDVAARGLDIPHVA 474
K D++ L + TT+HG +SQ++RE +L F++ +LVATDV RG+DIP VA
Sbjct: 587 KNCDSIAKNLDKAGYRVTTLHGGKSQEQREISLEGFRAKRYNVLVATDVVGRGIDIPDVA 646
Query: 475 HVVNFDLPNDIDDYVHRIGRTGRAGKKGLATAFFNENNTSMARSLQDLMQEANQEVPAWL 534
HV+N+D+P I+ Y HRIGRTGRAGK G+AT+F ++T + L+ ++ ++N VP L
Sbjct: 647 HVINYDMPKHIEMYTHRIGRTGRAGKSGVATSFLTLHDTEVFYDLKQMLVQSNSAVPPEL 706
Query: 535 SRFAA 539
+R A
Sbjct: 707 ARHEA 711
>At2g47330 putative ATP-dependent RNA helicase
Length = 760
Score = 305 bits (782), Expect = 4e-83
Identities = 199/505 (39%), Positives = 277/505 (54%), Gaps = 54/505 (10%)
Query: 95 DRREANPFADQDDSE---EPVTQE------------EQENTGINFDAYEDIPVETSGGNV 139
D+R+ P D S EP+ ++ EQE T D + + + SG +V
Sbjct: 168 DKRKIEPITALDHSSIDYEPINKDFYEELESISGMTEQETT----DYRQRLGIRVSGFDV 223
Query: 140 PPPVNTFAEIDLGEALNQNIRRCKYVKPTPVQRHAIPISLGGRDLMACAQTGSGKTAAFC 199
PV TF + + I++ Y KPT +Q A+PI L GRD++ A+TGSGKTAAF
Sbjct: 224 HRPVKTFEDCGFSSQIMSAIKKQAYEKPTAIQCQALPIVLSGRDVIGIAKTGSGKTAAFV 283
Query: 200 FPIISGIMTGQPAQRPPRGVRTVCPLALVLSPTRELSMQIHEEARKFSYQTGVRVVVAYG 259
P+I IM QP + G P+ ++ +PTREL+ QI EA+KFS G+RV YG
Sbjct: 284 LPMIVHIMD-QPELQRDEG-----PIGVICAPTRELAHQIFLEAKKFSKAYGLRVSAVYG 337
Query: 260 GAPINQQLRELERGVDILVATPGRLVDLLERARVSLSMIRYLALDEADRMLDMGFEPQIR 319
G ++Q +EL+ G +I+VATPGRL+D+L+ +++ YL LDEADRM D+GFEPQ+R
Sbjct: 338 GMSKHEQFKELKAGCEIVVATPGRLIDMLKMKALTMMRASYLVLDEADRMFDLGFEPQVR 397
Query: 320 KIVEQMDMPPAGVRQTMLFSATFPKEIQRLASDFLSNYIFLAVGRVGSSTDLIDQRVEYV 379
IV Q+ RQT+LFSAT P ++++LA + LS+ I + VG VG + + I Q V +
Sbjct: 398 SIVGQIRPD----RQTLLFSATMPWKVEKLAREILSDPIRVTVGEVGMANEDITQVVNVI 453
Query: 380 -QESDKRSHLMDLLHAQRANGVQGKQALTLVFVETKKGADALEHWLCLNNFPATTIHGDR 438
+++K L++ L G LVF K D +E L LN+F +HGD+
Sbjct: 454 PSDAEKLPWLLEKLPGMIDEGD------VLVFASKKATVDEIEAQLTLNSFKVAALHGDK 507
Query: 439 SQQEREAALRSFKSGNTPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRA 498
Q R L+ FKSG +L+ATDVAARGLDI + VVN+D+ D+D +VHRIGRTGRA
Sbjct: 508 DQASRMETLQKFKSGVHHVLIATDVAARGLDIKSLKTVVNYDIAKDMDMHVHRIGRTGRA 567
Query: 499 G-KKGLATAFFNENNTSMARSLQDLMQEANQEVPAWLSRFAARSSFGGGKNRRSGGGRF- 556
G + G+A + A L + + A Q VP L+ A + GRF
Sbjct: 568 GDRDGVAYTLVTQREARFAGELVNSLVAAGQNVPPELTDLAMKD------------GRFK 615
Query: 557 GGRDFRREGSFSRGGSDYHGAGNSG 581
RD R+ G RGG G GN G
Sbjct: 616 SKRDGRKGGKKGRGG----GGGNKG 636
>At1g20920 putative RNA helicase
Length = 1166
Score = 295 bits (754), Expect = 7e-80
Identities = 161/414 (38%), Positives = 247/414 (58%), Gaps = 19/414 (4%)
Query: 128 EDIPVETSGGNVPPPVNTFAEIDLGEALNQNIRRCKYVKPTPVQRHAIPISLGGRDLMAC 187
+++ ++ G +VP P+ + + L + +++ Y KP P+Q A+PI + GRD +
Sbjct: 513 KELELKVHGKDVPRPIKFWHQTGLTSKILDTMKKLNYEKPMPIQTQALPIIMSGRDCIGV 572
Query: 188 AQTGSGKTAAFCFPIISGIMTGQPAQRPPRGVRTVCPLALVLSPTRELSMQIHEEARKFS 247
A+TGSGKT F P++ I P + P+ LV++PTREL QIH + RKFS
Sbjct: 573 AKTGSGKTLGFVLPMLRHIKDQPPVEAGDG------PIGLVMAPTRELVQQIHSDIRKFS 626
Query: 248 YQTGVRVVVAYGGAPINQQLRELERGVDILVATPGRLVDLLERARVSLSMIR---YLALD 304
G+R V YGG+ + QQ+ EL+RG +I+V TPGR++D+L + ++ +R +L +D
Sbjct: 627 KPLGIRCVPVYGGSGVAQQISELKRGTEIVVCTPGRMIDILCTSSGKITNLRRVTFLVMD 686
Query: 305 EADRMLDMGFEPQIRKIVEQMDMPPAGVRQTMLFSATFPKEIQRLASDFLSNYIFLAVGR 364
EADRM DMGFEPQI +I++ + RQT+LFSATFP++++ LA L+ + + VG
Sbjct: 687 EADRMFDMGFEPQITRIIQNIRPE----RQTVLFSATFPRQVETLARKVLNKPVEIQVGG 742
Query: 365 VGSSTDLIDQRVEYVQESDKRSHLMDLLHAQRANGVQGKQALTLVFVETKKGADALEHWL 424
I Q VE ESD+ L++LL G ++ LVFV++++ DAL +
Sbjct: 743 RSVVNKDITQLVEVRPESDRFLRLLELL------GEWSEKGKILVFVQSQEKCDALYRDM 796
Query: 425 CLNNFPATTIHGDRSQQEREAALRSFKSGNTPILVATDVAARGLDIPHVAHVVNFDLPND 484
+++P ++HG + Q +RE+ + FK+ +L+AT VAARGLD+ + VVNFD PN
Sbjct: 797 IKSSYPCLSLHGGKDQTDRESTISDFKNDVCNLLIATSVAARGLDVKELELVVNFDAPNH 856
Query: 485 IDDYVHRIGRTGRAGKKGLATAFFNENNTSMARSLQDLMQEANQEVPAWLSRFA 538
+DYVHR+GRTGRAG+KG A F +E++ A L ++ + Q VP L A
Sbjct: 857 YEDYVHRVGRTGRAGRKGCAVTFISEDDAKYAPDLVKALELSEQPVPDDLKALA 910
>At5g51280 DEAD-box protein abstrakt
Length = 591
Score = 287 bits (734), Expect = 1e-77
Identities = 170/468 (36%), Positives = 258/468 (54%), Gaps = 35/468 (7%)
Query: 135 SGGNVPPPVNTFAEIDLGEALNQNIRRCKYVKPTPVQRHAIPISLGGRDLMACAQTGSGK 194
+G ++PPP+ F ++ + ++ V+PTP+Q +P+ L GRD++ A TGSGK
Sbjct: 137 NGDDIPPPIKNFKDMKFPRPVLDTLKEKGIVQPTPIQVQGLPVILAGRDMIGIAFTGSGK 196
Query: 195 TAAFCFPIISGIMTGQPAQRPPRGVRTVCPLALVLSPTRELSMQIHEEARKFS---YQTG 251
T F P+I + + G P+ L++ P+REL+ Q +E +F + G
Sbjct: 197 TLVFVLPMIMIALQEEMMMPIAAGEG---PIGLIVCPSRELARQTYEVVEQFVAPLVEAG 253
Query: 252 ---VRVVVAYGGAPINQQLRELERGVDILVATPGRLVDLLERARVSLSMIRYLALDEADR 308
+R ++ GG + QL ++RGV I+VATPGRL D+L + ++SL RYL LDEADR
Sbjct: 254 YPPLRSLLCIGGIDMRSQLEVVKRGVHIVVATPGRLKDMLAKKKMSLDACRYLTLDEADR 313
Query: 309 MLDMGFEPQIRKIVEQMDMPPAGVRQTMLFSATFPKEIQRLASDFLSNYIFLAVGRVGSS 368
++D+GFE IR++ + RQT+LFSAT P +IQ A L + + VGR G++
Sbjct: 314 LVDLGFEDDIREVFDHFKSQ----RQTLLFSATMPTKIQIFARSALVKPVTVNVGRAGAA 369
Query: 369 TDLIDQRVEYVQESDKRSHLMDLLHAQRANGVQGKQALTLVFVETKKGADALEHWLCLNN 428
+ Q VEYV++ K +L++ L Q L+F E K D + +L L
Sbjct: 370 NLDVIQEVEYVKQEAKIVYLLECL--------QKTSPPVLIFCENKADVDDIHEYLLLKG 421
Query: 429 FPATTIHGDRSQQEREAALRSFKSGNTPILVATDVAARGLDIPHVAHVVNFDLPNDIDDY 488
A IHG + Q++RE A+ SFK+G +LVATDVA++GLD P + HV+N+D+P +I++Y
Sbjct: 422 VEAVAIHGGKDQEDREYAISSFKAGKKDVLVATDVASKGLDFPDIQHVINYDMPAEIENY 481
Query: 489 VHRIGRTGRAGKKGLATAFFNENNT-SMARSLQDLMQEANQEVPAWLSRF------AARS 541
VHRIGRTGR GK G+AT F N+N + + L+ L+QEA Q +P L+ A
Sbjct: 482 VHRIGRTGRCGKTGIATTFINKNQSETTLLDLKHLLQEAKQRIPPVLAELNDPMEEAETI 541
Query: 542 SFGGGKNRRSGGGRFGGR-------DFRREGSFSRGGSDYHGAGNSGG 582
+ G + G G R + ++ + S DY G+G G
Sbjct: 542 ANASGVKGCAYCGGLGHRIRDCPKLEHQKSVAISNSRKDYFGSGGYRG 589
>At1g31970 unknown protein
Length = 537
Score = 280 bits (716), Expect = 2e-75
Identities = 179/451 (39%), Positives = 257/451 (56%), Gaps = 26/451 (5%)
Query: 96 RREANPFADQDDSEEPVTQEEQENTGINFDAYEDIPVETSGGNVPP--PVNTFAEIDLGE 153
+RE + P + E E+ G + + V G + TFAE +L E
Sbjct: 66 KREEKESEKNKKKDVPEKKLEAEDLGEGESEQQKVVVTGKGVEEAKYAALKTFAESNLPE 125
Query: 154 ALNQNIRRC--KYVKPTPVQRHAIPISLGGRDLMACAQTGSGKTAAFCFPIISGIMTGQP 211
N+ C + KP+P+Q H P L GRDL+ A+TGSGKT AF P I ++ +
Sbjct: 126 ----NVLDCCKTFEKPSPIQSHTWPFLLDGRDLIGIAKTGSGKTLAFGIPAIMHVL--KK 179
Query: 212 AQRPPRGVRTVCPLALVLSPTRELSMQIHEEARKFSYQTGVRVVVAYGGAPINQQLRELE 271
++ G + V P LVLSPTREL++QI + R+ G++ + YGG+ Q+ +
Sbjct: 180 NKKIGGGSKKVNPTCLVLSPTRELAVQISDVLREAGEPCGLKSICVYGGSSKGPQISAIR 239
Query: 272 RGVDILVATPGRLVDLLERARVSLSMIRYLALDEADRMLDMGFEPQIRKIVEQMDMPPAG 331
GVDI++ TPGRL DL+E + LS + ++ LDEADRMLDMGFE +R I+ +
Sbjct: 240 SGVDIVIGTPGRLRDLIESNVLRLSDVSFVVLDEADRMLDMGFEEPVRFILSNTNK---- 295
Query: 332 VRQTMLFSATFPKEIQRLASDFLS-NYIFLAVGRVGSSTDL-IDQRVEYVQESDKRSHLM 389
VRQ ++FSAT+P ++ +LA +F+ N I + +G V + + + Q +E + E + L+
Sbjct: 296 VRQMVMFSATWPLDVHKLAQEFMDPNPIKVIIGSVDLAANHDVMQIIEVLDERARDQRLI 355
Query: 390 DLL---HAQRANGVQGKQALTLVFVETKKGADALEHWLCLNNFPATTIHGDRSQQEREAA 446
LL H + N V LVF K A+ LE +L + A +IHG+++Q ER +
Sbjct: 356 ALLEKYHKSQKNRV-------LVFALYKVEAERLERFLQQRGWKAVSIHGNKAQSERTRS 408
Query: 447 LRSFKSGNTPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRAGKKGLATA 506
L FK G+ P+LVATDVAARGLDIP V V+N+ P +DYVHRIGRTGRAGKKG+A
Sbjct: 409 LSLFKEGSCPLLVATDVAARGLDIPDVEVVINYTFPLTTEDYVHRIGRTGRAGKKGVAHT 468
Query: 507 FFNENNTSMARSLQDLMQEANQEVPAWLSRF 537
FF N +A L ++++EA Q VPA L +F
Sbjct: 469 FFTPLNKGLAGELVNVLREAGQVVPADLLKF 499
>At4g33370 putative protein
Length = 542
Score = 278 bits (712), Expect = 5e-75
Identities = 157/411 (38%), Positives = 239/411 (57%), Gaps = 22/411 (5%)
Query: 132 VETSGGNVPPPVNTFAEIDLGEALNQNIRRCKYVKPTPVQRHAIPISLGGRDLMACAQTG 191
+ +G ++PPP+ F ++ L + ++ + PTP+Q +P+ L GRD++ A TG
Sbjct: 85 ITVNGEDIPPPIKNFMDMKFPSPLLRMLKDKGIMHPTPIQVQGLPVVLSGRDMIGIAFTG 144
Query: 192 SGKTAAFCFPIISGIMTGQPAQRPPRGVRTVCPLALVLSPTRELSMQIHEEARKFSYQT- 250
SGKT F P+I I+ Q P P+ALV+ P+REL+ Q ++ +F
Sbjct: 145 SGKTLVFVLPMI--ILALQEEIMMPIAAGEG-PIALVICPSRELAKQTYDVVEQFVASLV 201
Query: 251 -----GVRVVVAYGGAPINQQLRELERGVDILVATPGRLVDLLERARVSLSMIRYLALDE 305
+R ++ GG + QL +++GV I+VATPGRL D+L + ++SL R L LDE
Sbjct: 202 EDGYPRLRSLLCIGGVDMRSQLDVVKKGVHIVVATPGRLKDILAKKKMSLDACRLLTLDE 261
Query: 306 ADRMLDMGFEPQIRKIVEQMDMPPAGVRQTMLFSATFPKEIQRLASDFLSNYIFLAVGRV 365
ADR++D+GFE IR + + RQT+LFSAT P +IQ A+ L + + VGR
Sbjct: 262 ADRLVDLGFEDDIRHVFDHFKSQ----RQTLLFSATMPAKIQIFATSALVKPVTVNVGRA 317
Query: 366 GSSTDLIDQRVEYVQESDKRSHLMDLLHAQRANGVQGKQALTLVFVETKKGADALEHWLC 425
G++ + Q VEYV++ K +L++ L Q L+F E K D + +L
Sbjct: 318 GAANLDVIQEVEYVKQEAKIVYLLECL--------QKTTPPVLIFCENKADVDDIHEYLL 369
Query: 426 LNNFPATTIHGDRSQQEREAALRSFKSGNTPILVATDVAARGLDIPHVAHVVNFDLPNDI 485
L A IHG + Q++R+ A+ FK+G +LVATDVA++GLD P + HV+N+D+P +I
Sbjct: 370 LKGVEAVAIHGGKDQEDRDYAISLFKAGKKDVLVATDVASKGLDFPDIQHVINYDMPGEI 429
Query: 486 DDYVHRIGRTGRAGKKGLATAFFNENNTSMA-RSLQDLMQEANQEVPAWLS 535
++YVHRIGRTGR GK G+AT F N+N + + L+ L+QEA Q +P L+
Sbjct: 430 ENYVHRIGRTGRCGKTGIATTFINKNQSEITLLDLKHLLQEAKQRIPPVLA 480
>At3g22330 DEAD-Box RNA helicase like protein
Length = 616
Score = 275 bits (702), Expect = 7e-74
Identities = 175/442 (39%), Positives = 260/442 (58%), Gaps = 36/442 (8%)
Query: 169 PVQRHAIPISLGGRDLMACAQTGSGKTAAFCFPIISGIMTGQPAQRPPRGVRTVCPLALV 228
P+Q+ + ++ GRD++ A+TG+GKT AF PII I+ R PL LV
Sbjct: 129 PIQKAVLEPAMEGRDMIGRARTGTGKTLAFGIPIIDKIIKYNAKHGRGRN-----PLCLV 183
Query: 229 LSPTRELSMQIHEEARKFSYQTGVRVVVAYGGAPINQQLRELERGVDILVATPGRLVDLL 288
L+PTREL+ Q+ +E R+ + + + YGG PI QQ+R+L+ GVD+ V TPGR++DL+
Sbjct: 184 LAPTRELARQVEKEFRESA--PSLDTICLYGGTPIGQQMRQLDYGVDVAVGTPGRVIDLM 241
Query: 289 ERARVSLSMIRYLALDEADRMLDMGFEPQIRKIVEQMDMPPAGVRQTMLFSATFPKEIQR 348
+R ++LS ++++ LDEAD+ML +GF + I+E++ RQ+M+FSAT P I+
Sbjct: 242 KRGALNLSEVQFVVLDEADQMLQVGFAEDVEIILEKLPEK----RQSMMFSATMPSWIRS 297
Query: 349 LASDFLSNYIFLAVGRVGSSTD-LIDQRVEY--VQESDKRSHLMDLLHAQRANGVQGKQA 405
L +L+N L V VG S L D Y + +S R+ ++ L + A G GK
Sbjct: 298 LTKKYLNNP--LTVDLVGDSDQKLADGITTYSIIADSYGRASIIGPLVTEHAKG--GK-- 351
Query: 406 LTLVFVETKKGADALEHWLCLNNFPATTIHGDRSQQEREAALRSFKSGNTPILVATDVAA 465
+VF +TK+ AD L + L +F +HGD SQ +RE L F+ G+ ILVATDVAA
Sbjct: 352 -CIVFTQTKRDADRLSYALA-RSFKCEALHGDISQSQRERTLAGFRDGHFNILVATDVAA 409
Query: 466 RGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRAGKKGLATAFFNENNTSMARSLQDLMQE 525
RGLD+P+V +++++LPN+ + +VHR GRTGRAGKKG A ++++ + + ++ +
Sbjct: 410 RGLDVPNVDLIIHYELPNNTETFVHRTGRTGRAGKKGSAILIYSQDQSRAVKIIEREVGS 469
Query: 526 ANQEVPAWLSRFAARSSFGGGKNRRSGGGRFGGRDFRREGSFSRGGSDYHGAGNSGGGY- 584
E+P+ + S F G +R GG FGG G RG S G + GGGY
Sbjct: 470 RFTELPSIAVERGSASMFEGIGSR--SGGSFGG------GMRDRGSS--FGGRSGGGGYG 519
Query: 585 GASGGYGGGGYGGG---YAGNS 603
G+SGGYGGG GG Y+G+S
Sbjct: 520 GSSGGYGGGRSGGSSNRYSGDS 541
Score = 35.0 bits (79), Expect = 0.12
Identities = 22/53 (41%), Positives = 28/53 (52%), Gaps = 4/53 (7%)
Query: 540 RSSFGGGKNRRSGGGRFGGRDFRREGSFSRGGSD---YHGAGNSGGGYGASGG 589
RSS GG++ GGGR GG R F GSD G +S GG+G++ G
Sbjct: 561 RSSQSGGRSS-FGGGRSGGSSNNRSSGFGDFGSDRSSQSGGRSSFGGFGSNDG 612
>At3g22310 putative RNA helicase
Length = 610
Score = 272 bits (695), Expect = 5e-73
Identities = 171/435 (39%), Positives = 247/435 (56%), Gaps = 21/435 (4%)
Query: 169 PVQRHAIPISLGGRDLMACAQTGSGKTAAFCFPIISGIMTGQPAQRPPRGVRTVCPLALV 228
P+Q+ + ++ GRD++ A+TG+GKT AF PII I+ + RG C LV
Sbjct: 141 PIQKAVLEPAMEGRDMIGRARTGTGKTLAFGIPIIDKIIKFNA--KHGRGKNPQC---LV 195
Query: 229 LSPTRELSMQIHEEARKFSYQTGVRVVVAYGGAPINQQLRELERGVDILVATPGRLVDLL 288
L+PTREL+ Q+ +E R+ + + + YGG PI QQ+REL G+D+ V TPGR++DL+
Sbjct: 196 LAPTRELARQVEKEFRESA--PSLDTICLYGGTPIGQQMRELNYGIDVAVGTPGRIIDLM 253
Query: 289 ERARVSLSMIRYLALDEADRMLDMGFEPQIRKIVEQMDMPPAGVRQTMLFSATFPKEIQR 348
+R ++LS ++++ LDEAD+ML +GF + I++++ PA RQ+M+FSAT P I+
Sbjct: 254 KRGALNLSEVQFVVLDEADQMLQVGFAEDVEIILQKL---PAK-RQSMMFSATMPSWIRS 309
Query: 349 LASDFLSNYIFLAVGRVGSSTD-LIDQRVEYVQESDKRSHLMDLLHAQRANGVQGKQALT 407
L +L+N L + VG S L D Y +D + + +G GK
Sbjct: 310 LTKKYLNNP--LTIDLVGDSDQKLADGITMYSIAADSYGRASIIGPLVKEHGKGGK---C 364
Query: 408 LVFVETKKGADALEHWLCLNNFPATTIHGDRSQQEREAALRSFKSGNTPILVATDVAARG 467
+VF +TK+ AD L L ++ +HGD SQ +RE L F+ GN ILVATDVAARG
Sbjct: 365 IVFTQTKRDADRLAFGLA-KSYKCEALHGDISQAQRERTLAGFRDGNFSILVATDVAARG 423
Query: 468 LDIPHVAHVVNFDLPNDIDDYVHRIGRTGRAGKKGLATAFFNENNTSMARSLQDLMQEAN 527
LD+P+V V++++LPN+ + +VHR GRTGRAGKKG A ++ T + ++ +
Sbjct: 424 LDVPNVDLVIHYELPNNTETFVHRTGRTGRAGKKGSAILIHGQDQTRAVKMIEKEVGSRF 483
Query: 528 QEVPAWLSRFAARSSFGGGKNRRSGGGRFGGRDFRREGSFSRGGSDYHGAGNSGGGYGAS 587
E+P+ + S F G R GG FGG G S G S G S GGYG S
Sbjct: 484 NELPSIAVERGSASMFEGVGAR--SGGSFGGGRSGGGGYGSYGSSSGRSGGGSYGGYGGS 541
Query: 588 GGYGGGGYGGGYAGN 602
G GGG GG Y G+
Sbjct: 542 SGRSGGG-GGSYGGS 555
>At1g28180 hypothetical protein
Length = 622
Score = 246 bits (629), Expect = 2e-65
Identities = 157/401 (39%), Positives = 219/401 (54%), Gaps = 26/401 (6%)
Query: 153 EALNQNIRRCKYVKPTPVQRHAIPISLGGRDLMACAQTGSGKTAAFCFPIISGIMTGQPA 212
E N + R K P IP+ L RD++ + TGSGKTAAF P+++ I P
Sbjct: 219 EDFNISYRGSKIPHPMRNWEETIPLGLEQRDVIGISATGSGKTAAFVLPMLAYISRLPPM 278
Query: 213 QRPPRGVRTVCPLALVLSPTRELSMQIHEEARKFSYQTGVRVVVAYGGAPINQQLRELER 272
+ + T P ALV+ PTREL+ QI EE KFS G + V G I +Q +L +
Sbjct: 279 REENQ---TEGPYALVMVPTRELAHQIEEETVKFSRYLGFKAVSITGWESIEKQALKLSQ 335
Query: 273 GVDILVATPGRLVDLLERARVSLSMIRYLALDEADRMLDMGFEPQIRKIVEQM---DMPP 329
G +I++ATPGRL+D LER V L+ YL LDEADRM+DM FEPQ+ ++++ M ++ P
Sbjct: 336 GCEIVIATPGRLLDCLERRYVVLNQCNYLVLDEADRMIDMDFEPQVSEVLDVMPCSNLKP 395
Query: 330 AG----------VRQTMLFSATFPKEIQRLASDFLSNYIFLAVGRVGSSTDLIDQRVEYV 379
R T +FSAT ++RLA FL N + V +G +T I Q+V
Sbjct: 396 EKEDEELEEKKIYRTTYMFSATMLLSVERLARKFLRNPV---VVTIGETTKFITQQVIMT 452
Query: 380 QESDKRSHLMDLLHAQRANGVQGKQALTLVFVETKKGAD-ALEHWLCLNNFPATTIHGDR 438
+ESDK S L L+ G +VFV T+ D +++ + TT+H +
Sbjct: 453 KESDKFSRLKKLIDD------LGDDKTAIVFVNTRNKVDYIVKNLEKVGRCRVTTLHAGK 506
Query: 439 SQQEREAALRSFKSGNTPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRA 498
SQ++R+ +L FK +LV TDV RGLDI +A V+N+D+PN +D Y HRIGRTGRA
Sbjct: 507 SQEQRDYSLEEFKKKRFNVLVTTDVLGRGLDILDLAQVINYDMPNTMDLYTHRIGRTGRA 566
Query: 499 GKKGLATAFFNENNTSMARSLQDLMQEANQEVPAWLSRFAA 539
GK G+AT F + + L+ + E N VP L+R A
Sbjct: 567 GKTGVATTFLTLEDKDVFYGLKQKLNECNSLVPPELARHEA 607
>At5g60990 replication protein A1 - like
Length = 456
Score = 234 bits (596), Expect = 1e-61
Identities = 154/449 (34%), Positives = 227/449 (50%), Gaps = 21/449 (4%)
Query: 143 VNTFAEIDLGEALNQNIRRCKYVKPTPVQRHAIPISLGGRDLMACAQTGSGKTAAFCFPI 202
V TFAE+ + E L + R + P+ +Q A+P +L G+D++ AQTGSGKT AF PI
Sbjct: 8 VKTFAELGVREELVKACERLGWKNPSKIQAEALPFALEGKDVIGLAQTGSGKTGAFAIPI 67
Query: 203 ISGIMTGQPAQRPPRGVRT-VCPLALVLSPTRELSMQIHEEARKFSYQTGVRVVVAYGGA 261
+ ++ P +G R A VLSPTREL++QI E+ +R V GG
Sbjct: 68 LQALLEYVYDSEPKKGRRPDPAFFACVLSPTRELAIQIAEQFEALGADISLRCAVLVGGI 127
Query: 262 PINQQLRELERGVDILVATPGRLVDLLERAR-VSLSMIRYLALDEADRMLDMGFEPQIRK 320
QQ L + ++VATPGRL D + + SL ++YL LDEADR+L+ FE + +
Sbjct: 128 DRMQQTIALGKRPHVIVATPGRLWDHMSDTKGFSLKSLKYLVLDEADRLLNEDFEKSLNQ 187
Query: 321 IVEQMDMPPAGVRQTMLFSATFPKEIQRLASDFLSNYIFLAVGRVGSSTDLIDQRVEYVQ 380
I+E++ + R+T LFSAT K++++L L N + + S+ D + Q+ +V
Sbjct: 188 ILEEIPLE----RKTFLFSATMTKKVRKLQRACLRNPVKIEAASKYSTVDTLKQQYRFVA 243
Query: 381 ESDKRSHLMDLLHAQRANGVQGKQALTLVFVETKKGADALEHWLCLNNFPATTIHGDRSQ 440
K +L+ +L ++ +++F T G L L F A I G +Q
Sbjct: 244 AKYKDCYLVYILSEM-------PESTSMIFTRTCDGTRFLALVLRSLGFRAIPISGQMTQ 296
Query: 441 QEREAALRSFKSGNTPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRAGK 500
+R AL FK+G ILV TDVA+RGLDIP V V+N+D+P + DY+HR+GRT RAG+
Sbjct: 297 SKRLGALNKFKAGECNILVCTDVASRGLDIPSVDVVINYDIPTNSKDYIHRVGRTARAGR 356
Query: 501 KGLATAFFNENNTSMARSLQDLMQEANQEVPA-------WLSRFAARSSFGGGKNRRSGG 553
G+ + N+ ++ L+ + E PA L R A + SGG
Sbjct: 357 SGVGISLVNQYELEWYIQIEKLIGKKLPEYPAEEDEVLSLLERVAEAKKLSAMNMKESGG 416
Query: 554 GRFGGRDFRREGSFSRGGSDYHGAGNSGG 582
+ G D F G D G GG
Sbjct: 417 RKRRGEDDEESERFLGGNKD-RGNKERGG 444
>At5g26742 unknown protein
Length = 475
Score = 233 bits (593), Expect = 3e-61
Identities = 146/379 (38%), Positives = 218/379 (56%), Gaps = 22/379 (5%)
Query: 147 AEIDLGEALNQNIRRCKYVKPTPVQRHAIPISLGGRDLMACAQTGSGKTAAFCFPIISGI 206
+++ L + L +++ + P+QR + +L GRD++A A+TG+GKT AF PII +
Sbjct: 105 SKLSLPQRLEESLEKRGITHLFPIQRAVLVPALQGRDIIARAKTGTGKTLAFGIPIIKRL 164
Query: 207 MTGQPAQRPPRGVRTVCPLALVLSPTRELSMQIHEEARKFSYQTGVRVVVAYGGAPINQQ 266
T + P LVL+PTREL+ Q+ +E ++ + + V YGG Q
Sbjct: 165 -TEEAGDYTAFRRSGRLPKFLVLAPTRELAKQVEKEIKESAPY--LSTVCVYGGVSYTIQ 221
Query: 267 LRELERGVDILVATPGRLVDLLERARVSLSMIRYLALDEADRMLDMGFEPQIRKIVEQMD 326
L RGVD++V TPGR++DL+E + L + YL LDEAD+ML +GFE + I+E +
Sbjct: 222 QSALTRGVDVVVGTPGRIIDLIEGRSLKLGEVEYLVLDEADQMLAVGFEEAVESILENLP 281
Query: 327 MPPAGVRQTMLFSATFPKEIQRLASDFLSNYIFLAVGRVGSSTDLIDQRVEY----VQES 382
RQ+MLFSAT P +++LA +L N L + VG + + + ++ +
Sbjct: 282 TK----RQSMLFSATMPTWVKKLARKYLDNP--LNIDLVGDQDEKLAEGIKLYAIATTST 335
Query: 383 DKRSHLMDLLHAQRANGVQGKQALTLVFVETKKGADALEHWLCLNNFPAT-TIHGDRSQQ 441
KR+ L DL+ V K T+VF +TK+ AD + L L+N AT +HGD SQ
Sbjct: 336 SKRTILSDLIT------VYAKGGKTIVFTQTKRDADEVS--LALSNSIATEALHGDISQH 387
Query: 442 EREAALRSFKSGNTPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRAGKK 501
+RE L +F+ G +LVATDVA+RGLDIP+V V++++LPND + +VHR GRTGRAGK+
Sbjct: 388 QRERTLNAFRQGKFTVLVATDVASRGLDIPNVDLVIHYELPNDPETFVHRSGRTGRAGKE 447
Query: 502 GLATAFFNENNTSMARSLQ 520
G A + RSL+
Sbjct: 448 GSAILMHTSSQKRTVRSLE 466
>At3g09720 RNA helicase like protein
Length = 541
Score = 223 bits (568), Expect = 2e-58
Identities = 141/441 (31%), Positives = 230/441 (51%), Gaps = 36/441 (8%)
Query: 108 SEEPVTQEE-------QENTGINFDAY--EDIPVETSGGNVPPPVNTFAEIDLGEALN-- 156
+EE +T+ E + N + DA + + SG N+PPP+ +FAE+
Sbjct: 92 AEEQITKNEIVENPKKELNRQMERDALSRKQYSIHVSGNNIPPPLKSFAELSSRYGCEGY 151
Query: 157 --QNIRRCKYVKPTPVQRHAIPISLGGRDLMACAQTGSGKTAAFCFPIISGIMTGQPAQR 214
+N+ + +PTP+QR AIPI L GR+ ACA TGSGKT AF P++ + +R
Sbjct: 152 ILRNLAELGFKEPTPIQRQAIPILLSGRECFACAPTGSGKTFAFICPMLIKL------KR 205
Query: 215 PPR-GVRTVCPLALVLSPTRELSMQIHEEARKFSYQTGVRVVVAYGGAPINQQLRELERG 273
P G+R A++LSP REL+ Q E +K G + P+ + +
Sbjct: 206 PSTDGIR-----AVILSPARELAAQTAREGKKLI--KGSNFHIRLMTKPLVKTADFSKLW 258
Query: 274 VDILVATPGRLVDLLERARVSLSMIRYLALDEADRMLDMGFEPQIRKIVEQMDMPPAGVR 333
D+L++TP RL ++ ++ LS + YL LDE+D++ + QI +V+ P +R
Sbjct: 259 CDVLISTPMRLKRAIKAKKIDLSKVEYLVLDESDKLFEQSLLKQIDCVVKACSNPSI-IR 317
Query: 334 QTMLFSATFPKEIQRLASDFLSNYIFLAVGRVGSSTDLIDQRVEYVQESDKRSHLMDLLH 393
LFSAT P ++ LA + + + + +GR ++++ + Q++ + + + L
Sbjct: 318 S--LFSATLPDSVEELARSIMHDAVRVIIGRKNTASETVKQKLVFAGSEEGK------LL 369
Query: 394 AQRANGVQGKQALTLVFVETKKGADALEHWLCLNNFPATTIHGDRSQQEREAALRSFKSG 453
A R + + L+FV++K+ A L L N A IH D ERE A+ F++G
Sbjct: 370 ALRQSFAESLNPPVLIFVQSKERAKELYDELKCENIRAGVIHSDLPPGERENAVDQFRAG 429
Query: 454 NTPILVATDVAARGLDIPHVAHVVNFDLPNDIDDYVHRIGRTGRAGKKGLATAFFNENNT 513
+L+ATDV ARG+D + V+N+D P+ Y+HRIGR+GRAG+ G A F+ E +
Sbjct: 430 EKWVLIATDVIARGMDFKGINCVINYDFPDSASAYIHRIGRSGRAGRSGEAITFYTEQDV 489
Query: 514 SMARSLQDLMQEANQEVPAWL 534
R++ + M + EVP+W+
Sbjct: 490 PFLRNIANTMMSSGCEVPSWI 510
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,259,349
Number of Sequences: 26719
Number of extensions: 790381
Number of successful extensions: 18998
Number of sequences better than 10.0: 421
Number of HSP's better than 10.0 without gapping: 305
Number of HSP's successfully gapped in prelim test: 124
Number of HSP's that attempted gapping in prelim test: 4496
Number of HSP's gapped (non-prelim): 5862
length of query: 612
length of database: 11,318,596
effective HSP length: 105
effective length of query: 507
effective length of database: 8,513,101
effective search space: 4316142207
effective search space used: 4316142207
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146760.13


