

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146759.10 + phase: 1 /partial
(178 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
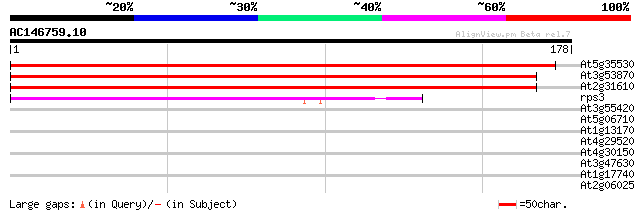
At5g35530 40S ribosomal protein S3 303 3e-83
At3g53870 ribosomal protein S3a homolog 298 1e-81
At2g31610 40S ribosomal protein; contains C-terminal domain 293 3e-80
rps3 -chloroplast genome- ribosomal protein S3 40 9e-04
At3g55420 unknown protein (At3g55420) 29 1.2
At5g06710 homeobox protein 27 4.4
At1g13170 putative oxysterol-binding protein 27 4.4
At4g29520 unknown protein 27 5.8
At4g30150 hypothetical protein 27 7.6
At3g47630 unknown protein 27 7.6
At1g17740 unknown protein 27 7.6
At2g06025 unknown protein 26 9.9
>At5g35530 40S ribosomal protein S3
Length = 248
Score = 303 bits (776), Expect = 3e-83
Identities = 150/173 (86%), Positives = 160/173 (91%)
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELTS+VQKRFKFP++SVELYAEKV NRGLCAIAQAESLRYKLLGGLAVRRACY
Sbjct: 61 EKGRRIRELTSLVQKRFKFPQDSVELYAEKVANRGLCAIAQAESLRYKLLGGLAVRRACY 120
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
GVLRFVMESGAKGCEVIVSGKLRA RAKSMKFKDGYM+SSGQP K+YID+AVRHVLLRQG
Sbjct: 121 GVLRFVMESGAKGCEVIVSGKLRAARAKSMKFKDGYMVSSGQPTKEYIDAAVRHVLLRQG 180
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVP 173
VLG+KVKIMLDWDPKGKQGP TPLPD+V IHTPKE++ YI PA VV VP
Sbjct: 181 VLGLKVKIMLDWDPKGKQGPMTPLPDVVIIHTPKEDDVYIAPAQVVTQAAFVP 233
>At3g53870 ribosomal protein S3a homolog
Length = 249
Score = 298 bits (762), Expect = 1e-81
Identities = 148/167 (88%), Positives = 155/167 (92%)
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELTS+VQKRFKFP +SVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY
Sbjct: 61 EKGRRIRELTSLVQKRFKFPVDSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 120
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
GVLRFVMESGAKGCEVIVSGKLRA RAKSMKFKDGYM+SSGQP K+YIDSAVRHVLLRQG
Sbjct: 121 GVLRFVMESGAKGCEVIVSGKLRAARAKSMKFKDGYMVSSGQPTKEYIDSAVRHVLLRQG 180
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVA 167
VLGIKVK+MLDWDPKG GPKTPLPD+V IH+PKEEE PA V A
Sbjct: 181 VLGIKVKVMLDWDPKGISGPKTPLPDVVIIHSPKEEEAIYAPAQVAA 227
>At2g31610 40S ribosomal protein; contains C-terminal domain
Length = 250
Score = 293 bits (750), Expect = 3e-80
Identities = 145/167 (86%), Positives = 153/167 (90%)
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELTS+VQKRFKFP +SVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY
Sbjct: 61 EKGRRIRELTSLVQKRFKFPVDSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 120
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
GVLRFVMESGAKGCEVIVSGKLRA RAKSMKFKDGYM+SSGQP K+YID+AVRHVLLRQG
Sbjct: 121 GVLRFVMESGAKGCEVIVSGKLRAARAKSMKFKDGYMVSSGQPTKEYIDAAVRHVLLRQG 180
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVA 167
VLGIKVKIMLDWDP GK GPKTPLPD+V IH PK++ Y PA A
Sbjct: 181 VLGIKVKIMLDWDPTGKSGPKTPLPDVVIIHAPKDDVVYSAPAQAAA 227
>rps3 -chloroplast genome- ribosomal protein S3
Length = 218
Score = 39.7 bits (91), Expect = 9e-04
Identities = 31/133 (23%), Positives = 64/133 (47%), Gaps = 5/133 (3%)
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
+K RR+ EL VQK + + +++N AE + +L ++ R+A
Sbjct: 87 DKPRRVEELQMNVQKELNCVNRKLNIAITRISNPYGDPNILAEFIAGQLKNRVSFRKAMK 146
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKF-KDGYM-ISSGQPVKDYIDSAVRHVLLR 118
+ ++ KG +V ++G++ + +++ ++G + + + + DY VR +
Sbjct: 147 KAIELTEQANTKGIQVQIAGRIDGKEIARVEWIREGRVPLQTIEAKIDYCSYTVRTIY-- 204
Query: 119 QGVLGIKVKIMLD 131
GVLGIK+ I +D
Sbjct: 205 -GVLGIKIWIFVD 216
>At3g55420 unknown protein (At3g55420)
Length = 216
Score = 29.3 bits (64), Expect = 1.2
Identities = 17/53 (32%), Positives = 26/53 (48%)
Query: 4 RRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVR 56
+ IR+L + RFKF + + Y V N G A AE L+ L +A++
Sbjct: 135 KMIRKLKNWEMARFKFRKGCITFYVYAVRNAGNEGFAAAEDLKVILQAVVALK 187
>At5g06710 homeobox protein
Length = 336
Score = 27.3 bits (59), Expect = 4.4
Identities = 38/156 (24%), Positives = 62/156 (39%), Gaps = 28/156 (17%)
Query: 14 QKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACYGVLRFVMESGAKG 73
+K+F F E + K+NN + + + +E K LG +G R + S +
Sbjct: 12 KKQFSFMEKN-----SKINNPSVSSTSTSE----KDLGFCMALDVAFGGHRSLSSSSSPS 62
Query: 74 CEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQGVLGIKVKIMLDWD 133
E K A RAK D + +SS +D ++ L + L +
Sbjct: 63 VED--EKKKPAPRAKK---SDEFRVSSS------VDPPLQ--------LQLHFPNWLPEN 103
Query: 134 PKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVAND 169
KG+QG + PL + +EEEE + +V D
Sbjct: 104 SKGRQGGRMPLGAATVVEEEEEEEEAVPSMSVSPPD 139
>At1g13170 putative oxysterol-binding protein
Length = 816
Score = 27.3 bits (59), Expect = 4.4
Identities = 41/206 (19%), Positives = 78/206 (36%), Gaps = 37/206 (17%)
Query: 2 KGRRIRELTSVVQKRFK--FPENSVELYAEKVNNRGLCAIAQAESLRYKLLG-------- 51
+GR+ + ++ + ++ +P+ + ++EKV++ + E + G
Sbjct: 506 EGRQCKPFNPLLGETYEADYPDKGLRFFSEKVSHHPMIVACHCEGQGWNFWGDSNIKGKF 565
Query: 52 -GLAVRRACYGVLRFVMESG--------AKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQ 102
G +++ GVL + G I+ GKL +M+ K G S
Sbjct: 566 WGRSIQLDPVGVLTLKFDDGEIYQWSKVTTSIYNIILGKLYCDHYGTMRIKGGSNYSCRL 625
Query: 103 PVKD--YIDSAVR--HVLLRQGVLGIKVKIML-DW---------DPKGKQGPKTPLPDIV 148
K+ ID R H ++ G KV I++ W DP K P+ + V
Sbjct: 626 KFKEQSVIDRNPRQVHGFVQDNRTGEKVAILIGKWDEAMYYVLGDPTTKPKGYDPMTEAV 685
Query: 149 TI----HTPKEEEEYIRPAAVVANDI 170
+ +P + + P A+ N+I
Sbjct: 686 LLWERDKSPTKTRYNLSPFAISLNEI 711
>At4g29520 unknown protein
Length = 306
Score = 26.9 bits (58), Expect = 5.8
Identities = 12/34 (35%), Positives = 20/34 (58%)
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNR 34
EK + EL + K FK +++ +A+KV+NR
Sbjct: 250 EKESKTEELKKTITKEFKKKGEALKRHAQKVSNR 283
>At4g30150 hypothetical protein
Length = 1966
Score = 26.6 bits (57), Expect = 7.6
Identities = 19/68 (27%), Positives = 32/68 (46%), Gaps = 4/68 (5%)
Query: 39 IAQAESLRYKLLGGLAVRRACYGVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMI 98
+ ++E KLL A+R A + ++ + E A GC L A +K+MK+
Sbjct: 697 LERSEKSVEKLLSSQALRLAIHKAIKVIPEGQASGC----IKSLTADVSKTMKWIKQVCC 752
Query: 99 SSGQPVKD 106
S+G +D
Sbjct: 753 STGATEQD 760
>At3g47630 unknown protein
Length = 320
Score = 26.6 bits (57), Expect = 7.6
Identities = 23/112 (20%), Positives = 47/112 (41%), Gaps = 17/112 (15%)
Query: 8 ELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLL--GGLAVRRACYGVL-- 63
++ +V+ +F ++ + + E+ + L + AE+ KL+ L+ R+ L
Sbjct: 179 KVNKIVKGQFDLFQSMYKPFLEECETKNLLRFSSAEASHTKLVQDSSLSATRSLVSSLPA 238
Query: 64 -------------RFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQ 102
+FV E+G EV +S + A + + M+SSG+
Sbjct: 239 SVRSQMGKSLGEKKFVSETGRVMGEVCISSREEAAKCMEKVMRRRVMVSSGR 290
>At1g17740 unknown protein
Length = 624
Score = 26.6 bits (57), Expect = 7.6
Identities = 36/140 (25%), Positives = 58/140 (40%), Gaps = 16/140 (11%)
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVR-RAC 59
+KG RI E VV ++P +S+++ V + A++ A G +++ +
Sbjct: 479 QKGLRISEERMVVDSSPEYPVDSIQVQILNVESNFAGAVSDA--------GDISIEGKVK 530
Query: 60 YGVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYM--ISSGQPVKDYIDSAVRHVLL 117
YGV G+ G +V + G L R G + I Q V S R VL
Sbjct: 531 YGVPHLTC-VGSFGVDVSLEGNLILCRQVDQPGMIGQVGNILGEQNVNVNFMSVGRTVLR 589
Query: 118 RQGVLGIKVKIMLDWDPKGK 137
+Q ++ I V D +P K
Sbjct: 590 KQAIMAIGV----DEEPDNK 605
>At2g06025 unknown protein
Length = 288
Score = 26.2 bits (56), Expect = 9.9
Identities = 18/48 (37%), Positives = 25/48 (51%), Gaps = 8/48 (16%)
Query: 29 EKVNNRGLCAIAQAESLRYKLLGGLAVRR-------ACYGVLRFVMES 69
EKV ++ C+I Q S RY + L V + AC +LRF +ES
Sbjct: 191 EKVKSQLFCSINQEGSNRYGYIANLCVAKSARRQGIAC-NMLRFAVES 237
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.138 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,883,811
Number of Sequences: 26719
Number of extensions: 159418
Number of successful extensions: 363
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 356
Number of HSP's gapped (non-prelim): 12
length of query: 178
length of database: 11,318,596
effective HSP length: 93
effective length of query: 85
effective length of database: 8,833,729
effective search space: 750866965
effective search space used: 750866965
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146759.10


