

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146751.11 - phase: 0
(399 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
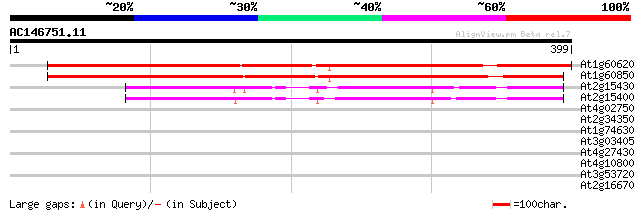
Score E
Sequences producing significant alignments: (bits) Value
At1g60620 RNA polymerase subunit (isoform B 475 e-134
At1g60850 RNA polymerase subunit 431 e-121
At2g15430 DNA-directed RNA polymerase II, third largest subunit 147 8e-36
At2g15400 DNA-directed RNA polymerase II, third largest subunit 143 2e-34
At4g02750 hypothetical protein 30 2.3
At2g34350 nodulin-like protein 30 3.1
At1g74630 hypothetical protein 30 3.1
At3g03405 putative protein 28 6.8
At4g27430 COP1-interacting protein 7 (CIP7) 28 8.9
At4g10800 unknown protein (At4g10800) 28 8.9
At3g53720 unknown protein 28 8.9
At2g16670 putative retroelement pol polyprotein 28 8.9
>At1g60620 RNA polymerase subunit (isoform B
Length = 385
Score = 475 bits (1223), Expect = e-134
Identities = 242/375 (64%), Positives = 295/375 (78%), Gaps = 15/375 (4%)
Query: 28 IMNLEDVPSTLPTHLELQKTRVFCNLDAPQHTDTIQYSGAYAALGVDNSVRLDNFYQNFK 87
I +L DVP+ LP HLELQ+TRV C D+ H I +SGAY+++GVDNSVRL+NF ++FK
Sbjct: 23 IFDLPDVPTGLPPHLELQRTRVVCKKDSNIHPTAITFSGAYSSMGVDNSVRLENFSEDFK 82
Query: 88 VEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRL 147
V+V +T+ +M FDMIG+ +ANAFRRIL+AE+P+MAIE+VY+ANNTS++QDEVL+HRL
Sbjct: 83 VDVISLTETDMVFDMIGVHAGIANAFRRILLAELPSMAIEKVYVANNTSVIQDEVLAHRL 142
Query: 148 GLIPIDADPKLFEYPDDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDTLKWLPNGSELI 207
GLIPI ADP+LFEY + + NEKNTIVFK HV+C KG PR V + LKWLPNGSELI
Sbjct: 143 GLIPIAADPRLFEYLSE-NDQPNEKNTIVFKLHVKCLKGDPRRKVLTSELKWLPNGSELI 201
Query: 208 AEGTKSAADPAPKTFTTFN--QKSLPKFSKDP-APYNLDVIVAKLGPGQEIELEAHAVKG 264
E S PKT+T+FN Q S P+F+++P P D+++AKLGPGQEIELEAHAVKG
Sbjct: 202 KESGGSTT--TPKTYTSFNHSQDSFPEFAENPIRPTLKDILIAKLGPGQEIELEAHAVKG 259
Query: 265 IGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVVK 324
IGKTHAKWSPVATAWYRMLPEVVL K+ + + AEELV CP KVFDIED+G+GRK+A V
Sbjct: 260 IGKTHAKWSPVATAWYRMLPEVVLLKEFEGKHAEELVKVCPKKVFDIEDMGQGRKRATVA 319
Query: 325 NARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEAV 384
R C+LCRECIR +G +W D+V LRRVK+HFIFTIESTG+ PP+VLF EAV
Sbjct: 320 RPRDCSLCRECIR---------DGVEWEDQVDLRRVKNHFIFTIESTGSQPPEVLFNEAV 370
Query: 385 KILEDKCERVITELS 399
KILEDKCERVI+ELS
Sbjct: 371 KILEDKCERVISELS 385
>At1g60850 RNA polymerase subunit
Length = 375
Score = 431 bits (1109), Expect = e-121
Identities = 225/371 (60%), Positives = 278/371 (74%), Gaps = 17/371 (4%)
Query: 28 IMNLEDVPSTLPTHLELQKTRVFCNLDAPQHTDTIQYSGAY-AALGVDNSVRLDNFYQNF 86
I +L DVP+ LP HL+ Q+TRV +AP HT + YSG Y ++ D++V+L NFY NF
Sbjct: 15 IDDLPDVPAGLPPHLKAQQTRVVSKNNAPAHTASAIYSGTYVSSTEEDDNVKLGNFYDNF 74
Query: 87 KVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHR 146
KV+V +T +MEFDMIGID A ANAFRRILIAEVP+MAIE+V IA NTS++ DEVL+HR
Sbjct: 75 KVDVVSLTKTDMEFDMIGIDAAFANAFRRILIAEVPSMAIEKVLIAYNTSVIIDEVLAHR 134
Query: 147 LGLIPIDADPKLFEYPDDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDTLKWLPNGSEL 206
+GLIPI ADP+LFEY + + NEKNTIVFK HV+C K +PRL V + LKWLPNGSEL
Sbjct: 135 MGLIPIAADPRLFEYLSEHD-QANEKNTIVFKLHVKCPKNRPRLKVLTSDLKWLPNGSEL 193
Query: 207 IAEGTKSAADPAPKTFTTFN--QKSLPKFSKDP-APYNLDVIVAKLGPGQEIELEAHAVK 263
+ E + P KT+T+F+ Q SLP+F+ +P P +LD+++AKL PGQEIELEAHAVK
Sbjct: 194 LRESENKTSKP--KTYTSFSCSQDSLPEFANNPITPCDLDILIAKLAPGQEIELEAHAVK 251
Query: 264 GIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVV 323
GIGKTHAKWSPV TAWYRM PEVVL +V+DELAE LV+ CP VFDIED+G+G+K+A V
Sbjct: 252 GIGKTHAKWSPVGTAWYRMHPEVVLRGEVEDELAERLVNVCPQNVFDIEDMGKGKKRATV 311
Query: 324 KNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEA 383
R CTLC+EC+R+ D D V L VK+HFIF IESTG++PP+VLFTEA
Sbjct: 312 AQPRKCTLCKECVRDDD----------LVDHVDLGSVKNHFIFNIESTGSLPPEVLFTEA 361
Query: 384 VKILEDKCERV 394
VKILE KCE +
Sbjct: 362 VKILEAKCEAI 372
>At2g15430 DNA-directed RNA polymerase II, third largest subunit
Length = 319
Score = 147 bits (372), Expect = 8e-36
Identities = 107/324 (33%), Positives = 171/324 (52%), Gaps = 45/324 (13%)
Query: 83 YQNF-KVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDE 141
YQ F K++++ + D+ +F++ D ++ANA RR++I+EVPT+AI+ V I N+S++ DE
Sbjct: 6 YQRFPKIKIRELKDDYAKFELRETDVSMANALRRVMISEVPTVAIDLVEIEVNSSVLNDE 65
Query: 142 VLSHRLGLIPIDADPKL---FEYPDDA--GGENNEKNTIVFKQHVRCEKGQPRLTVKSDT 196
++HRLGLIP+ ++ + F DA G E ++ F+ +C Q L V S
Sbjct: 66 FIAHRLGLIPLTSERAMSMRFSRDCDACDGDGQCEFCSVEFRLSSKCVTDQ-TLDVTSRD 124
Query: 197 LKWLPNGSELIAEGTKSAADP--APKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQE 254
L +ADP P FT + S + + +I+ KL GQE
Sbjct: 125 L---------------YSADPTVTPVDFTIDS-------SVSDSSEHKGIIIVKLRRGQE 162
Query: 255 IELEAHAVKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEE----LVSKCPAKVFD 310
++L A A KGIGK HAKWSP AT + P++++ +D+ D L++E L+ P KVF
Sbjct: 163 LKLRAIARKGIGKDHAKWSPAATVTFMYEPDIIINEDMMDTLSDEEKIDLIESSPTKVFG 222
Query: 311 IEDIGRGRKKAVVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIES 370
++ + R + VV + A T E I++A+ + G + + D FIFT+ES
Sbjct: 223 MDPVTR---QVVVVDPEAYTYDEEVIKKAEAMGKPG-------LIEISPKDDSFIFTVES 272
Query: 371 TGAIPPDVLFTEAVKILEDKCERV 394
TGA+ L A+ +L+ K + V
Sbjct: 273 TGAVKASQLVLNAIDLLKQKLDAV 296
>At2g15400 DNA-directed RNA polymerase II, third largest subunit
Length = 319
Score = 143 bits (360), Expect = 2e-34
Identities = 104/324 (32%), Positives = 167/324 (51%), Gaps = 45/324 (13%)
Query: 83 YQNFK-VEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDE 141
YQ F V+++ + D+ +F++ D ++ANA RR++I+EVPTMAI V I N+S++ DE
Sbjct: 6 YQRFPTVKIRELKDDYAKFELRETDVSMANALRRVMISEVPTMAIHLVKIEVNSSVLNDE 65
Query: 142 VLSHRLGLIPIDADPKLF-----EYPDDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDT 196
++ RL LIP+ ++ + + D G E+ E ++ F +C Q L V S
Sbjct: 66 FIAQRLSLIPLTSERAMSMRFCQDCEDCNGDEHCEFCSVEFPLSAKCVTDQ-TLDVTSRD 124
Query: 197 LKWLPNGSELIAEGTKSAADP--APKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQE 254
L +ADP P FT+ S + + +I+AKL GQE
Sbjct: 125 L---------------YSADPTVTPVDFTS-------NSSTSDSSEHKGIIIAKLRRGQE 162
Query: 255 IELEAHAVKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEE----LVSKCPAKVFD 310
++L+A A KGIGK HAKWSP AT Y P++++ +++ + L +E L+ P KVF
Sbjct: 163 LKLKALARKGIGKDHAKWSPAATVTYMYEPDIIINEEMMNTLTDEEKIDLIESSPTKVFG 222
Query: 311 IEDIGRGRKKAVVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIES 370
I+ + + VV + A T E I++A+ + G + + D F+FT+ES
Sbjct: 223 IDPV---TGQVVVVDPEAYTYDEEVIKKAEAMGKPG-------LIEIHPKHDSFVFTVES 272
Query: 371 TGAIPPDVLFTEAVKILEDKCERV 394
TGA+ L A+ IL+ K + +
Sbjct: 273 TGALKASQLVLNAIDILKQKLDAI 296
>At4g02750 hypothetical protein
Length = 781
Score = 30.0 bits (66), Expect = 2.3
Identities = 18/54 (33%), Positives = 27/54 (49%), Gaps = 3/54 (5%)
Query: 287 VLTKDVKDELAEELV---SKCPAKVFDIEDIGRGRKKAVVKNARACTLCRECIR 337
V+ DV++E E +V S+ A + I + GR V+KN R C C I+
Sbjct: 696 VVLHDVEEEEKERMVRYHSERLAVAYGIMRVSSGRPIRVIKNLRVCEDCHNAIK 749
>At2g34350 nodulin-like protein
Length = 2301
Score = 29.6 bits (65), Expect = 3.1
Identities = 19/69 (27%), Positives = 37/69 (53%), Gaps = 8/69 (11%)
Query: 243 DVIVAKLGPGQEIELEAHAVKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDE------- 295
D++V L G++ ++ + AVKG+ + ++S + ++ Y +LP L K++
Sbjct: 1944 DMVVGCLA-GEKPQMISAAVKGVARLTYEFSDLISSAYNLLPSTFLLLQRKNKEITKVRL 2002
Query: 296 LAEELVSKC 304
L E L+ KC
Sbjct: 2003 LLEMLIKKC 2011
>At1g74630 hypothetical protein
Length = 643
Score = 29.6 bits (65), Expect = 3.1
Identities = 30/89 (33%), Positives = 46/89 (50%), Gaps = 21/89 (23%)
Query: 255 IELEAHA-VKGIG---KTHAKWSP-VATAWYRMLPEVVLTKDVKDELAEELVSKCPAKV- 308
I++EAH +K I K A ++P VA+A Y DV++E E+ VSK K+
Sbjct: 531 IDIEAHEKLKEIILRLKDEAGYTPEVASALY----------DVEEEEKEDQVSKHSEKLA 580
Query: 309 --FDIEDIGRGRKKAVVKNARACTLCREC 335
F + + +G +VKN R +CR+C
Sbjct: 581 LAFALARLSKGANIRIVKNLR---ICRDC 606
>At3g03405 putative protein
Length = 193
Score = 28.5 bits (62), Expect = 6.8
Identities = 27/119 (22%), Positives = 48/119 (39%), Gaps = 12/119 (10%)
Query: 291 DVKDELAEELVSKCPAKVFDIED------------IGRGRKKAVVKNARACTLCRECIRE 338
D+ EL EE++S P K+ IG+ R +VK C+ C+ +
Sbjct: 6 DLPKELVEEIISGVPTKLMRAVRGDNTRLVVWNPYIGQTRSIKLVKGWDTCSYAFGCLEK 65
Query: 339 ADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEAVKILEDKCERVITE 397
+ + ++ D +L +D+F F P L E+ I E+K +++E
Sbjct: 66 NKKSCRSHKILRFLDDDTLYNGRDNFRFKFLKVNTGPEVSLVLESFIIDEEKKVVMVSE 124
>At4g27430 COP1-interacting protein 7 (CIP7)
Length = 1058
Score = 28.1 bits (61), Expect = 8.9
Identities = 25/83 (30%), Positives = 40/83 (48%), Gaps = 6/83 (7%)
Query: 290 KDVKDELAEELVSKCPAKVFDIEDIGRGRK---KAVVKNARACTLCRECIREADEGEEGG 346
K +DE AE S + + ED G+K K V++N T R +E+D E G
Sbjct: 352 KSKQDESAEP--SDNSSTETESEDGNEGKKQSRKVVIRNINYITSKRNGAKESDSD-ESG 408
Query: 347 EGEKWTDRVSLRRVKDHFIFTIE 369
E E + D S+++ + I ++E
Sbjct: 409 EEEGFVDGDSIKQQVEEAIGSVE 431
>At4g10800 unknown protein (At4g10800)
Length = 283
Score = 28.1 bits (61), Expect = 8.9
Identities = 29/102 (28%), Positives = 50/102 (48%), Gaps = 17/102 (16%)
Query: 53 LDAPQHTDTIQYSGAYAALGVD---NSVRLDNFYQNFKVEVKRITDEEMEFDMIGIDPAV 109
+D +T + S + A+ +D NS L + E+K E++EF M+ D V
Sbjct: 6 IDVSLNTKNVFLSALHFAMSIDTLKNSPELVD-------ELKTSAQEQVEF-MLSEDEEV 57
Query: 110 ANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRLGLIP 151
R+LI+E ++ R+ I+N S++ D LS L ++P
Sbjct: 58 -----RLLISEEEVKSVVRLGISNVISMLLDR-LSSLLVILP 93
>At3g53720 unknown protein
Length = 842
Score = 28.1 bits (61), Expect = 8.9
Identities = 40/159 (25%), Positives = 65/159 (40%), Gaps = 31/159 (19%)
Query: 201 PNGSELIAEGTKSAADPAPKT----FTTFNQKSLPKFSKDPAPY-----NLDVIVAKLGP 251
P+ E I G + A PA K F + PAP N + + P
Sbjct: 659 PDDRESIELGGRMAEHPAVKVTVIRFLVRETLRSTAVTLRPAPSKGKEKNYAFLTTNVDP 718
Query: 252 GQEIELEAHAVKGIGKTHAKWSPVATAWYRMLPEVVLTKDVK-DELAEELVSKCPAKVFD 310
+E EL+ A++ +KW E+V K+ + + + EE++S +K FD
Sbjct: 719 EKEKELDEGALEDF---KSKWK-----------EMVEYKEKEPNNIIEEILSIGQSKDFD 764
Query: 311 IEDIGRGRKKAVVKNARACTLCRECIREADEGEEGGEGE 349
+ +GRGR + +A L R+A+ E G G+
Sbjct: 765 LIVVGRGR----IPSAEVAALAE---RQAEHPELGPIGD 796
>At2g16670 putative retroelement pol polyprotein
Length = 1333
Score = 28.1 bits (61), Expect = 8.9
Identities = 23/101 (22%), Positives = 43/101 (41%), Gaps = 19/101 (18%)
Query: 89 EVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVY---------IANNTSLVQ 139
E R D+ EF + G+D A+ R L++ +P +E VY + N +++
Sbjct: 89 EKDREEDKVHEF-LSGLDDALFRTVRSSLVSRIPVQPLEEVYNIVRQEEDLLRNGANVLD 147
Query: 140 DEVLSHRLGLIPIDADPKLFEYPDDAGGENNEKNTIVFKQH 180
D+ + PKL++ G +EK+ + +H
Sbjct: 148 DQ---REVNAFAAQMRPKLYQ------GRGDEKDKSMVCKH 179
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.134 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,422,560
Number of Sequences: 26719
Number of extensions: 421784
Number of successful extensions: 1029
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 1009
Number of HSP's gapped (non-prelim): 15
length of query: 399
length of database: 11,318,596
effective HSP length: 101
effective length of query: 298
effective length of database: 8,619,977
effective search space: 2568753146
effective search space used: 2568753146
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146751.11


