

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146750.9 + phase: 0
(214 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
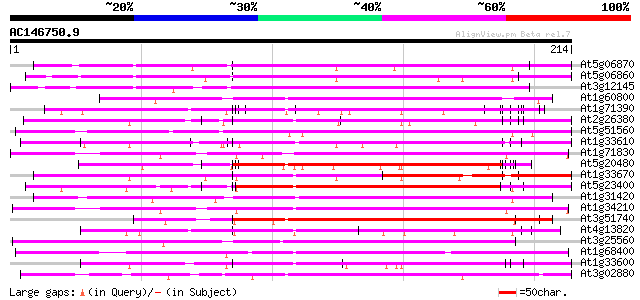
Score E
Sequences producing significant alignments: (bits) Value
At5g06870 polygalacturonase inhibiting protein 130 5e-31
At5g06860 polygalacturonase inhibiting protein 1; PGIP1 (gb|AAF6... 126 7e-30
At3g12145 leucine-rich repeat protein FLR1 114 4e-26
At1g60800 Putative Serine/Threonine protein kinase 113 7e-26
At1g71390 putative disease resistance protein 109 1e-24
At2g26380 putative disease resistance protein 108 2e-24
At5g51560 receptor-like protein kinase 106 8e-24
At1g33610 hypothetical protein 105 2e-23
At1g71830 protein kinase, putative 104 4e-23
At5g20480 receptor protein kinase - like 103 7e-23
At1g33670 disease resistance protein, putative 103 7e-23
At5g23400 disease resistance protein-like 103 9e-23
At1g31420 unknown protein 102 2e-22
At1g34210 putative protein 101 3e-22
At3g51740 unknown protein 100 4e-22
At4g13820 disease resistance protein - like 100 6e-22
At3g25560 unknown protein 100 7e-22
At1g68400 receptor kinase like protein 100 7e-22
At1g33600 unknown protein 100 7e-22
At3g02880 putative protein kinase 100 1e-21
>At5g06870 polygalacturonase inhibiting protein
Length = 330
Score = 130 bits (327), Expect = 5e-31
Identities = 77/190 (40%), Positives = 108/190 (56%), Gaps = 5/190 (2%)
Query: 10 LFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGCG- 68
L LS++ + S A D C+ DDK LLKI+ P L WD T+CC W + CG
Sbjct: 9 LLLSTLLLTTSLAKD--LCHKDDKTTLLKIKKSLNNPY-HLASWDPKTDCCSWYCLECGD 65
Query: 69 RPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNL 128
RVT + I G +SG +P E G+LPYL+ L ++ +TG I + +KL+ L L
Sbjct: 66 ATVNHRVTSLIIQDG-EISGQIPPEVGDLPYLTSLIFRKLTNLTGHIQPTIAKLKNLTFL 124
Query: 129 DLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
L +L+GP+P FL +LK L+ +DLS N LSG+IP+SL +L+ L +S N+L G IP
Sbjct: 125 RLSWTNLTGPVPEFLSQLKNLEYIDLSFNDLSGSIPSSLSSLRKLEYLELSRNKLTGPIP 184
Query: 189 AGLKKFNKNV 198
F+ V
Sbjct: 185 ESFGTFSGKV 194
Score = 82.8 bits (203), Expect = 1e-16
Identities = 58/178 (32%), Positives = 80/178 (44%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
LSG++P+ +L L L L+ K+TGPIP SF ++ +L L N LSG IP LG
Sbjct: 155 LSGSIPSSLSSLRKLEYLELSRN-KLTGPIPESFGTFSGKVPSLFLSHNQLSGTIPKSLG 213
Query: 145 K----------------------------------------------LKRLKEVDLSNNK 158
K L +D+++N
Sbjct: 214 NPDFYRIDLSRNKLQGDASILFGAKKTTWIVDISRNMFQFDLSKVKLAKTLNNLDMNHNG 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
++G+IPA NVS+N+LCG IP G +++F+ F HNKCLCGAPL CK
Sbjct: 274 ITGSIPAEWSKAY-FQLLNVSYNRLCGRIPKGEYIQRFDSYSFFHNKCLCGAPLPSCK 330
>At5g06860 polygalacturonase inhibiting protein 1; PGIP1
(gb|AAF69827.1)
Length = 330
Score = 126 bits (317), Expect = 7e-30
Identities = 78/189 (41%), Positives = 108/189 (56%), Gaps = 6/189 (3%)
Query: 7 LTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVG 66
L LLFL + F + S D CN +DK LLKI+ P L WD T+CC W +
Sbjct: 7 LCLLFLFT--FLTTCLSKDL-CNQNDKNTLLKIKKSLNNPY-HLASWDPQTDCCSWYCLE 62
Query: 67 CGRPYPG-RVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRL 125
CG RVT +TI G +SG +PAE G+LPYL L ++ +TG I + +KL+ L
Sbjct: 63 CGDATVNHRVTALTIFSGQ-ISGQIPAEVGDLPYLETLVFRKLSNLTGTIQPTIAKLKNL 121
Query: 126 QNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCG 185
+ L L +L+GPIP F+ +LK L+ ++LS N LSG+IP+SL L + +S N+L G
Sbjct: 122 RMLRLSWTNLTGPIPDFISQLKNLEFLELSFNDLSGSIPSSLSTLPKILALELSRNKLTG 181
Query: 186 AIPAGLKKF 194
+IP F
Sbjct: 182 SIPESFGSF 190
Score = 86.7 bits (213), Expect = 9e-18
Identities = 64/178 (35%), Positives = 80/178 (43%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
LSG++P+ LP + L L+ K+TG IP SF + +L L N LSGPIP LG
Sbjct: 155 LSGSIPSSLSTLPKILALELSRN-KLTGSIPESFGSFPGTVPDLRLSHNQLSGPIPKSLG 213
Query: 145 KLKRLKEVDLSNNKLSGT-----------------------------IPASLGNLQ---- 171
+ +DLS NKL G IP +LG L
Sbjct: 214 NID-FNRIDLSRNKLQGDASMLFGSNKTTWSIDLSRNMFQFDISKVDIPKTLGILDLNHN 272
Query: 172 -------------SLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
L FNVS+N+LCG IP G L+ F+ + HNKCLCGAPL CK
Sbjct: 273 GITGNIPVQWTEAPLQFFNVSYNKLCGHIPTGGKLQTFDSYSYFHNKCLCGAPLEICK 330
>At3g12145 leucine-rich repeat protein FLR1
Length = 203
Score = 114 bits (285), Expect = 4e-26
Identities = 74/199 (37%), Positives = 110/199 (55%), Gaps = 10/199 (5%)
Query: 1 MFLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECC 60
M L +HL++ F SI F +S + C +DK ALL+I+ G P L W+ T+CC
Sbjct: 1 MKLFVHLSIFF--SILFITLPSS--YSCTENDKNALLQIKKALGNPP-LLSSWNPRTDCC 55
Query: 61 D-WSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSF 119
W+ V C RVT ++++ G +SG + + G+L L L + +P +TG IP +
Sbjct: 56 TGWTGVECTNR---RVTGLSVTSG-EVSGQISYQIGDLVDLRTLDFSYLPHLTGNIPRTI 111
Query: 120 SKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVS 179
+KL+ L L L SLSGPIP ++ +LK L +DLS N+ +G IP SL + L ++
Sbjct: 112 TKLKNLNTLYLKHTSLSGPIPDYISELKSLTFLDLSFNQFTGPIPGSLSQMPKLEAIQIN 171
Query: 180 FNQLCGAIPAGLKKFNKNV 198
N+L G+IP F NV
Sbjct: 172 DNKLTGSIPNSFGSFVGNV 190
Score = 34.3 bits (77), Expect = 0.050
Identities = 22/78 (28%), Positives = 38/78 (48%), Gaps = 1/78 (1%)
Query: 123 QRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSN-NKLSGTIPASLGNLQSLSQFNVSFN 181
+R+ L + S +SG I +G L L+ +D S L+G IP ++ L++L+ +
Sbjct: 66 RRVTGLSVTSGEVSGQISYQIGDLVDLRTLDFSYLPHLTGNIPRTITKLKNLNTLYLKHT 125
Query: 182 QLCGAIPAGLKKFNKNVF 199
L G IP + + F
Sbjct: 126 SLSGPIPDYISELKSLTF 143
>At1g60800 Putative Serine/Threonine protein kinase
Length = 632
Score = 113 bits (283), Expect = 7e-26
Identities = 65/176 (36%), Positives = 101/176 (56%), Gaps = 12/176 (6%)
Query: 35 ALLKIRDHFGGPKGRLDDWD-NNTECCDWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAE 93
AL+ +++ P L++WD N+ + C W V C Y + + + S LSGTL
Sbjct: 38 ALVAVKNELNDPYKVLENWDVNSVDPCSWRMVSCTDGYVSSLDLPSQS----LSGTLSPR 93
Query: 94 FGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVD 153
GNL YL + L + +TGPIP + +L++LQ+LDL +NS +G IP+ LG+LK L +
Sbjct: 94 IGNLTYLQSVVL-QNNAITGPIPETIGRLEKLQSLDLSNNSFTGEIPASLGELKNLNYLR 152
Query: 154 LSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFE--HNKCLCG 207
L+NN L GT P SL ++ L+ ++S+N L G++P K + F+ N +CG
Sbjct: 153 LNNNSLIGTCPESLSKIEGLTLVDISYNNLSGSLP----KVSARTFKVIGNALICG 204
>At1g71390 putative disease resistance protein
Length = 784
Score = 109 bits (272), Expect = 1e-24
Identities = 74/212 (34%), Positives = 98/212 (45%), Gaps = 31/212 (14%)
Query: 14 SIYFS---KSNASDDF-FCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGCGR 69
+IYFS S AS FC D + LLK RD F + + W+ T+CC W V C
Sbjct: 14 TIYFSFLIHSLASPSLHFCRHDQRDGLLKFRDEFPIFESKSSPWNKTTDCCSWDGVTCD- 72
Query: 70 PYPGRVTVVTIS-------------------------RGWGLSGTLPAEFGNLPYLSMLS 104
G+V + + G L G +P+ GNL L L
Sbjct: 73 DKSGQVISLDLRSTLLNSSLKTNSSLFRLQYLRHLDLSGCNLHGEIPSSLGNLSRLENLE 132
Query: 105 LAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIP 164
L+ ++ G IP S L++L+NL LG N L G IPS LG L L ++DL NN L G +P
Sbjct: 133 LSS-NRLVGEIPYSIGNLKQLRNLSLGDNDLIGEIPSSLGNLSLLLDLDLWNNSLVGEVP 191
Query: 165 ASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
AS+GNL L ++ N L G+IP K
Sbjct: 192 ASIGNLNELRVMSLDRNSLSGSIPISFTNLTK 223
Score = 67.8 bits (164), Expect = 4e-12
Identities = 37/95 (38%), Positives = 52/95 (53%), Gaps = 6/95 (6%)
Query: 110 KVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGN 169
++ G IP S L+ L+ L+L N+ + IP L +L+ +DLS NKLSG IP LG
Sbjct: 609 RIYGEIPESIGCLEELRLLNLSGNAFTSDIPRVWENLTKLETLDLSRNKLSGQIPQDLGK 668
Query: 170 LQSLSQFNVSFNQLCGAIPAGLKKFNKNVFEHNKC 204
L LS N S N+L G +P G + F+ +C
Sbjct: 669 LSFLSYMNFSHNRLQGPVPRGTQ------FQRQRC 697
Score = 66.6 bits (161), Expect = 9e-12
Identities = 40/117 (34%), Positives = 64/117 (54%), Gaps = 2/117 (1%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIP-NSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
SG P ++P L+ +S+ + + +GPI + S +LQNL L N L G IP + K
Sbjct: 258 SGHFPKFLFSIPSLAWVSM-DRNQFSGPIEFANISSSSKLQNLILTRNKLDGSIPESISK 316
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFEHN 202
L +D+++N +SG +P S+ L SL F S N+L G +P+ L + + + HN
Sbjct: 317 FLNLVLLDVAHNNISGPVPRSMSKLVSLRIFGFSNNKLEGEVPSWLWRLSSTMLSHN 373
Score = 60.5 bits (145), Expect = 7e-10
Identities = 36/93 (38%), Positives = 49/93 (51%), Gaps = 2/93 (2%)
Query: 91 PAEFGNLPYLSMLS--LAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKR 148
P EF N+ S L + K+ G IP S SK L LD+ N++SGP+P + KL
Sbjct: 284 PIEFANISSSSKLQNLILTRNKLDGSIPESISKFLNLVLLDVAHNNISGPVPRSMSKLVS 343
Query: 149 LKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFN 181
L+ SNNKL G +P+ L L S + SF+
Sbjct: 344 LRIFGFSNNKLEGEVPSWLWRLSSTMLSHNSFS 376
Score = 56.6 bits (135), Expect = 9e-09
Identities = 38/130 (29%), Positives = 59/130 (45%), Gaps = 21/130 (16%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L G++P L +L +A ++GP+P S SKL L+ +N L G +PS+L +
Sbjct: 306 LDGSIPESISKFLNLVLLDVAHN-NISGPVPRSMSKLVSLRIFGFSNNKLEGEVPSWLWR 364
Query: 146 LKR--------------------LKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCG 185
L ++ +DLS N GT P + L+ L ++S N G
Sbjct: 365 LSSTMLSHNSFSSFEKIYSKETMIQVLDLSFNSFRGTFPVWICKLKGLHFLDLSNNLFNG 424
Query: 186 AIPAGLKKFN 195
+IP L+ FN
Sbjct: 425 SIPLCLRNFN 434
Score = 55.1 bits (131), Expect = 3e-08
Identities = 40/133 (30%), Positives = 61/133 (45%), Gaps = 25/133 (18%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNL----------------- 128
L G +PA GNL L ++SL + ++G IP SF+ L +L
Sbjct: 186 LVGEVPASIGNLNELRVMSL-DRNSLSGSIPISFTNLTKLSEFRIFFNNFTSLPSDLSGF 244
Query: 129 ------DLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIP-ASLGNLQSLSQFNVSFN 181
D+ +NS SG P FL + L V + N+ SG I A++ + L ++ N
Sbjct: 245 HNLVTFDISANSFSGHFPKFLFSIPSLAWVSMDRNQFSGPIEFANISSSSKLQNLILTRN 304
Query: 182 QLCGAIPAGLKKF 194
+L G+IP + KF
Sbjct: 305 KLDGSIPESISKF 317
Score = 50.4 bits (119), Expect = 7e-07
Identities = 27/78 (34%), Positives = 41/78 (51%)
Query: 110 KVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGN 169
K +G +P+ F+ LQ+LD+ N L G P L K L V++ +NK+ T P+ LG+
Sbjct: 444 KFSGTLPDIFANNTNLQSLDVSGNQLEGKFPKSLINCKGLHFVNVESNKIKDTFPSWLGS 503
Query: 170 LQSLSQFNVSFNQLCGAI 187
L SL + N G +
Sbjct: 504 LPSLQVLILRSNDFYGPL 521
Score = 49.3 bits (116), Expect = 2e-06
Identities = 38/123 (30%), Positives = 57/123 (45%), Gaps = 22/123 (17%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIP-------------NSFSKLQRL------- 125
+SG +P L L + + K+ G +P NSFS +++
Sbjct: 330 ISGPVPRSMSKLVSLRIFGFSNN-KLEGEVPSWLWRLSSTMLSHNSFSSFEKIYSKETMI 388
Query: 126 QNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCG 185
Q LDL NS G P ++ KLK L +DLSNN +G+IP L N +L+ + N+ G
Sbjct: 389 QVLDLSFNSFRGTFPVWICKLKGLHFLDLSNNLFNGSIPLCLRNF-NLTGLILGNNKFSG 447
Query: 186 AIP 188
+P
Sbjct: 448 TLP 450
Score = 47.8 bits (112), Expect = 4e-06
Identities = 34/104 (32%), Positives = 45/104 (42%), Gaps = 2/104 (1%)
Query: 88 GTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLK 147
GT P L L L L+ G IP L L LG+N SG +P
Sbjct: 400 GTFPVWICKLKGLHFLDLSNN-LFNGSIPLCLRNFN-LTGLILGNNKFSGTLPDIFANNT 457
Query: 148 RLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
L+ +D+S N+L G P SL N + L NV N++ P+ L
Sbjct: 458 NLQSLDVSGNQLEGKFPKSLINCKGLHFVNVESNKIKDTFPSWL 501
Score = 39.7 bits (91), Expect = 0.001
Identities = 33/116 (28%), Positives = 53/116 (45%), Gaps = 15/116 (12%)
Query: 77 VVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLS 136
++ IS G SG LP F + + M++L + S+ ++ +QN L S+
Sbjct: 535 IIDISHN-GFSGVLPPNFFS-SWREMITL---------VHGSYEYIEDIQNYSLIYRSME 583
Query: 137 ----GPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
G SF + + +D S N++ G IP S+G L+ L N+S N IP
Sbjct: 584 MVNKGVEMSFERIRQDFRAIDFSENRIYGEIPESIGCLEELRLLNLSGNAFTSDIP 639
Score = 36.6 bits (83), Expect = 0.010
Identities = 26/81 (32%), Positives = 38/81 (46%), Gaps = 3/81 (3%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPI--PSFL 143
L G P N L +++ E K+ P+ L LQ L L SN GP+ PS
Sbjct: 469 LEGKFPKSLINCKGLHFVNV-ESNKIKDTFPSWLGSLPSLQVLILRSNDFYGPLYHPSMS 527
Query: 144 GKLKRLKEVDLSNNKLSGTIP 164
+ L+ +D+S+N SG +P
Sbjct: 528 IGFQGLRIIDISHNGFSGVLP 548
>At2g26380 putative disease resistance protein
Length = 480
Score = 108 bits (271), Expect = 2e-24
Identities = 64/192 (33%), Positives = 95/192 (49%), Gaps = 8/192 (4%)
Query: 6 HLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFG-GPKGRLDDWDNNTECCDWSF 64
H +F + I+ N + C+ DD+A LL + P G L W T+CC W+
Sbjct: 7 HNFFIFTAVIFLRCLNPTAAATCHPDDEAGLLAFKSGITKDPSGILSTWKKGTDCCSWNG 66
Query: 65 VGCGRPYPGRVTVVTI-----SRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSF 119
V C P RV V+TI G LSGT+ L +L + + +TGP P
Sbjct: 67 VSC--PNGNRVVVLTIRIESDDAGIFLSGTISPSLAKLQHLEGVVFINLKNITGPFPPFL 124
Query: 120 SKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVS 179
+L L+ + L + LSGP+P+ +G L RL + + N+ G+IP+S+ NL L+ N+
Sbjct: 125 FRLPHLKYVYLENTRLSGPLPANIGALNRLDTLTVKGNRFIGSIPSSISNLTRLNYLNLG 184
Query: 180 FNQLCGAIPAGL 191
N L G IP G+
Sbjct: 185 GNLLTGTIPLGI 196
Score = 71.2 bits (173), Expect = 4e-13
Identities = 41/104 (39%), Positives = 61/104 (58%), Gaps = 2/104 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQR-LQNLDLGSNSLSGPIPSFLG 144
LSGT+P F ++ L +L+L+ + +G +P S + L L L+LG N+LSG IPS+L
Sbjct: 212 LSGTIPDIFKSMTNLRILTLSRN-RFSGKLPPSIASLAPVLAFLELGQNNLSGSIPSYLS 270
Query: 145 KLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
+ L +DLS N+ SG +P SL L ++ N+S N L P
Sbjct: 271 RFVALDTLDLSKNRFSGAVPKSLAKLTKIANINLSHNLLTNPFP 314
Score = 66.2 bits (160), Expect = 1e-11
Identities = 41/122 (33%), Positives = 67/122 (54%), Gaps = 3/122 (2%)
Query: 74 RVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSN 133
R+ +T+ +G G++P+ NL L+ L+L +TG IP + L+ + NL+L N
Sbjct: 153 RLDTLTV-KGNRFIGSIPSSISNLTRLNYLNLGGN-LLTGTIPLGIANLKLISNLNLDGN 210
Query: 134 SLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQF-NVSFNQLCGAIPAGLK 192
LSG IP + L+ + LS N+ SG +P S+ +L + F + N L G+IP+ L
Sbjct: 211 RLSGTIPDIFKSMTNLRILTLSRNRFSGKLPPSIASLAPVLAFLELGQNNLSGSIPSYLS 270
Query: 193 KF 194
+F
Sbjct: 271 RF 272
Score = 65.5 bits (158), Expect = 2e-11
Identities = 39/112 (34%), Positives = 61/112 (53%), Gaps = 2/112 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+GT+P NL +S L+L + +++G IP+ F + L+ L L N SG +P +
Sbjct: 188 LTGTIPLGIANLKLISNLNL-DGNRLSGTIPDIFKSMTNLRILTLSRNRFSGKLPPSIAS 246
Query: 146 LKR-LKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
L L ++L N LSG+IP+ L +L ++S N+ GA+P L K K
Sbjct: 247 LAPVLAFLELGQNNLSGSIPSYLSRFVALDTLDLSKNRFSGAVPKSLAKLTK 298
Score = 60.1 bits (144), Expect = 9e-10
Identities = 38/111 (34%), Positives = 57/111 (51%), Gaps = 28/111 (25%)
Query: 127 NLDLGSNSLSGPIPSFLGKLKRLKE-----------------------VDLSNNKLSGTI 163
++DL N +SG FL ++L+E +DLS N + G +
Sbjct: 374 SIDLSDNEISGSPLRFLKGAEQLREFRMSGNKLRFDLRKLSFSTTLETLDLSRNLVFGKV 433
Query: 164 PASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFEHNKCLCGAPLAPCK 214
PA + L++L N+S N LCG +P + KF ++VF N CLCG+PL+ CK
Sbjct: 434 PARVAGLKTL---NLSQNHLCGKLP--VTKFPESVFAGNDCLCGSPLSHCK 479
>At5g51560 receptor-like protein kinase
Length = 680
Score = 106 bits (265), Expect = 8e-24
Identities = 80/260 (30%), Positives = 127/260 (48%), Gaps = 54/260 (20%)
Query: 3 LLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDW 62
+LLHL LL L ++FS ++ D+ A L++++ L W N + C
Sbjct: 6 MLLHLLLLPLLFLHFSNQVMAEI----TDELATLMEVKTELDPEDKHLASWSVNGDLCK- 60
Query: 63 SFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSL------AEMPK------ 110
F G G + GRV+ +++ +G GLSG + G L +L+ L L ++P+
Sbjct: 61 DFEGVGCDWKGRVSNISL-QGKGLSGKISPNIGKLKHLTGLFLHYNALVGDIPRELGNLS 119
Query: 111 -----------VTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKL 159
++G IP++ K+Q LQ L L N+L+G IP L L++L + L +NKL
Sbjct: 120 ELTDLYLNVNNLSGEIPSNIGKMQGLQVLQLCYNNLTGSIPRELSSLRKLSVLALQSNKL 179
Query: 160 SGTIPASLGNLQSLSQFNVSFNQLCGAIPAG------------------------LKKFN 195
+G IPASLG+L +L + ++S+N L G++P LK+ N
Sbjct: 180 TGAIPASLGDLSALERLDLSYNHLFGSVPGKLASPPLLRVLDIRNNSLTGNVPPVLKRLN 239
Query: 196 KNV-FEHNKCLCGAPLAPCK 214
+ FE+N LCGA +P K
Sbjct: 240 EGFSFENNLGLCGAEFSPLK 259
>At1g33610 hypothetical protein
Length = 907
Score = 105 bits (261), Expect = 2e-23
Identities = 68/194 (35%), Positives = 95/194 (48%), Gaps = 9/194 (4%)
Query: 5 LHLTLLFLSSIYFSKSNASDDFF-CNADDKAALLKIRDHFG-GPKGRLDDWDNNTECCDW 62
L TL S I F + +S C+ DD+A LL + P G L W T CC W
Sbjct: 4 LSFTLFIFSVITFLQCLSSTGAATCHPDDEAGLLAFKSGITQDPSGMLSSWKKGTSCCSW 63
Query: 63 SFVGCGRPYPGRVTVVTI-----SRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPN 117
+ C RVT++ + LSGTL L +LS++SL +TG P
Sbjct: 64 KGIICFNS--DRVTMLELVGFPKKPERSLSGTLSPSLAKLQHLSVISLGGHVNITGSFPK 121
Query: 118 SFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFN 177
+L +L+ +D+ +N LSGP+P+ +G L L+E+ L NK +G IP S+ NL LS
Sbjct: 122 FLLQLPKLRYVDIQNNRLSGPLPANIGVLSLLEEIFLQGNKFTGPIPNSISNLTRLSYLI 181
Query: 178 VSFNQLCGAIPAGL 191
N L G IP G+
Sbjct: 182 FGGNLLTGTIPLGI 195
Score = 100 bits (250), Expect = 4e-22
Identities = 57/167 (34%), Positives = 88/167 (52%), Gaps = 8/167 (4%)
Query: 28 CNADDKAALLKIRDHFG-GPKGRLDDWDNNTECCDWSFVGCGRPYPGRVTVVTISRGWGL 86
C+ DD+A LL + P G L W T+CC WS V C RVT +++ + L
Sbjct: 480 CDPDDEAGLLGFKSGITKDPSGILSSWKKGTDCCFWSGVFCVNN--DRVTQLSVDGDFSL 537
Query: 87 -----SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPS 141
SGT+ L +L + L + K+TGP P +L +L +++ LSGP+P+
Sbjct: 538 DGNSPSGTISPMLAKLQHLERILLTSLRKITGPFPQFIFRLPKLNYINIQGCLLSGPLPA 597
Query: 142 FLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
+G+L +LK + + N +G IP+S+ NL L+ N+ N+L G IP
Sbjct: 598 NIGELSQLKTLVIDGNMFTGHIPSSIANLTRLTWLNLGNNRLSGTIP 644
Score = 66.6 bits (161), Expect = 9e-12
Identities = 43/111 (38%), Positives = 62/111 (55%), Gaps = 3/111 (2%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
LSGT+P F ++ L+ L L+ G +P S + L L LDL N+LSG IP++L
Sbjct: 639 LSGTIPNIFKSMKELNSLDLSRNG-FFGRLPPSIASLAPTLYYLDLSQNNLSGTIPNYLS 697
Query: 145 KLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFN 195
+ + L + LS NK SG +P S NL +++ ++S N L G P LK N
Sbjct: 698 RFEALSTLVLSKNKYSGVVPMSFTNLINITNLDLSHNLLTGPFPV-LKSIN 747
Score = 66.2 bits (160), Expect = 1e-11
Identities = 43/149 (28%), Positives = 73/149 (48%), Gaps = 31/149 (20%)
Query: 70 PYPGRVTVVT-----ISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQR 124
P P ++ +T I G L+GT+P NL + L L + +++G IP+ F ++
Sbjct: 166 PIPNSISNLTRLSYLIFGGNLLTGTIPLGIANLKLMQNLQLGDN-RLSGTIPDIFESMKL 224
Query: 125 LQNLDLGSN-------------------------SLSGPIPSFLGKLKRLKEVDLSNNKL 159
L+ LDL SN +LSG IP+++ + +L+++DLS N+
Sbjct: 225 LKFLDLSSNEFYGKLPLSIATLAPTLLALQVSQNNLSGAIPNYISRFNKLEKLDLSKNRF 284
Query: 160 SGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
SG +P NL +++ ++S N L G P
Sbjct: 285 SGVVPQGFVNLTNINNLDLSHNLLTGQFP 313
Score = 61.6 bits (148), Expect = 3e-10
Identities = 45/134 (33%), Positives = 67/134 (49%), Gaps = 21/134 (15%)
Query: 84 WGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFL 143
W L+GT Y + L+E +++G S+ + L N L L
Sbjct: 791 WKLAGTY--------YYDSIDLSEN-EISGSPAKFLSQXKYLMEFRAAGNKLRFD----L 837
Query: 144 GKL---KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFE 200
GKL + L+ +DLS N + G + A+ L+++ NVS N LCG +P + KF + F
Sbjct: 838 GKLTFVRTLETLDLSRNLIFGRVLATFAGLKTM---NVSQNHLCGKLP--VTKFPASXFA 892
Query: 201 HNKCLCGAPLAPCK 214
N CLCG+PL+PCK
Sbjct: 893 GNDCLCGSPLSPCK 906
Score = 38.9 bits (89), Expect = 0.002
Identities = 28/95 (29%), Positives = 41/95 (42%), Gaps = 28/95 (29%)
Query: 128 LDLGSNSLSGPIPSFLGKL-----------------------KRLKEVDLSNNKLSGTIP 164
+DL N +SG + FL + + LK +DLS N + G +P
Sbjct: 373 IDLSKNEISGSLERFLNETRYLLEFRAAENKLRFDMGNLTFPRTLKTLDLSRNLVFGKVP 432
Query: 165 ASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVF 199
++ LQ L N+S N LCG +P KF + F
Sbjct: 433 VTVAGLQRL---NLSQNHLCGELPT--TKFPASAF 462
>At1g71830 protein kinase, putative
Length = 625
Score = 104 bits (259), Expect = 4e-23
Identities = 79/221 (35%), Positives = 109/221 (48%), Gaps = 25/221 (11%)
Query: 1 MFLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNN-TEC 59
+F+LL L LL S++ + +N D AL +R P L WD
Sbjct: 7 VFILLSLILLPNHSLWLASANLEGD---------ALHTLRVTLVDPNNVLQSWDPTLVNP 57
Query: 60 CDWSFVGCGRPYPGRVTVVTISRGWG-LSGTLPAEFG---NLPYLSMLSLAEMPKVTGPI 115
C W V C +V+ + G LSG L E G NL YL + S +TGPI
Sbjct: 58 CTWFHVTCNNEN----SVIRVDLGNAELSGHLVPELGVLKNLQYLELYS----NNITGPI 109
Query: 116 PNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQ 175
P++ L L +LDL NS SGPIP LGKL +L+ + L+NN L+G+IP SL N+ +L
Sbjct: 110 PSNLGNLTNLVSLDLYLNSFSGPIPESLGKLSKLRFLRLNNNSLTGSIPMSLTNITTLQV 169
Query: 176 FNVSFNQLCGAIP--AGLKKFNKNVFEHNKCLCGAPLA-PC 213
++S N+L G++P F F +N LCG + PC
Sbjct: 170 LDLSNNRLSGSVPDNGSFSLFTPISFANNLDLCGPVTSHPC 210
>At5g20480 receptor protein kinase - like
Length = 1031
Score = 103 bits (257), Expect = 7e-23
Identities = 68/199 (34%), Positives = 103/199 (51%), Gaps = 29/199 (14%)
Query: 27 FCNADDKAALLKIRDHFGGPKGR--LDDWDNNTECCDWSFVGCGRPYPGRVTVVTISRG- 83
F N D ALL+ + R L W++++ C+W V CGR R V++++ G
Sbjct: 26 FSNETDMQALLEFKSQVSENNKREVLASWNHSSPFCNWIGVTCGRR---RERVISLNLGG 82
Query: 84 WGLSGTLPAEFGNLPYLSMLSLAE------MPK-----------------VTGPIPNSFS 120
+ L+G + GNL +L +L+LA+ +P+ + G IP+S S
Sbjct: 83 FKLTGVISPSIGNLSFLRLLNLADNSFGSTIPQKVGRLFRLQYLNMSYNLLEGRIPSSLS 142
Query: 121 KLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSF 180
RL +DL SN L +PS LG L +L +DLS N L+G PASLGNL SL + + ++
Sbjct: 143 NCSRLSTVDLSSNHLGHGVPSELGSLSKLAILDLSKNNLTGNFPASLGNLTSLQKLDFAY 202
Query: 181 NQLCGAIPAGLKKFNKNVF 199
NQ+ G IP + + + VF
Sbjct: 203 NQMRGEIPDEVARLTQMVF 221
Score = 82.8 bits (203), Expect = 1e-16
Identities = 46/103 (44%), Positives = 65/103 (62%), Gaps = 1/103 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
+SGT+P + GNL L LSL E ++G +P SF KL LQ +DL SN++SG IPS+ G
Sbjct: 381 ISGTIPHDIGNLVSLQELSL-ETNMLSGELPVSFGKLLNLQVVDLYSNAISGEIPSYFGN 439
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
+ RL+++ L++N G IP SLG + L + N+L G IP
Sbjct: 440 MTRLQKLHLNSNSFHGRIPQSLGRCRYLLDLWMDTNRLNGTIP 482
Score = 72.8 bits (177), Expect = 1e-13
Identities = 42/108 (38%), Positives = 61/108 (55%), Gaps = 1/108 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG LP FG L L ++ L ++G IP+ F + RLQ L L SNS G IP LG+
Sbjct: 405 LSGELPVSFGKLLNLQVVDLYSNA-ISGEIPSYFGNMTRLQKLHLNSNSFHGRIPQSLGR 463
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
+ L ++ + N+L+GTIP + + SL+ ++S N L G P + K
Sbjct: 464 CRYLLDLWMDTNRLNGTIPQEILQIPSLAYIDLSNNFLTGHFPEEVGK 511
Score = 70.9 bits (172), Expect = 5e-13
Identities = 40/106 (37%), Positives = 54/106 (50%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L G LPA NL ++G IP+ L LQ L L +N LSG +P GK
Sbjct: 356 LGGELPASIANLSTTLTSLFLGQNLISGTIPHDIGNLVSLQELSLETNMLSGELPVSFGK 415
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
L L+ VDL +N +SG IP+ GN+ L + +++ N G IP L
Sbjct: 416 LLNLQVVDLYSNAISGEIPSYFGNMTRLQKLHLNSNSFHGRIPQSL 461
Score = 64.3 bits (155), Expect = 5e-11
Identities = 40/103 (38%), Positives = 54/103 (51%), Gaps = 2/103 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+G P E G L L L A K++G +P + ++ L + NS G IP + +
Sbjct: 501 LTGHFPEEVGKLELLVGLG-ASYNKLSGKMPQAIGGCLSMEFLFMQGNSFDGAIPD-ISR 558
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
L LK VD SNN LSG IP L +L SL N+S N+ G +P
Sbjct: 559 LVSLKNVDFSNNNLSGRIPRYLASLPSLRNLNLSMNKFEGRVP 601
Score = 60.5 bits (145), Expect = 7e-10
Identities = 42/145 (28%), Positives = 68/145 (45%), Gaps = 27/145 (18%)
Query: 74 RVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSN 133
++ ++ +S+ L+G PA GNL L L A ++ G IP+ ++L ++ + N
Sbjct: 170 KLAILDLSKN-NLTGNFPASLGNLTSLQKLDFA-YNQMRGEIPDEVARLTQMVFFQIALN 227
Query: 134 SLSGPIP-----------------SFLGKLK--------RLKEVDLSNNKLSGTIPASLG 168
S SG P SF G L+ L+ + L N+ +G IP +L
Sbjct: 228 SFSGGFPPALYNISSLESLSLADNSFSGNLRADFGYLLPNLRRLLLGTNQFTGAIPKTLA 287
Query: 169 NLQSLSQFNVSFNQLCGAIPAGLKK 193
N+ SL +F++S N L G+IP K
Sbjct: 288 NISSLERFDISSNYLSGSIPLSFGK 312
Score = 57.4 bits (137), Expect = 6e-09
Identities = 38/114 (33%), Positives = 55/114 (47%), Gaps = 6/114 (5%)
Query: 86 LSGTLPAEFGNLPYLSML-----SLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIP 140
LSG++P FG L L L SL + + +L+ LD+G N L G +P
Sbjct: 302 LSGSIPLSFGKLRNLWWLGIRNNSLGNNSSSGLEFIGAVANCTQLEYLDVGYNRLGGELP 361
Query: 141 SFLGKLKR-LKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
+ + L L + L N +SGTIP +GNL SL + ++ N L G +P K
Sbjct: 362 ASIANLSTTLTSLFLGQNLISGTIPHDIGNLVSLQELSLETNMLSGELPVSFGK 415
Score = 57.4 bits (137), Expect = 6e-09
Identities = 36/116 (31%), Positives = 60/116 (51%), Gaps = 2/116 (1%)
Query: 74 RVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSN 133
R++ V +S L +P+E G+L L++L L++ +TG P S L LQ LD N
Sbjct: 146 RLSTVDLSSNH-LGHGVPSELGSLSKLAILDLSKN-NLTGNFPASLGNLTSLQKLDFAYN 203
Query: 134 SLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPA 189
+ G IP + +L ++ ++ N SG P +L N+ SL +++ N G + A
Sbjct: 204 QMRGEIPDEVARLTQMVFFQIALNSFSGGFPPALYNISSLESLSLADNSFSGNLRA 259
Score = 57.0 bits (136), Expect = 7e-09
Identities = 34/101 (33%), Positives = 52/101 (50%), Gaps = 1/101 (0%)
Query: 88 GTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLK 147
G +P G YL L + + ++ G IP ++ L +DL +N L+G P +GKL+
Sbjct: 455 GRIPQSLGRCRYLLDLWM-DTNRLNGTIPQEILQIPSLAYIDLSNNFLTGHFPEEVGKLE 513
Query: 148 RLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
L + S NKLSG +P ++G S+ + N GAIP
Sbjct: 514 LLVGLGASYNKLSGKMPQAIGGCLSMEFLFMQGNSFDGAIP 554
Score = 56.6 bits (135), Expect = 9e-09
Identities = 38/107 (35%), Positives = 55/107 (50%), Gaps = 2/107 (1%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKL-QRLQNLDLGSNSLSGPIPSFLGK 145
SG P N+ L LSLA+ +G + F L L+ L LG+N +G IP L
Sbjct: 230 SGGFPPALYNISSLESLSLADN-SFSGNLRADFGYLLPNLRRLLLGTNQFTGAIPKTLAN 288
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLK 192
+ L+ D+S+N LSG+IP S G L++L + N L +GL+
Sbjct: 289 ISSLERFDISSNYLSGSIPLSFGKLRNLWWLGIRNNSLGNNSSSGLE 335
Score = 52.4 bits (124), Expect = 2e-07
Identities = 34/103 (33%), Positives = 56/103 (54%), Gaps = 4/103 (3%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG +P G + L + + G IP+ S+L L+N+D +N+LSG IP +L
Sbjct: 525 LSGKMPQAIGGCLSMEFLFM-QGNSFDGAIPD-ISRLVSLKNVDFSNNNLSGRIPRYLAS 582
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFN-QLCGAI 187
L L+ ++LS NK G +P + G ++ + +V N +CG +
Sbjct: 583 LPSLRNLNLSMNKFEGRVPTT-GVFRNATAVSVFGNTNICGGV 624
>At1g33670 disease resistance protein, putative
Length = 455
Score = 103 bits (257), Expect = 7e-23
Identities = 61/186 (32%), Positives = 98/186 (51%), Gaps = 7/186 (3%)
Query: 10 LFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFG-GPKGRLDDWDNNTECCDWSFVGCG 68
LFL I+ +++ C+ DDKA LL + P G L W + +CC W + C
Sbjct: 8 LFLFVIFLRCLSSTGAATCHPDDKAGLLAFKSGITQDPSGILSSWQKDIDCCSWYGIFCL 67
Query: 69 RPYPG-RVTVVTISRGWG-----LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKL 122
G RVT++ + LSGT+ L +L+ + L + K+TG P+ KL
Sbjct: 68 PTIHGDRVTMMALDGNTDVGETFLSGTISPLLAKLHHLNEIRLTNLRKITGSFPHFLFKL 127
Query: 123 QRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
+L+ + L +N LSGP+P+ +G L L+ + ++ N+ SG+IP+S+ L SL Q ++ N+
Sbjct: 128 PKLRTVYLENNRLSGPLPANIGALSNLEILSVAGNRFSGSIPSSMSKLTSLLQLKLNGNR 187
Query: 183 LCGAIP 188
L G P
Sbjct: 188 LSGIFP 193
Score = 79.3 bits (194), Expect = 1e-15
Identities = 47/110 (42%), Positives = 66/110 (59%), Gaps = 2/110 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG LPA G L L +LS+A + +G IP+S SKL L L L N LSG P
Sbjct: 140 LSGPLPANIGALSNLEILSVAGN-RFSGSIPSSMSKLTSLLQLKLNGNRLSGIFPDIFKS 198
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNL-QSLSQFNVSFNQLCGAIPAGLKKF 194
+++L+ +DLS+N+ SG +P+S+ +L +LS V N+L G IP L +F
Sbjct: 199 MRQLRFLDLSSNRFSGNLPSSIASLAPTLSTLEVGHNKLSGTIPDYLSRF 248
Score = 65.5 bits (158), Expect = 2e-11
Identities = 37/75 (49%), Positives = 48/75 (63%), Gaps = 8/75 (10%)
Query: 143 LGKLKR---LKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVF 199
LGKLK LK +DLS N + G +P ++ LQ+L N+S N LCG +P+ KF + F
Sbjct: 385 LGKLKFGIFLKTLDLSRNLVFGKVPVTVTRLQTL---NLSQNHLCGKLPS--TKFPASAF 439
Query: 200 EHNKCLCGAPLAPCK 214
NKCLCG PL+PCK
Sbjct: 440 VDNKCLCGFPLSPCK 454
Score = 62.0 bits (149), Expect = 2e-10
Identities = 36/104 (34%), Positives = 57/104 (54%), Gaps = 2/104 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
LSG P F ++ L L L+ + +G +P+S + L L L++G N LSG IP +L
Sbjct: 188 LSGIFPDIFKSMRQLRFLDLSSN-RFSGNLPSSIASLAPTLSTLEVGHNKLSGTIPDYLS 246
Query: 145 KLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
+ + L ++LS N +G +P S NL ++ ++S N L G P
Sbjct: 247 RFELLSALNLSRNGYTGVVPMSFANLTNIIFLDLSHNLLTGPFP 290
>At5g23400 disease resistance protein-like
Length = 589
Score = 103 bits (256), Expect = 9e-23
Identities = 70/199 (35%), Positives = 101/199 (50%), Gaps = 11/199 (5%)
Query: 7 LTLLFLSSIYFS-KSNASDDFFCNADDKAALLKIRDHF-GGPKGRLDDWDNNTECC--DW 62
+ LLF+S++ + ++S C++ D+A LL + G LD W +CC DW
Sbjct: 9 MNLLFVSALVRNFVLSSSQQVICSSQDRATLLGFKSSIIEDTTGVLDSWVGK-DCCNGDW 67
Query: 63 SFVGCGRPYPGRVTVVTISRGWG-----LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPN 117
V C P G+VT + + + GTL GNL L +L + +TG IPN
Sbjct: 68 EGVQCN-PATGKVTGLVLQSAVNEPTLYMKGTLSPSLGNLRSLELLLITGNKFITGSIPN 126
Query: 118 SFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFN 177
SFS L L+ L L NSL G + S LG L L+ + L+ N+ SG +PAS G+L+ L+ N
Sbjct: 127 SFSNLTSLRQLILDDNSLQGNVLSSLGHLPLLEILSLAGNRFSGLVPASFGSLRRLTTMN 186
Query: 178 VSFNQLCGAIPAGLKKFNK 196
++ N G IP K K
Sbjct: 187 LARNSFSGPIPVTFKNLLK 205
Score = 94.7 bits (234), Expect = 3e-20
Identities = 53/131 (40%), Positives = 78/131 (59%), Gaps = 4/131 (3%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
+SG +P +FG L +L++ K++G IP+S S L L LD+ N ++G IP +G+
Sbjct: 457 ISGRIP-DFGESLNLKVLNIGSN-KISGQIPSSISNLVELVRLDISRNHITGGIPQAIGQ 514
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNK 203
L +LK +DLS N L+G IP SL N++++ + N+LCG IP G F + HN
Sbjct: 515 LAQLKWLDLSINALTGRIPDSLLNIKTIKHASFRANRLCGQIPQGRPFNIFPAAAYLHNL 574
Query: 204 CLCGAPLAPCK 214
CLCG PL C+
Sbjct: 575 CLCGKPLPACR 585
Score = 85.5 bits (210), Expect = 2e-17
Identities = 44/101 (43%), Positives = 66/101 (64%), Gaps = 1/101 (0%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
SG +PA FG+L L+ ++LA +GPIP +F L +L+NLDL SN LSGPIP F+G+
Sbjct: 169 SGLVPASFGSLRRLTTMNLARN-SFSGPIPVTFKNLLKLENLDLSSNLLSGPIPDFIGQF 227
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
+ L + LS+N+ SG +P S+ +L+ L ++ N L G +
Sbjct: 228 QNLTNLYLSSNRFSGVLPVSVYSLRKLQTMSLERNGLTGPL 268
Score = 67.8 bits (164), Expect = 4e-12
Identities = 41/118 (34%), Positives = 64/118 (53%), Gaps = 2/118 (1%)
Query: 74 RVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSN 133
R+T + ++R SG +P F NL L L L+ ++GPIP+ + Q L NL L SN
Sbjct: 181 RLTTMNLARN-SFSGPIPVTFKNLLKLENLDLSSN-LLSGPIPDFIGQFQNLTNLYLSSN 238
Query: 134 SLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
SG +P + L++L+ + L N L+G + L+SL+ +S N+ G IPA +
Sbjct: 239 RFSGVLPVSVYSLRKLQTMSLERNGLTGPLSDRFSYLKSLTSLQLSGNKFIGHIPASI 296
Score = 36.2 bits (82), Expect = 0.013
Identities = 28/99 (28%), Positives = 48/99 (48%), Gaps = 28/99 (28%)
Query: 118 SFSKLQR---LQNLDLGSNSLSGPIPSFLGKLKRLKEV---------------------- 152
+F KL R L +LDL N L+G + +FL L +++V
Sbjct: 364 TFPKLTRPTTLTSLDLSDNFLTGDVSAFLTSLTNVQKVKLSKNQLRFDLSKLKLPEGVAS 423
Query: 153 -DLSNNKLSGTIPASLGNLQS--LSQFNVSFNQLCGAIP 188
DLS+N ++G++ + + N S L + +++ NQ+ G IP
Sbjct: 424 IDLSSNLVTGSLSSLINNKTSSFLEEIHLTNNQISGRIP 462
>At1g31420 unknown protein
Length = 592
Score = 102 bits (253), Expect = 2e-22
Identities = 74/202 (36%), Positives = 105/202 (51%), Gaps = 11/202 (5%)
Query: 10 LFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDW-DNNTECCDWSFVGCG 68
L L S+ S SN S + D ALL R+ + W + + C+W+ V C
Sbjct: 14 LLLISLLCSLSNESQAI---SPDGEALLSFRNAVTRSDSFIHQWRPEDPDPCNWNGVTCD 70
Query: 69 RPYPGRVTV-VTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN 127
+T+ +T + + G LP + G L +L +L L + G IP + L+
Sbjct: 71 AKTKRVITLNLTYHK---IMGPLPPDIGKLDHLRLLMLHNNA-LYGAIPTALGNCTALEE 126
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
+ L SN +GPIP+ +G L L+++D+S+N LSG IPASLG L+ LS FNVS N L G I
Sbjct: 127 IHLQSNYFTGPIPAEMGDLPGLQKLDMSSNTLSGPIPASLGQLKKLSNFNVSNNFLVGQI 186
Query: 188 PAG--LKKFNKNVFEHNKCLCG 207
P+ L F+KN F N LCG
Sbjct: 187 PSDGVLSGFSKNSFIGNLNLCG 208
>At1g34210 putative protein
Length = 628
Score = 101 bits (252), Expect = 3e-22
Identities = 74/217 (34%), Positives = 104/217 (47%), Gaps = 19/217 (8%)
Query: 2 FLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNN-TECC 60
F+ L LL +S++ + SN D AL +R + P L WD C
Sbjct: 11 FVCLISLLLLFNSLWLASSNMEGD---------ALHSLRANLVDPNNVLQSWDPTLVNPC 61
Query: 61 DWSFVGCGRPYPGRVTVVTISRGWG-LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSF 119
W V C +V+ + G LSG L + G L L L L +TGP+P+
Sbjct: 62 TWFHVTCNNEN----SVIRVDLGNADLSGQLVPQLGQLKNLQYLELYSN-NITGPVPSDL 116
Query: 120 SKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVS 179
L L +LDL NS +GPIP LGKL +L+ + L+NN L+G IP SL N+ +L ++S
Sbjct: 117 GNLTNLVSLDLYLNSFTGPIPDSLGKLFKLRFLRLNNNSLTGPIPMSLTNIMTLQVLDLS 176
Query: 180 FNQLCGAIP--AGLKKFNKNVFEHNKCLCGAPLA-PC 213
N+L G++P F F +N LCG + PC
Sbjct: 177 NNRLSGSVPDNGSFSLFTPISFANNLDLCGPVTSRPC 213
>At3g51740 unknown protein
Length = 836
Score = 100 bits (250), Expect = 4e-22
Identities = 55/123 (44%), Positives = 76/123 (61%), Gaps = 2/123 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
++GT+P F NL L L+L E + GPIP++ +L L L+L N ++GPIP +G
Sbjct: 299 INGTIPDSFSNLSSLVSLNL-ESNHLKGPIPDAIDRLHNLTELNLKRNKINGPIPETIGN 357
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL-KKFNKNVFEHNKC 204
+ +K++DLS N +G IP SL +L LS FNVS+N L G +P L KKFN + F N
Sbjct: 358 ISGIKKLDLSENNFTGPIPLSLVHLAKLSSFNVSYNTLSGPVPPVLSKKFNSSSFLGNIQ 417
Query: 205 LCG 207
LCG
Sbjct: 418 LCG 420
Score = 84.0 bits (206), Expect = 6e-17
Identities = 55/150 (36%), Positives = 77/150 (50%), Gaps = 10/150 (6%)
Query: 48 GRLDDWDNNTE---CCDWSFVGCGRPYPGRVTVVTISRGW-GLSGTLPAEFGNLPYLSML 103
G L W+N+ C W+ + C R VV I W GL GT+ + G L L L
Sbjct: 69 GVLKSWNNSASSQVCSGWAGIKCLRGQ-----VVAIQLPWKGLGGTISEKIGQLGSLRKL 123
Query: 104 SLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTI 163
SL + G +P S L+ L+ + L +N LSG IP LG L+ +DLS+N+L+G I
Sbjct: 124 SLHNNV-IAGSVPRSLGYLKSLRGVYLFNNRLSGSIPVSLGNCPLLQNLDLSSNQLTGAI 182
Query: 164 PASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
P SL L + N+SFN L G +P + +
Sbjct: 183 PPSLTESTRLYRLNLSFNSLSGPLPVSVAR 212
Score = 67.0 bits (162), Expect = 7e-12
Identities = 44/118 (37%), Positives = 64/118 (53%), Gaps = 3/118 (2%)
Query: 86 LSGTLPAEFGNLPY-LSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLG 144
LSG++P F N + L L+L + + +G +P S K L+ + + N LSG IP G
Sbjct: 226 LSGSIPDFFVNGSHPLKTLNL-DHNRFSGAVPVSLCKHSLLEEVSISHNQLSGSIPRECG 284
Query: 145 KLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFEHN 202
L L+ +D S N ++GTIP S NL SL N+ N L G IP + + + N+ E N
Sbjct: 285 GLPHLQSLDFSYNSINGTIPDSFSNLSSLVSLNLESNHLKGPIPDAIDRLH-NLTELN 341
>At4g13820 disease resistance protein - like
Length = 823
Score = 100 bits (249), Expect = 6e-22
Identities = 64/206 (31%), Positives = 95/206 (46%), Gaps = 36/206 (17%)
Query: 28 CNADDKAALLKIRDHF-------GGPKG--RLDDWDNNTECCDWSFVGCGRPYPGRVTVV 78
C D K ALL+ ++ F G G + + W NNT+CC W + C P G+V +
Sbjct: 29 CRQDQKNALLEFKNEFYVHEFNSNGIVGVKKTEKWRNNTDCCSWDGISCD-PKTGKVVEL 87
Query: 79 TISRGW-------------------------GLSGTLPAEFGNLPYLSMLSLAEMPKVTG 113
+ + SG LP G+L YL +LSL + + G
Sbjct: 88 DLMNSFLNGPLRYDSSLFRLQHLHNLDLGSNNFSGILPDSIGSLKYLRVLSLGDC-NLFG 146
Query: 114 PIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSL 173
IP+S L L NLDL N +G +P +G L +L E+ L + KLSG P+ L NL L
Sbjct: 147 KIPSSLGNLTYLTNLDLSVNDFTGELPDSMGHLNKLTELHLGSAKLSGNFPSMLLNLSEL 206
Query: 174 SQFNVSFNQLCGAIPAGLKKFNKNVF 199
+ ++ NQ G +P+ + +K V+
Sbjct: 207 TLIDLGSNQFGGMLPSNMSSLSKLVY 232
Score = 48.9 bits (115), Expect = 2e-06
Identities = 30/88 (34%), Positives = 46/88 (52%), Gaps = 11/88 (12%)
Query: 134 SLSGPIPSFLGKLKRL-KEVDLSNNKLSGTIPASLG--------NLQSLSQFNVSFNQLC 184
++ G I +G + + K +D+S N+ G IP S+G N+ + +Q N S+N L
Sbjct: 670 TVKGSIIELVGSVFTIYKTIDVSGNRFEGRIPESIGLLKELIVLNMSNNAQMNFSYNMLE 729
Query: 185 GAIPAG--LKKFNKNVFEHNKCLCGAPL 210
G IP G ++ N + F N LCG PL
Sbjct: 730 GPIPQGTQIQSQNSSSFAENLGLCGVPL 757
Score = 41.2 bits (95), Expect = 4e-04
Identities = 34/119 (28%), Positives = 53/119 (43%), Gaps = 20/119 (16%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAE--MPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPS-- 141
+ G +P +LP L +++++ GP + + L LD+ SN+ P P
Sbjct: 409 IGGQVPQWLWSLPELQYVNISQNSFSGFEGPA-DVIQRCGELLMLDISSNTFQDPFPLLP 467
Query: 142 -----FLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFN 195
FLG S+N+ SG IP ++ L SL +S N G+IP +KFN
Sbjct: 468 NSTTIFLG----------SDNRFSGEIPKTICKLVSLDTLVLSNNNFNGSIPRCFEKFN 516
Score = 38.5 bits (88), Expect = 0.003
Identities = 23/73 (31%), Positives = 36/73 (48%), Gaps = 8/73 (10%)
Query: 89 TLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKR 148
+LP+ G L +LS +P+ PN L LD+ +N + G +P +L L
Sbjct: 371 SLPSPMGTL----ILSSCNIPE----FPNFLENQTTLYYLDISANKIGGQVPQWLWSLPE 422
Query: 149 LKEVDLSNNKLSG 161
L+ V++S N SG
Sbjct: 423 LQYVNISQNSFSG 435
Score = 37.0 bits (84), Expect = 0.008
Identities = 31/102 (30%), Positives = 50/102 (48%), Gaps = 3/102 (2%)
Query: 87 SGTLPAEFGNL-PYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
+G++P F LS+L L ++G P S L++LD+G N LSG +P L
Sbjct: 505 NGSIPRCFEKFNTTLSVLHLRNN-NLSGEFPEE-SISDHLRSLDVGRNRLSGELPKSLIN 562
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
RL+ +++ +N ++ P L L L F + N+ G I
Sbjct: 563 CTRLEFLNVEDNIINDKFPFWLRMLPKLQIFVLRSNEFHGPI 604
Score = 35.0 bits (79), Expect = 0.030
Identities = 31/97 (31%), Positives = 48/97 (48%), Gaps = 4/97 (4%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIP-NSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
SG++P+ LP L+ L L GP+ + S L L L N+ +GPIP + K
Sbjct: 241 SGSIPSSLFMLPSLTSLVLGRND-FNGPLDFGNISSPSNLGVLSLLENNFNGPIPESISK 299
Query: 146 LKRLKEVDLS--NNKLSGTIPASLGNLQSLSQFNVSF 180
L L +DLS N K + +L+SL+ ++S+
Sbjct: 300 LVGLFYLDLSLWNTKRGMVDFNTFLHLKSLTFLDLSY 336
Score = 33.9 bits (76), Expect = 0.066
Identities = 17/46 (36%), Positives = 24/46 (51%)
Query: 140 PSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCG 185
P+FL L +D+S NK+ G +P L +L L N+S N G
Sbjct: 390 PNFLENQTTLYYLDISANKIGGQVPQWLWSLPELQYVNISQNSFSG 435
>At3g25560 unknown protein
Length = 635
Score = 100 bits (248), Expect = 7e-22
Identities = 63/193 (32%), Positives = 100/193 (51%), Gaps = 6/193 (3%)
Query: 2 FLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNT-ECC 60
+ L T F + S S+A + AL+ I+ P G L +WD+ + C
Sbjct: 12 YALFSSTFFFFFICFLSSSSAELTDKGVNFEVVALIGIKSSLTDPHGVLMNWDDTAVDPC 71
Query: 61 DWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFS 120
W+ + C + R+ + + LSGTL + GNL L + L + +TG IP+
Sbjct: 72 SWNMITCSDGFVIRLEAPSQN----LSGTLSSSIGNLTNLQTV-LLQNNYITGNIPHEIG 126
Query: 121 KLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSF 180
KL +L+ LDL +N+ +G IP L K L+ + ++NN L+GTIP+SL N+ L+ ++S+
Sbjct: 127 KLMKLKTLDLSTNNFTGQIPFTLSYSKNLQYLRVNNNSLTGTIPSSLANMTQLTFLDLSY 186
Query: 181 NQLCGAIPAGLKK 193
N L G +P L K
Sbjct: 187 NNLSGPVPRSLAK 199
>At1g68400 receptor kinase like protein
Length = 670
Score = 100 bits (248), Expect = 7e-22
Identities = 76/230 (33%), Positives = 101/230 (43%), Gaps = 33/230 (14%)
Query: 3 LLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDW 62
LLL L +L S + S S+ + N A G+L+ W+ T C W
Sbjct: 11 LLLSLLILLQSCLLSSSSSTDSETLLNFKLTA----------DSTGKLNSWNTTTNPCQW 60
Query: 63 SFVGCGRPYPGRVTVVTIS-------------------RGWGLSGTLPAEFGNLPYLSML 103
+ V C R R+ + I+ + LSG +P NL L +L
Sbjct: 61 TGVSCNRNRVTRLVLEDINLTGSISSLTSLTSLRVLSLKHNNLSGPIP-NLSNLTALKLL 119
Query: 104 SLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTI 163
L+ + +G P S + L RL LDL N+ SG IP L L L + L +N+ SG I
Sbjct: 120 FLSNN-QFSGNFPTSITSLTRLYRLDLSFNNFSGQIPPDLTDLTHLLTLRLESNRFSGQI 178
Query: 164 PASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFEHNKCLCGAPLAPC 213
P NL L FNVS N G IP L +F ++VF N LCGAPL C
Sbjct: 179 PNI--NLSDLQDFNVSGNNFNGQIPNSLSQFPESVFTQNPSLCGAPLLKC 226
>At1g33600 unknown protein
Length = 478
Score = 100 bits (248), Expect = 7e-22
Identities = 59/170 (34%), Positives = 84/170 (48%), Gaps = 9/170 (5%)
Query: 28 CNADDKAALLKIRDHFG-GPKGRLDDWDNNTECCDWSFVGCGRPYPGRVTVVTIS----- 81
C+ DD+A LL + P G L W T+CC W VGC RVT +TI+
Sbjct: 28 CHPDDEAGLLAFKSGITQDPTGILSSWKKGTDCCSWKGVGC---LTNRVTGLTINGQSDV 84
Query: 82 RGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPS 141
G LSGT+ L +L + + +TG P +L ++ + ++ LSGP+P+
Sbjct: 85 TGSFLSGTISPSLAKLQHLVGIYFTNLRNITGSFPQFLFQLPNVKQVYFTNSRLSGPLPA 144
Query: 142 FLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
+G L L E+ L N +G IP+S+ NL L N+ N L G IP GL
Sbjct: 145 NIGALSELGELSLDGNLFTGPIPSSISNLTRLYLLNLGDNLLTGTIPLGL 194
Score = 77.8 bits (190), Expect = 4e-15
Identities = 45/105 (42%), Positives = 63/105 (59%), Gaps = 2/105 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN-LDLGSNSLSGPIPSFLG 144
LS T+P F ++ L L+L+ K +G +P S + L+ + N LDL N+LSG IP+FL
Sbjct: 210 LSETIPDIFKSMQKLQSLTLSRN-KFSGNLPPSIASLKPILNYLDLSQNNLSGTIPTFLS 268
Query: 145 KLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPA 189
K L +DLS N+ SG +P SL N+ L N+S N L G +PA
Sbjct: 269 NFKVLDSLDLSRNRFSGVVPKSLANMPKLFHLNLSHNFLTGPLPA 313
Score = 77.0 bits (188), Expect = 7e-15
Identities = 47/106 (44%), Positives = 60/106 (56%), Gaps = 1/106 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG LPA G L L LSL + TGPIP+S S L RL L+LG N L+G IP L
Sbjct: 138 LSGPLPANIGALSELGELSL-DGNLFTGPIPSSISNLTRLYLLNLGDNLLTGTIPLGLAN 196
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
LK L ++ NN+LS TIP ++Q L +S N+ G +P +
Sbjct: 197 LKILLSLNFGNNRLSETIPDIFKSMQKLQSLTLSRNKFSGNLPPSI 242
Score = 72.0 bits (175), Expect = 2e-13
Identities = 42/112 (37%), Positives = 60/112 (53%), Gaps = 2/112 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+GT+P NL L L+ +++ IP+ F +Q+LQ+L L N SG +P +
Sbjct: 186 LTGTIPLGLANLKILLSLNFGNN-RLSETIPDIFKSMQKLQSLTLSRNKFSGNLPPSIAS 244
Query: 146 LKR-LKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
LK L +DLS N LSGTIP L N + L ++S N+ G +P L K
Sbjct: 245 LKPILNYLDLSQNNLSGTIPTFLSNFKVLDSLDLSRNRFSGVVPKSLANMPK 296
Score = 65.5 bits (158), Expect = 2e-11
Identities = 40/110 (36%), Positives = 57/110 (51%), Gaps = 28/110 (25%)
Query: 128 LDLGSNSLSGPIPSF--------------------LGKL---KRLKEVDLSNNKLSGTIP 164
+DL N +SG + F +GKL +RL+ +DLS N + G +P
Sbjct: 373 IDLSENEISGSLTWFFNLAHNLYEFQASGNKLRFDMGKLNLSERLESLDLSRNLIFGKVP 432
Query: 165 ASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFEHNKCLCGAPLAPCK 214
++ LQ L N+S N LCG +P + KF + F N CLCG+PL+PCK
Sbjct: 433 MTVAKLQKL---NLSHNHLCGKLP--VTKFPASAFVGNDCLCGSPLSPCK 477
>At3g02880 putative protein kinase
Length = 627
Score = 99.8 bits (247), Expect = 1e-21
Identities = 81/233 (34%), Positives = 113/233 (47%), Gaps = 37/233 (15%)
Query: 5 LHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTEC-CDWS 63
L L+++FL Y + + + D+ ALL +R+ +GR W+ + C+W
Sbjct: 7 LSLSVVFLFVFYLAAVTSDLE-----SDRRALLAVRNSV---RGRPLLWNMSASSPCNWH 58
Query: 64 FVGCGRPYPGRVTVVTISRGWGLSGTLP-AEFGNLPYLSMLSLAEMPKVTGPIPNSFSKL 122
V C GRVT + + G GL G+LP GNL L LSL ++GPIP+ FS L
Sbjct: 59 GVHCDA---GRVTALRLP-GSGLFGSLPIGGIGNLTQLKTLSL-RFNSLSGPIPSDFSNL 113
Query: 123 QRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQ----------- 171
L+ L L N+ SG IPS L L + ++L NK SG IP ++ +
Sbjct: 114 VLLRYLYLQGNAFSGEIPSLLFTLPSIIRINLGENKFSGRIPDNVNSATRLVTLYLERNQ 173
Query: 172 ----------SLSQFNVSFNQLCGAIPAGLKKFNKNVFEHNKCLCGAPLAPCK 214
L QFNVS NQL G+IP+ L + + FE N LCG PL C+
Sbjct: 174 LSGPIPEITLPLQQFNVSSNQLNGSIPSSLSSWPRTAFEGN-TLCGKPLDTCE 225
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.139 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,146,533
Number of Sequences: 26719
Number of extensions: 233710
Number of successful extensions: 5933
Number of sequences better than 10.0: 439
Number of HSP's better than 10.0 without gapping: 393
Number of HSP's successfully gapped in prelim test: 48
Number of HSP's that attempted gapping in prelim test: 541
Number of HSP's gapped (non-prelim): 2649
length of query: 214
length of database: 11,318,596
effective HSP length: 95
effective length of query: 119
effective length of database: 8,780,291
effective search space: 1044854629
effective search space used: 1044854629
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146750.9


