

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146750.10 + phase: 0
(221 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
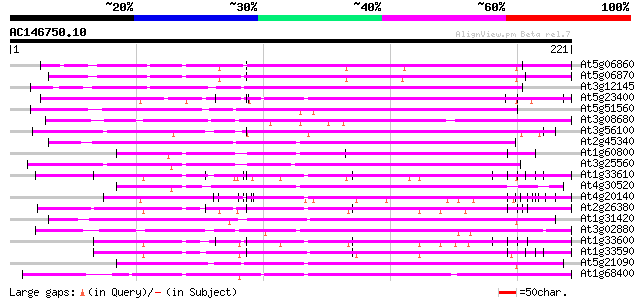
Sequences producing significant alignments: (bits) Value
At5g06860 polygalacturonase inhibiting protein 1; PGIP1 (gb|AAF6... 130 4e-31
At5g06870 polygalacturonase inhibiting protein 126 1e-29
At3g12145 leucine-rich repeat protein FLR1 124 5e-29
At5g23400 disease resistance protein-like 107 6e-24
At5g51560 receptor-like protein kinase 104 4e-23
At3g08680 putative protein kinase 102 2e-22
At3g56100 unknown protein 101 4e-22
At2g45340 putative receptor-like protein kinase 99 1e-21
At1g60800 Putative Serine/Threonine protein kinase 99 1e-21
At3g25560 unknown protein 99 2e-21
At1g33610 hypothetical protein 97 7e-21
At4g30520 receptor-like kinase homolog 97 9e-21
At4g20140 leucine rich repeat-like protein 96 1e-20
At2g26380 putative disease resistance protein 96 1e-20
At1g31420 unknown protein 96 1e-20
At3g02880 putative protein kinase 96 1e-20
At1g33600 unknown protein 96 1e-20
At1g33590 unknown protein 96 2e-20
At5g21090 leucine-rich repeat protein 95 3e-20
At1g68400 receptor kinase like protein 95 3e-20
>At5g06860 polygalacturonase inhibiting protein 1; PGIP1
(gb|AAF69827.1)
Length = 330
Score = 130 bits (328), Expect = 4e-31
Identities = 80/191 (41%), Positives = 111/191 (57%), Gaps = 8/191 (4%)
Query: 13 LTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSF 72
L LLFL + + +T + D+ CN NDK TLLKI+ P L+ WD TDCCS W
Sbjct: 7 LCLLFLFT-FLTTCLSKDL---CNQNDKNTLLKIKKSLNNPY-HLASWDPQTDCCS-WYC 60
Query: 73 VGCGKPLPG-RITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ 131
+ CG R+T +TI G +SG +PAE GDLPYL L ++ +TG I + +KL+
Sbjct: 61 LECGDATVNHRVTALTIFSGQ-ISGQIPAEVGDLPYLETLVFRKLSNLTGTIQPTIAKLK 119
Query: 132 RLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQL 191
L+ L L +L+G IP F+ QLK L+ +LS N L+G IP+S +LP + +S N+L
Sbjct: 120 NLRMLRLSWTNLTGPIPDFISQLKNLEFLELSFNDLSGSIPSSLSTLPKILALELSRNKL 179
Query: 192 CGAIPTGLSKF 202
G+IP F
Sbjct: 180 TGSIPESFGSF 190
Score = 82.0 bits (201), Expect = 2e-16
Identities = 62/177 (35%), Positives = 77/177 (43%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ-RLQKLDLGSNSLSGSIPTFLG 152
LSG++P+ LP + L L+ K+TG IP SF + L L N LSG IP LG
Sbjct: 155 LSGSIPSSLSTLPKILALELSRN-KLTGSIPESFGSFPGTVPDLRLSHNQLSGPIPKSLG 213
Query: 153 QL----------------------------------------------KGLQEFDLSNNK 166
+ K L DL++N
Sbjct: 214 NIDFNRIDLSRNKLQGDASMLFGSNKTTWSIDLSRNMFQFDISKVDIPKTLGILDLNHNG 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+TG IP + P L FNVS+N+LCG IPTG L F S+ HNKCLCGAPL C
Sbjct: 274 ITGNIPVQWTEAP-LQFFNVSYNKLCGHIPTGGKLQTFDSYSYFHNKCLCGAPLEIC 329
>At5g06870 polygalacturonase inhibiting protein
Length = 330
Score = 126 bits (316), Expect = 1e-29
Identities = 78/189 (41%), Positives = 106/189 (55%), Gaps = 7/189 (3%)
Query: 16 LFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVGC 75
L LS++ +TS A D+ C+ +DK TLLKI+ P L+ WD TDCCS W + C
Sbjct: 9 LLLSTLLLTTSLAKDL---CHKDDKTTLLKIKKSLNNPY-HLASWDPKTDCCS-WYCLEC 63
Query: 76 GKPLPG-RITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQ 134
G R+T + I G +SG +P E GDLPYL L ++ +TG I + +KL+ L
Sbjct: 64 GDATVNHRVTSLIIQDG-EISGQIPPEVGDLPYLTSLIFRKLTNLTGHIQPTIAKLKNLT 122
Query: 135 KLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGA 194
L L +L+G +P FL QLK L+ DLS N L+G IP+S SL L +S N+L G
Sbjct: 123 FLRLSWTNLTGPVPEFLSQLKNLEYIDLSFNDLSGSIPSSLSSLRKLEYLELSRNKLTGP 182
Query: 195 IPTGLSKFA 203
IP F+
Sbjct: 183 IPESFGTFS 191
Score = 84.7 bits (208), Expect = 3e-17
Identities = 60/177 (33%), Positives = 79/177 (43%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ-RLQKLDLGSNSLSGSIPTFLG 152
LSG++P+ L L +L L+ K+TGPIP SF ++ L L N LSG+IP LG
Sbjct: 155 LSGSIPSSLSSLRKLEYLELSRN-KLTGPIPESFGTFSGKVPSLFLSHNQLSGTIPKSLG 213
Query: 153 Q----------------------------------------------LKGLQEFDLSNNK 166
K L D+++N
Sbjct: 214 NPDFYRIDLSRNKLQGDASILFGAKKTTWIVDISRNMFQFDLSKVKLAKTLNNLDMNHNG 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+TG IPA + S NVS+N+LCG IP G + +F SF HNKCLCGAPL +C
Sbjct: 274 ITGSIPAEW-SKAYFQLLNVSYNRLCGRIPKGEYIQRFDSYSFFHNKCLCGAPLPSC 329
>At3g12145 leucine-rich repeat protein FLR1
Length = 203
Score = 124 bits (310), Expect = 5e-29
Identities = 74/194 (38%), Positives = 107/194 (55%), Gaps = 10/194 (5%)
Query: 9 LLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCS 68
L +HL++ F SI F T + +SC NDK LL+I+ G P LS W+ TDCC+
Sbjct: 3 LFVHLSIFF--SILFITLPSS---YSCTENDKNALLQIKKALGNPP-LLSSWNPRTDCCT 56
Query: 69 DWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFS 128
W+ V C R+T ++++ G +SG + + GDL L L + +P +TG IP + +
Sbjct: 57 GWTGVECTNR---RVTGLSVTSG-EVSGQISYQIGDLVDLRTLDFSYLPHLTGNIPRTIT 112
Query: 129 KLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSF 188
KL+ L L L SLSG IP ++ +LK L DLS N+ TG IP S +P L ++
Sbjct: 113 KLKNLNTLYLKHTSLSGPIPDYISELKSLTFLDLSFNQFTGPIPGSLSQMPKLEAIQIND 172
Query: 189 NQLCGAIPTGLSKF 202
N+L G+IP F
Sbjct: 173 NKLTGSIPNSFGSF 186
>At5g23400 disease resistance protein-like
Length = 589
Score = 107 bits (266), Expect = 6e-24
Identities = 71/214 (33%), Positives = 108/214 (50%), Gaps = 10/214 (4%)
Query: 13 LTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHF-GGPNGRLSDWDNGTDCCS-DW 70
+ LLF+S++ + + + C++ D+ATLL + G L W G DCC+ DW
Sbjct: 9 MNLLFVSALVRNFVLSSSQQVICSSQDRATLLGFKSSIIEDTTGVLDSWV-GKDCCNGDW 67
Query: 71 SFVGCGKPLPGRITVVTISRGWG-----LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPN 125
V C P G++T + + + GTL G+L L L + +TG IPN
Sbjct: 68 EGVQCN-PATGKVTGLVLQSAVNEPTLYMKGTLSPSLGNLRSLELLLITGNKFITGSIPN 126
Query: 126 SFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFN 185
SFS L L++L L NSL G++ + LG L L+ L+ N+ +G++PASFGSL L+ N
Sbjct: 127 SFSNLTSLRQLILDDNSLQGNVLSSLGHLPLLEILSLAGNRFSGLVPASFGSLRRLTTMN 186
Query: 186 VSFNQLCGAIPTGLSKFAK-SSFDHNKCLCGAPL 218
++ N G IP K + D + L P+
Sbjct: 187 LARNSFSGPIPVTFKNLLKLENLDLSSNLLSGPI 220
Score = 94.0 bits (232), Expect = 6e-20
Identities = 53/130 (40%), Positives = 78/130 (59%), Gaps = 4/130 (3%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
+SG +P +FG+ L L++ K++G IP+S S L L +LD+ N ++G IP +GQ
Sbjct: 457 ISGRIP-DFGESLNLKVLNIGSN-KISGQIPSSISNLVELVRLDISRNHITGGIPQAIGQ 514
Query: 154 LKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNK 211
L L+ DLS N LTG IP S ++ ++ + N+LCG IP G + F +++ HN
Sbjct: 515 LAQLKWLDLSINALTGRIPDSLLNIKTIKHASFRANRLCGQIPQGRPFNIFPAAAYLHNL 574
Query: 212 CLCGAPLAAC 221
CLCG PL AC
Sbjct: 575 CLCGKPLPAC 584
Score = 79.3 bits (194), Expect = 1e-15
Identities = 44/101 (43%), Positives = 60/101 (58%), Gaps = 1/101 (0%)
Query: 95 SGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQL 154
SG +PA FG L L ++LA +GPIP +F L +L+ LDL SN LSG IP F+GQ
Sbjct: 169 SGLVPASFGSLRRLTTMNLARN-SFSGPIPVTFKNLLKLENLDLSSNLLSGPIPDFIGQF 227
Query: 155 KGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAI 195
+ L LS+N+ +GV+P S SL L ++ N L G +
Sbjct: 228 QNLTNLYLSSNRFSGVLPVSVYSLRKLQTMSLERNGLTGPL 268
Score = 65.5 bits (158), Expect = 2e-11
Identities = 41/119 (34%), Positives = 61/119 (50%), Gaps = 2/119 (1%)
Query: 82 RITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSN 141
R+T + ++R SG +P F +L L L L+ ++GPIP+ + Q L L L SN
Sbjct: 181 RLTTMNLARN-SFSGPIPVTFKNLLKLENLDLSSN-LLSGPIPDFIGQFQNLTNLYLSSN 238
Query: 142 SLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLS 200
SG +P + L+ LQ L N LTG + F L SL+ +S N+ G IP ++
Sbjct: 239 RFSGVLPVSVYSLRKLQTMSLERNGLTGPLSDRFSYLKSLTSLQLSGNKFIGHIPASIT 297
Score = 33.9 bits (76), Expect = 0.070
Identities = 28/99 (28%), Positives = 43/99 (43%), Gaps = 28/99 (28%)
Query: 126 SFSKLQR---LQKLDLGSNSLSGSIPTFLGQL-----------------------KGLQE 159
+F KL R L LDL N L+G + FL L +G+
Sbjct: 364 TFPKLTRPTTLTSLDLSDNFLTGDVSAFLTSLTNVQKVKLSKNQLRFDLSKLKLPEGVAS 423
Query: 160 FDLSNNKLTGVIPASFGSLPS--LSQFNVSFNQLCGAIP 196
DLS+N +TG + + + S L + +++ NQ+ G IP
Sbjct: 424 IDLSSNLVTGSLSSLINNKTSSFLEEIHLTNNQISGRIP 462
>At5g51560 receptor-like protein kinase
Length = 680
Score = 104 bits (259), Expect = 4e-23
Identities = 70/215 (32%), Positives = 112/215 (51%), Gaps = 31/215 (14%)
Query: 9 LLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCS 68
+LLHL LL L ++FS ++ ++ ATL++++ + L+ W D C
Sbjct: 6 MLLHLLLLPLLFLHFSNQVMAEI-----TDELATLMEVKTELDPEDKHLASWSVNGDLCK 60
Query: 69 DWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSL------AEMPK---- 118
D+ VGC GR++ +++ +G GLSG + G L +L L L ++P+
Sbjct: 61 DFEGVGCD--WKGRVSNISL-QGKGLSGKISPNIGKLKHLTGLFLHYNALVGDIPRELGN 117
Query: 119 -------------VTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNN 165
++G IP++ K+Q LQ L L N+L+GSIP L L+ L L +N
Sbjct: 118 LSELTDLYLNVNNLSGEIPSNIGKMQGLQVLQLCYNNLTGSIPRELSSLRKLSVLALQSN 177
Query: 166 KLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLS 200
KLTG IPAS G L +L + ++S+N L G++P L+
Sbjct: 178 KLTGAIPASLGDLSALERLDLSYNHLFGSVPGKLA 212
>At3g08680 putative protein kinase
Length = 640
Score = 102 bits (253), Expect = 2e-22
Identities = 79/229 (34%), Positives = 109/229 (47%), Gaps = 34/229 (14%)
Query: 15 LLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVG 74
L L + + S + D+E +DK LL+ P+ R +W++ C+ W+ +
Sbjct: 9 LFLLVTTFVSRCLSADIE-----SDKQALLEFASLV--PHSRKLNWNSTIPICASWTGIT 61
Query: 75 CGKPLPGRITVVTISRGWGLSGTLPAE-FGDLPYLNFLSL------AEMPKVTGPIP--- 124
C K R+T + + G GL G LP + F L L +SL +P V +P
Sbjct: 62 CSKN-NARVTALRLP-GSGLYGPLPEKTFEKLDALRIISLRSNHLQGNIPSVILSLPFIR 119
Query: 125 ------NSFSKL------QRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIP 172
N+FS RL LDL +NSLSG+IPT L L L + L NN L+G IP
Sbjct: 120 SLYFHENNFSGTIPPVLSHRLVNLDLSANSLSGNIPTSLQNLTQLTDLSLQNNSLSGPIP 179
Query: 173 ASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 221
P L N+SFN L G++P+ + F SSF N LCGAPL C
Sbjct: 180 ---NLPPRLKYLNLSFNNLNGSVPSSVKSFPASSFQGNSLLCGAPLTPC 225
>At3g56100 unknown protein
Length = 719
Score = 101 bits (251), Expect = 4e-22
Identities = 60/134 (44%), Positives = 74/134 (54%), Gaps = 12/134 (8%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEM-----------PKVTGPIPNSFSKLQRLQKLDLGSNS 142
LSG +P L FL+L K+ G +P+ SKL +L+K+D+ NS
Sbjct: 209 LSGQIPVSLSRSSSLQFLALDHNNLSGPILDTWGSKIRGTLPSELSKLTKLRKMDISGNS 268
Query: 143 LSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLS-K 201
+SG IP LG + L DLS NKLTG IP S L SL+ FNVS+N L G +PT LS K
Sbjct: 269 VSGHIPETLGNISSLIHLDLSQNKLTGEIPISISDLESLNFFNVSYNNLSGPVPTLLSQK 328
Query: 202 FAKSSFDHNKCLCG 215
F SSF N LCG
Sbjct: 329 FNSSSFVGNSLLCG 342
Score = 87.8 bits (216), Expect = 4e-18
Identities = 68/207 (32%), Positives = 98/207 (46%), Gaps = 13/207 (6%)
Query: 10 LLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNG--TDCC 67
LLHL + L + +S A D A D L ++ P G L W+ + C
Sbjct: 32 LLHLIICLLFFVPPCSSQAWDGVVITQA-DYQGLQAVKQELIDPRGFLRSWNGSGFSACS 90
Query: 68 SDWSFVGCGKPLPGRITVVTISRGW-GLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNS 126
W+ + C + G++ V+ + W L G + + G L L LSL + + G IP S
Sbjct: 91 GGWAGIKCAQ---GQVIVIQLP--WKSLGGRISEKIGQLQALRKLSLHDN-NLGGSIPMS 144
Query: 127 FSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNV 186
+ L+ + L +N L+GSIP LG LQ DLSNN L+ +IP + L + N+
Sbjct: 145 LGLIPNLRGVQLFNNRLTGSIPASLGVSHFLQTLDLSNNLLSEIIPPNLADSSKLLRLNL 204
Query: 187 SFNQLCGAIPTGLSKFAKSSF---DHN 210
SFN L G IP LS+ + F DHN
Sbjct: 205 SFNSLSGQIPVSLSRSSSLQFLALDHN 231
>At2g45340 putative receptor-like protein kinase
Length = 691
Score = 99.4 bits (246), Expect = 1e-21
Identities = 68/184 (36%), Positives = 91/184 (48%), Gaps = 6/184 (3%)
Query: 16 LFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVGC 75
LFL +F S+A S ++++ LL I+ L+ W D CS SF G
Sbjct: 7 LFLILFFFFLSSA----LSSSSSELDILLDIKSSLDPEKRFLTSWTPDADPCSSGSFDGV 62
Query: 76 GKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQK 135
R+ +++ +G GL+GT+P G L L L L +TG IP S L L
Sbjct: 63 ACDGNRRVANISL-QGMGLTGTIPPSIGLLTSLTGLYL-HFNSLTGHIPKDISNLPLLTD 120
Query: 136 LDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAI 195
L L N+LSG IP +G L LQ L NKL+G IP FGSL ++ + +NQL GAI
Sbjct: 121 LYLNVNNLSGEIPPLIGNLDNLQVIQLCYNKLSGSIPTQFGSLKKITVLALQYNQLSGAI 180
Query: 196 PTGL 199
P L
Sbjct: 181 PASL 184
>At1g60800 Putative Serine/Threonine protein kinase
Length = 632
Score = 99.4 bits (246), Expect = 1e-21
Identities = 57/155 (36%), Positives = 84/155 (53%), Gaps = 7/155 (4%)
Query: 43 LLKIRDHFGGPNGRLSDWD-NGTDCCSDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAE 101
L+ +++ P L +WD N D CS W V C + + + S LSGTL
Sbjct: 39 LVAVKNELNDPYKVLENWDVNSVDPCS-WRMVSCTDGYVSSLDLPSQS----LSGTLSPR 93
Query: 102 FGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFD 161
G+L YL + L + +TGPIP + +L++LQ LDL +NS +G IP LG+LK L
Sbjct: 94 IGNLTYLQSVVL-QNNAITGPIPETIGRLEKLQSLDLSNNSFTGEIPASLGELKNLNYLR 152
Query: 162 LSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIP 196
L+NN L G P S + L+ ++S+N L G++P
Sbjct: 153 LNNNSLIGTCPESLSKIEGLTLVDISYNNLSGSLP 187
Score = 52.8 bits (125), Expect = 1e-07
Identities = 28/75 (37%), Positives = 39/75 (51%)
Query: 133 LQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLC 192
+ LDL S SLSG++ +G L LQ L NN +TG IP + G L L ++S N
Sbjct: 76 VSSLDLPSQSLSGTLSPRIGNLTYLQSVVLQNNAITGPIPETIGRLEKLQSLDLSNNSFT 135
Query: 193 GAIPTGLSKFAKSSF 207
G IP L + ++
Sbjct: 136 GEIPASLGELKNLNY 150
>At3g25560 unknown protein
Length = 635
Score = 98.6 bits (244), Expect = 2e-21
Identities = 63/195 (32%), Positives = 102/195 (52%), Gaps = 8/195 (4%)
Query: 8 FLLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDN-GTDC 66
+ L T F + S+S+A+ + N + L+ I+ P+G L +WD+ D
Sbjct: 12 YALFSSTFFFFFICFLSSSSAELTDKGVNF-EVVALIGIKSSLTDPHGVLMNWDDTAVDP 70
Query: 67 CSDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNS 126
CS W+ + C R+ + + LSGTL + G+L L + L + +TG IP+
Sbjct: 71 CS-WNMITCSDGFVIRLEAPSQN----LSGTLSSSIGNLTNLQTV-LLQNNYITGNIPHE 124
Query: 127 FSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNV 186
KL +L+ LDL +N+ +G IP L K LQ ++NN LTG IP+S ++ L+ ++
Sbjct: 125 IGKLMKLKTLDLSTNNFTGQIPFTLSYSKNLQYLRVNNNSLTGTIPSSLANMTQLTFLDL 184
Query: 187 SFNQLCGAIPTGLSK 201
S+N L G +P L+K
Sbjct: 185 SYNNLSGPVPRSLAK 199
>At1g33610 hypothetical protein
Length = 907
Score = 97.1 bits (240), Expect = 7e-21
Identities = 66/196 (33%), Positives = 94/196 (47%), Gaps = 9/196 (4%)
Query: 11 LHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFG-GPNGRLSDWDNGTDCCSD 69
L TL S I F + +C+ +D+A LL + P+G LS W GT CCS
Sbjct: 4 LSFTLFIFSVITFLQCLSSTGAATCHPDDEAGLLAFKSGITQDPSGMLSSWKKGTSCCS- 62
Query: 70 WSFVGCGKPLPGRITVVTI-----SRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIP 124
W + C R+T++ + LSGTL L +L+ +SL +TG P
Sbjct: 63 WKGIICFNS--DRVTMLELVGFPKKPERSLSGTLSPSLAKLQHLSVISLGGHVNITGSFP 120
Query: 125 NSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQF 184
+L +L+ +D+ +N LSG +P +G L L+E L NK TG IP S +L LS
Sbjct: 121 KFLLQLPKLRYVDIQNNRLSGPLPANIGVLSLLEEIFLQGNKFTGPIPNSISNLTRLSYL 180
Query: 185 NVSFNQLCGAIPTGLS 200
N L G IP G++
Sbjct: 181 IFGGNLLTGTIPLGIA 196
Score = 90.9 bits (224), Expect = 5e-19
Identities = 56/169 (33%), Positives = 87/169 (51%), Gaps = 9/169 (5%)
Query: 34 SCNANDKATLLKIRDHFG-GPNGRLSDWDNGTDCCSDWSFVGCGKPLPGRITVVTISRGW 92
+C+ +D+A LL + P+G LS W GTDCC WS V C R+T +++ +
Sbjct: 479 TCDPDDEAGLLGFKSGITKDPSGILSSWKKGTDCCF-WSGVFCVNN--DRVTQLSVDGDF 535
Query: 93 GL-----SGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSI 147
L SGT+ L +L + L + K+TGP P +L +L +++ LSG +
Sbjct: 536 SLDGNSPSGTISPMLAKLQHLERILLTSLRKITGPFPQFIFRLPKLNYINIQGCLLSGPL 595
Query: 148 PTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIP 196
P +G+L L+ + N TG IP+S +L L+ N+ N+L G IP
Sbjct: 596 PANIGELSQLKTLVIDGNMFTGHIPSSIANLTRLTWLNLGNNRLSGTIP 644
Score = 78.6 bits (192), Expect = 2e-15
Identities = 46/110 (41%), Positives = 62/110 (55%), Gaps = 1/110 (0%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
LSG LPA G L L + L + K TGPIPNS S L RL L G N L+G+IP +
Sbjct: 139 LSGPLPANIGVLSLLEEIFL-QGNKFTGPIPNSISNLTRLSYLIFGGNLLTGTIPLGIAN 197
Query: 154 LKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFA 203
LK +Q L +N+L+G IP F S+ L ++S N+ G +P ++ A
Sbjct: 198 LKLMQNLQLGDNRLSGTIPDIFESMKLLKFLDLSSNEFYGKLPLSIATLA 247
Score = 67.4 bits (163), Expect = 6e-12
Identities = 41/104 (39%), Positives = 59/104 (56%), Gaps = 2/104 (1%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ-RLQKLDLGSNSLSGSIPTFLG 152
LSGT+P F + LN L L+ G +P S + L L LDL N+LSG+IP +L
Sbjct: 639 LSGTIPNIFKSMKELNSLDLSRNG-FFGRLPPSIASLAPTLYYLDLSQNNLSGTIPNYLS 697
Query: 153 QLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIP 196
+ + L LS NK +GV+P SF +L +++ ++S N L G P
Sbjct: 698 RFEALSTLVLSKNKYSGVVPMSFTNLINITNLDLSHNLLTGPFP 741
Score = 67.0 bits (162), Expect = 7e-12
Identities = 37/104 (35%), Positives = 60/104 (57%), Gaps = 2/104 (1%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ-RLQKLDLGSNSLSGSIPTFLG 152
LSGT+P F + L FL L+ + G +P S + L L L + N+LSG+IP ++
Sbjct: 211 LSGTIPDIFESMKLLKFLDLSSN-EFYGKLPLSIATLAPTLLALQVSQNNLSGAIPNYIS 269
Query: 153 QLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIP 196
+ L++ DLS N+ +GV+P F +L +++ ++S N L G P
Sbjct: 270 RFNKLEKLDLSKNRFSGVVPQGFVNLTNINNLDLSHNLLTGQFP 313
Score = 63.2 bits (152), Expect = 1e-10
Identities = 41/109 (37%), Positives = 53/109 (48%), Gaps = 28/109 (25%)
Query: 136 LDLGSNSLSGSIPTFLGQLKGLQEF-----------------------DLSNNKLTGVIP 172
+DL N +SGS FL Q K L EF DLS N + G +
Sbjct: 802 IDLSENEISGSPAKFLSQXKYLMEFRAAGNKLRFDLGKLTFVRTLETLDLSRNLIFGRVL 861
Query: 173 ASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 221
A+F L ++ NVS N LCG +P ++KF S F N CLCG+PL+ C
Sbjct: 862 ATFAGLKTM---NVSQNHLCGKLP--VTKFPASXFAGNDCLCGSPLSPC 905
Score = 58.5 bits (140), Expect = 3e-09
Identities = 42/142 (29%), Positives = 67/142 (46%), Gaps = 10/142 (7%)
Query: 78 PLPGRITVVT-----ISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQR 132
P+P I+ +T I G L+GT+P +L + L L + +++G IP+ F ++
Sbjct: 166 PIPNSISNLTRLSYLIFGGNLLTGTIPLGIANLKLMQNLQLGDN-RLSGTIPDIFESMKL 224
Query: 133 LQKLDLGSNSLSGSIPTFLGQLKG-LQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQL 191
L+ LDL SN G +P + L L +S N L+G IP L + ++S N+
Sbjct: 225 LKFLDLSSNEFYGKLPLSIATLAPTLLALQVSQNNLSGAIPNYISRFNKLEKLDLSKNRF 284
Query: 192 CGAIPTG---LSKFAKSSFDHN 210
G +P G L+ HN
Sbjct: 285 SGVVPQGFVNLTNINNLDLSHN 306
Score = 51.6 bits (122), Expect = 3e-07
Identities = 33/99 (33%), Positives = 49/99 (49%), Gaps = 3/99 (3%)
Query: 93 GLSGTLPAEFGDL-PYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFL 151
G G LP L P L +L L++ ++G IPN S+ + L L L N SG +P
Sbjct: 662 GFFGRLPPSIASLAPTLYYLDLSQN-NLSGTIPNYLSRFEALSTLVLSKNKYSGVVPMSF 720
Query: 152 GQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQ 190
L + DLS+N LTG P S+ + ++S+N+
Sbjct: 721 TNLINITNLDLSHNLLTGPFPV-LKSINGIESLDLSYNK 758
Score = 43.1 bits (100), Expect = 1e-04
Identities = 30/95 (31%), Positives = 44/95 (45%), Gaps = 28/95 (29%)
Query: 136 LDLGSNSLSGSIPTFLGQLKGLQEF-----------------------DLSNNKLTGVIP 172
+DL N +SGS+ FL + + L EF DLS N + G +P
Sbjct: 373 IDLSKNEISGSLERFLNETRYLLEFRAAENKLRFDMGNLTFPRTLKTLDLSRNLVFGKVP 432
Query: 173 ASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSF 207
++ L + N+S N LCG +PT +KF S+F
Sbjct: 433 V---TVAGLQRLNLSQNHLCGELPT--TKFPASAF 462
>At4g30520 receptor-like kinase homolog
Length = 648
Score = 96.7 bits (239), Expect = 9e-21
Identities = 63/177 (35%), Positives = 92/177 (51%), Gaps = 14/177 (7%)
Query: 43 LLKIRDHFGGPNGRLSDWDN-GTDCCSDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAE 101
L+ IR++ P+G L++WD D CS W+ + C P + + + LSG L
Sbjct: 41 LISIRNNLHDPHGALNNWDEFSVDPCS-WAMITCS---PDNLVIGLGAPSQSLSGGLSES 96
Query: 102 FGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFD 161
G+L L +SL + ++G IP L +LQ LDL +N SG IP + QL LQ
Sbjct: 97 IGNLTNLRQVSL-QNNNISGKIPPELGFLPKLQTLDLSNNRFSGDIPVSIDQLSSLQYLR 155
Query: 162 LSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPL 218
L+NN L+G PAS +P LS ++S+N L G +P KF +F+ + G PL
Sbjct: 156 LNNNSLSGPFPASLSQIPHLSFLDLSYNNLSGPVP----KFPARTFN----VAGNPL 204
>At4g20140 leucine rich repeat-like protein
Length = 1232
Score = 96.3 bits (238), Expect = 1e-20
Identities = 58/142 (40%), Positives = 80/142 (55%), Gaps = 3/142 (2%)
Query: 81 GRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQK-LDLG 139
G + V+ + + SG+LP G L L L L+ +TG IP +LQ LQ LDL
Sbjct: 719 GALNVLNLDKNQ-FSGSLPQAMGKLSKLYELRLSRN-SLTGEIPVEIGQLQDLQSALDLS 776
Query: 140 SNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGL 199
N+ +G IP+ +G L L+ DLS+N+LTG +P S G + SL NVSFN L G +
Sbjct: 777 YNNFTGDIPSTIGTLSKLETLDLSHNQLTGEVPGSVGDMKSLGYLNVSFNNLGGKLKKQF 836
Query: 200 SKFAKSSFDHNKCLCGAPLAAC 221
S++ SF N LCG+PL+ C
Sbjct: 837 SRWPADSFLGNTGLCGSPLSRC 858
Score = 78.2 bits (191), Expect = 3e-15
Identities = 47/117 (40%), Positives = 65/117 (55%), Gaps = 2/117 (1%)
Query: 83 ITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNS 142
+TV T + L+GT+PAE G L L L+LA +TG IP+ ++ +LQ L L +N
Sbjct: 217 LTVFTAAENM-LNGTIPAELGRLENLEILNLANN-SLTGEIPSQLGEMSQLQYLSLMANQ 274
Query: 143 LSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGL 199
L G IP L L LQ DLS N LTG IP F ++ L ++ N L G++P +
Sbjct: 275 LQGLIPKSLADLGNLQTLDLSANNLTGEIPEEFWNMSQLLDLVLANNHLSGSLPKSI 331
Score = 77.0 bits (188), Expect = 7e-15
Identities = 44/114 (38%), Positives = 61/114 (52%), Gaps = 1/114 (0%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
L G +P G+L L L+LA ++TGPIP+ +L R+Q L L N L G IP LG
Sbjct: 155 LVGDIPETLGNLVNLQMLALASC-RLTGPIPSQLGRLVRVQSLILQDNYLEGPIPAELGN 213
Query: 154 LKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSF 207
L F + N L G IPA G L +L N++ N L G IP+ L + ++ +
Sbjct: 214 CSDLTVFTAAENMLNGTIPAELGRLENLEILNLANNSLTGEIPSQLGEMSQLQY 267
Score = 76.3 bits (186), Expect = 1e-14
Identities = 51/170 (30%), Positives = 85/170 (50%), Gaps = 5/170 (2%)
Query: 38 NDKATLLKIRDHF---GGPNGRLSDWDNGTDCCSDWSFVGCGKPLPGRITVVTISRGWGL 94
ND TLL+++ + L W++ W+ V C R+ + ++ G GL
Sbjct: 25 NDLQTLLEVKKSLVTNPQEDDPLRQWNSDNINYCSWTGVTCDNTGLFRVIALNLT-GLGL 83
Query: 95 SGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQL 154
+G++ FG L L L+ + GPIP + S L L+ L L SN L+G IP+ LG L
Sbjct: 84 TGSISPWFGRFDNLIHLDLSSN-NLVGPIPTALSNLTSLESLFLFSNQLTGEIPSQLGSL 142
Query: 155 KGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAK 204
++ + +N+L G IP + G+L +L ++ +L G IP+ L + +
Sbjct: 143 VNIRSLRIGDNELVGDIPETLGNLVNLQMLALASCRLTGPIPSQLGRLVR 192
Score = 74.7 bits (182), Expect = 4e-14
Identities = 46/113 (40%), Positives = 62/113 (54%), Gaps = 1/113 (0%)
Query: 96 GTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLK 155
G +P G L LN L L + ++ G +P S +L LDL N LSGSIP+ G LK
Sbjct: 470 GEIPPSIGRLKELNLLHLRQN-ELVGGLPASLGNCHQLNILDLADNQLSGSIPSSFGFLK 528
Query: 156 GLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSFD 208
GL++ L NN L G +P S SL +L++ N+S N+L G I + SFD
Sbjct: 529 GLEQLMLYNNSLQGNLPDSLISLRNLTRINLSHNRLNGTIHPLCGSSSYLSFD 581
Score = 73.2 bits (178), Expect = 1e-13
Identities = 41/103 (39%), Positives = 59/103 (56%), Gaps = 1/103 (0%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
L G +PAE G+ L + AE + G IP +L+ L+ L+L +NSL+G IP+ LG+
Sbjct: 203 LEGPIPAELGNCSDLTVFTAAEN-MLNGTIPAELGRLENLEILNLANNSLTGEIPSQLGE 261
Query: 154 LKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIP 196
+ LQ L N+L G+IP S L +L ++S N L G IP
Sbjct: 262 MSQLQYLSLMANQLQGLIPKSLADLGNLQTLDLSANNLTGEIP 304
Score = 70.5 bits (171), Expect = 7e-13
Identities = 42/112 (37%), Positives = 61/112 (53%), Gaps = 1/112 (0%)
Query: 93 GLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLG 152
G +P E G+ L+ L L + ++TG IP + K++ L LD+ SN+L+G+IP L
Sbjct: 586 GFEDEIPLELGNSQNLDRLRLGKN-QLTGKIPWTLGKIRELSLLDMSSNALTGTIPLQLV 644
Query: 153 QLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAK 204
K L DL+NN L+G IP G L L + +S NQ ++PT L K
Sbjct: 645 LCKKLTHIDLNNNFLSGPIPPWLGKLSQLGELKLSSNQFVESLPTELFNCTK 696
Score = 68.9 bits (167), Expect = 2e-12
Identities = 42/109 (38%), Positives = 63/109 (57%), Gaps = 2/109 (1%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
L+G +P++ G++ L +LSL ++ G IP S + L LQ LDL +N+L+G IP
Sbjct: 251 LTGEIPSQLGEMSQLQYLSLMAN-QLQGLIPKSLADLGNLQTLDLSANNLTGEIPEEFWN 309
Query: 154 LKGLQEFDLSNNKLTGVIPASF-GSLPSLSQFNVSFNQLCGAIPTGLSK 201
+ L + L+NN L+G +P S + +L Q +S QL G IP LSK
Sbjct: 310 MSQLLDLVLANNHLSGSLPKSICSNNTNLEQLVLSGTQLSGEIPVELSK 358
Score = 66.2 bits (160), Expect = 1e-11
Identities = 39/108 (36%), Positives = 58/108 (53%), Gaps = 1/108 (0%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
L+G +P++ G L + L + + ++ G IP + L LQ L L S L+G IP+ LG+
Sbjct: 131 LTGEIPSQLGSLVNIRSLRIGDN-ELVGDIPETLGNLVNLQMLALASCRLTGPIPSQLGR 189
Query: 154 LKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSK 201
L +Q L +N L G IPA G+ L+ F + N L G IP L +
Sbjct: 190 LVRVQSLILQDNYLEGPIPAELGNCSDLTVFTAAENMLNGTIPAELGR 237
Score = 65.9 bits (159), Expect = 2e-11
Identities = 46/136 (33%), Positives = 67/136 (48%), Gaps = 24/136 (17%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
L G LPA G+ LN L LA+ +++G IP+SF L+ L++L L +NSL G++P L
Sbjct: 492 LVGGLPASLGNCHQLNILDLADN-QLSGSIPSSFGFLKGLEQLMLYNNSLQGNLPDSLIS 550
Query: 154 LKGLQEFDLSNNKLTGVI-----------------------PASFGSLPSLSQFNVSFNQ 190
L+ L +LS+N+L G I P G+ +L + + NQ
Sbjct: 551 LRNLTRINLSHNRLNGTIHPLCGSSSYLSFDVTNNGFEDEIPLELGNSQNLDRLRLGKNQ 610
Query: 191 LCGAIPTGLSKFAKSS 206
L G IP L K + S
Sbjct: 611 LTGKIPWTLGKIRELS 626
Score = 65.9 bits (159), Expect = 2e-11
Identities = 45/119 (37%), Positives = 58/119 (47%), Gaps = 5/119 (4%)
Query: 97 TLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKG 156
+LP E + L LSL + + G IP L L L+L N SGS+P +G+L
Sbjct: 686 SLPTELFNCTKLLVLSL-DGNSLNGSIPQEIGNLGALNVLNLDKNQFSGSLPQAMGKLSK 744
Query: 157 LQEFDLSNNKLTGVIPASFGSLPSL-SQFNVSFNQLCGAIPT---GLSKFAKSSFDHNK 211
L E LS N LTG IP G L L S ++S+N G IP+ LSK HN+
Sbjct: 745 LYELRLSRNSLTGEIPVEIGQLQDLQSALDLSYNNFTGDIPSTIGTLSKLETLDLSHNQ 803
Score = 64.3 bits (155), Expect = 5e-11
Identities = 43/124 (34%), Positives = 61/124 (48%), Gaps = 3/124 (2%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
L G LP E L L L L E + +G IP L+ +D+ N G IP +G+
Sbjct: 420 LEGKLPKEISALRKLEVLFLYEN-RFSGEIPQEIGNCTSLKMIDMFGNHFEGEIPPSIGR 478
Query: 154 LKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPT--GLSKFAKSSFDHNK 211
LK L L N+L G +PAS G+ L+ +++ NQL G+IP+ G K + +N
Sbjct: 479 LKELNLLHLRQNELVGGLPASLGNCHQLNILDLADNQLSGSIPSSFGFLKGLEQLMLYNN 538
Query: 212 CLCG 215
L G
Sbjct: 539 SLQG 542
Score = 61.2 bits (147), Expect = 4e-10
Identities = 38/111 (34%), Positives = 55/111 (49%), Gaps = 1/111 (0%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
L+GT+P + L + L ++GPIP KL +L +L L SN S+PT L
Sbjct: 635 LTGTIPLQLVLCKKLTHIDLNNN-FLSGPIPPWLGKLSQLGELKLSSNQFVESLPTELFN 693
Query: 154 LKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAK 204
L L N L G IP G+L +L+ N+ NQ G++P + K +K
Sbjct: 694 CTKLLVLSLDGNSLNGSIPQEIGNLGALNVLNLDKNQFSGSLPQAMGKLSK 744
Score = 60.5 bits (145), Expect = 7e-10
Identities = 45/135 (33%), Positives = 64/135 (47%), Gaps = 24/135 (17%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAE------MPK------------------VTGPIPNSFSK 129
L+G +P EF ++ L L LA +PK ++G IP SK
Sbjct: 299 LTGEIPEEFWNMSQLLDLVLANNHLSGSLPKSICSNNTNLEQLVLSGTQLSGEIPVELSK 358
Query: 130 LQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFN 189
Q L++LDL +NSL+GSIP L +L L + L NN L G + S +L +L + N
Sbjct: 359 CQSLKQLDLSNNSLAGSIPEALFELVELTDLYLHNNTLEGTLSPSISNLTNLQWLVLYHN 418
Query: 190 QLCGAIPTGLSKFAK 204
L G +P +S K
Sbjct: 419 NLEGKLPKEISALRK 433
Score = 60.5 bits (145), Expect = 7e-10
Identities = 37/108 (34%), Positives = 52/108 (47%), Gaps = 1/108 (0%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
LSG +P G L L L L+ V +P +L L L NSL+GSIP +G
Sbjct: 659 LSGPIPPWLGKLSQLGELKLSSNQFVES-LPTELFNCTKLLVLSLDGNSLNGSIPQEIGN 717
Query: 154 LKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSK 201
L L +L N+ +G +P + G L L + +S N L G IP + +
Sbjct: 718 LGALNVLNLDKNQFSGSLPQAMGKLSKLYELRLSRNSLTGEIPVEIGQ 765
Score = 56.2 bits (134), Expect = 1e-08
Identities = 41/129 (31%), Positives = 59/129 (44%), Gaps = 24/129 (18%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSI------ 147
LSG++P+ FG L L L L + G +P+S L+ L +++L N L+G+I
Sbjct: 516 LSGSIPSSFGFLKGLEQLMLYNN-SLQGNLPDSLISLRNLTRINLSHNRLNGTIHPLCGS 574
Query: 148 -----------------PTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQ 190
P LG + L L N+LTG IP + G + LS ++S N
Sbjct: 575 SSYLSFDVTNNGFEDEIPLELGNSQNLDRLRLGKNQLTGKIPWTLGKIRELSLLDMSSNA 634
Query: 191 LCGAIPTGL 199
L G IP L
Sbjct: 635 LTGTIPLQL 643
Score = 48.1 bits (113), Expect = 4e-06
Identities = 33/106 (31%), Positives = 48/106 (45%), Gaps = 1/106 (0%)
Query: 91 GWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTF 150
G LSG +P E L L L+ + G IP + +L L L L +N+L G++
Sbjct: 345 GTQLSGEIPVELSKCQSLKQLDLSNN-SLAGSIPEALFELVELTDLYLHNNTLEGTLSPS 403
Query: 151 LGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIP 196
+ L LQ L +N L G +P +L L + N+ G IP
Sbjct: 404 ISNLTNLQWLVLYHNNLEGKLPKEISALRKLEVLFLYENRFSGEIP 449
>At2g26380 putative disease resistance protein
Length = 480
Score = 96.3 bits (238), Expect = 1e-20
Identities = 63/195 (32%), Positives = 94/195 (47%), Gaps = 10/195 (5%)
Query: 12 HLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFG-GPNGRLSDWDNGTDCCSDW 70
H +F + I+ N +C+ +D+A LL + P+G LS W GTDCCS W
Sbjct: 7 HNFFIFTAVIFLRCLNPTAAA-TCHPDDEAGLLAFKSGITKDPSGILSTWKKGTDCCS-W 64
Query: 71 SFVGCGKPLPGRITVVTI-----SRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPN 125
+ V C P R+ V+TI G LSGT+ L +L + + +TGP P
Sbjct: 65 NGVSC--PNGNRVVVLTIRIESDDAGIFLSGTISPSLAKLQHLEGVVFINLKNITGPFPP 122
Query: 126 SFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFN 185
+L L+ + L + LSG +P +G L L + N+ G IP+S +L L+ N
Sbjct: 123 FLFRLPHLKYVYLENTRLSGPLPANIGALNRLDTLTVKGNRFIGSIPSSISNLTRLNYLN 182
Query: 186 VSFNQLCGAIPTGLS 200
+ N L G IP G++
Sbjct: 183 LGGNLLTGTIPLGIA 197
Score = 68.9 bits (167), Expect = 2e-12
Identities = 39/104 (37%), Positives = 58/104 (55%), Gaps = 2/104 (1%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQR-LQKLDLGSNSLSGSIPTFLG 152
LSGT+P F + L L+L+ + +G +P S + L L L+LG N+LSGSIP++L
Sbjct: 212 LSGTIPDIFKSMTNLRILTLSRN-RFSGKLPPSIASLAPVLAFLELGQNNLSGSIPSYLS 270
Query: 153 QLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIP 196
+ L DLS N+ +G +P S L ++ N+S N L P
Sbjct: 271 RFVALDTLDLSKNRFSGAVPKSLAKLTKIANINLSHNLLTNPFP 314
Score = 68.2 bits (165), Expect = 3e-12
Identities = 45/132 (34%), Positives = 71/132 (53%), Gaps = 9/132 (6%)
Query: 78 PLPG------RITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ 131
PLP R+ +T+ +G G++P+ +L LN+L+L +TG IP + L+
Sbjct: 143 PLPANIGALNRLDTLTV-KGNRFIGSIPSSISNLTRLNYLNLGGN-LLTGTIPLGIANLK 200
Query: 132 RLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSL-PSLSQFNVSFNQ 190
+ L+L N LSG+IP + L+ LS N+ +G +P S SL P L+ + N
Sbjct: 201 LISNLNLDGNRLSGTIPDIFKSMTNLRILTLSRNRFSGKLPPSIASLAPVLAFLELGQNN 260
Query: 191 LCGAIPTGLSKF 202
L G+IP+ LS+F
Sbjct: 261 LSGSIPSYLSRF 272
Score = 60.1 bits (144), Expect = 9e-10
Identities = 35/112 (31%), Positives = 60/112 (53%), Gaps = 2/112 (1%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
L+GT+P +L ++ L+L + +++G IP+ F + L+ L L N SG +P +
Sbjct: 188 LTGTIPLGIANLKLISNLNL-DGNRLSGTIPDIFKSMTNLRILTLSRNRFSGKLPPSIAS 246
Query: 154 LKGLQEF-DLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAK 204
L + F +L N L+G IP+ +L ++S N+ GA+P L+K K
Sbjct: 247 LAPVLAFLELGQNNLSGSIPSYLSRFVALDTLDLSKNRFSGAVPKSLAKLTK 298
Score = 58.2 bits (139), Expect = 3e-09
Identities = 39/109 (35%), Positives = 53/109 (47%), Gaps = 28/109 (25%)
Query: 136 LDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLT-----------------------GVIP 172
+DL N +SGS FL + L+EF +S NKL G +P
Sbjct: 375 IDLSDNEISGSPLRFLKGAEQLREFRMSGNKLRFDLRKLSFSTTLETLDLSRNLVFGKVP 434
Query: 173 ASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 221
A L +L N+S N LCG +P ++KF +S F N CLCG+PL+ C
Sbjct: 435 ARVAGLKTL---NLSQNHLCGKLP--VTKFPESVFAGNDCLCGSPLSHC 478
>At1g31420 unknown protein
Length = 592
Score = 96.3 bits (238), Expect = 1e-20
Identities = 73/203 (35%), Positives = 100/203 (48%), Gaps = 11/203 (5%)
Query: 16 LFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVGC 75
L L S+ S SN E + D LL R+ + + W +W+ V C
Sbjct: 14 LLLISLLCSLSN----ESQAISPDGEALLSFRNAVTRSDSFIHQWRPEDPDPCNWNGVTC 69
Query: 76 GKPLPGRITV-VTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQ 134
IT+ +T + + G LP + G L +L L L + G IP + L+
Sbjct: 70 DAKTKRVITLNLTYHK---IMGPLPPDIGKLDHLRLLMLHNNA-LYGAIPTALGNCTALE 125
Query: 135 KLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGA 194
++ L SN +G IP +G L GLQ+ D+S+N L+G IPAS G L LS FNVS N L G
Sbjct: 126 EIHLQSNYFTGPIPAEMGDLPGLQKLDMSSNTLSGPIPASLGQLKKLSNFNVSNNFLVGQ 185
Query: 195 IPTG--LSKFAKSSFDHNKCLCG 215
IP+ LS F+K+SF N LCG
Sbjct: 186 IPSDGVLSGFSKNSFIGNLNLCG 208
>At3g02880 putative protein kinase
Length = 627
Score = 95.9 bits (237), Expect = 1e-20
Identities = 78/233 (33%), Positives = 112/233 (47%), Gaps = 37/233 (15%)
Query: 11 LHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDW 70
L L+++FL Y + +D +D+ LL +R+ GR W+ +W
Sbjct: 7 LSLSVVFLFVFYLAAVTSD------LESDRRALLAVRNSV---RGRPLLWNMSASSPCNW 57
Query: 71 SFVGCGKPLPGRITVVTISRGWGLSGTLP-AEFGDLPYLNFLSLAEMPKVTGPIPNSFSK 129
V C GR+T + + G GL G+LP G+L L LSL ++GPIP+ FS
Sbjct: 58 HGVHCDA---GRVTALRLP-GSGLFGSLPIGGIGNLTQLKTLSL-RFNSLSGPIPSDFSN 112
Query: 130 LQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASF-------------- 175
L L+ L L N+ SG IP+ L L + +L NK +G IP +
Sbjct: 113 LVLLRYLYLQGNAFSGEIPSLLFTLPSIIRINLGENKFSGRIPDNVNSATRLVTLYLERN 172
Query: 176 ---GSLPS----LSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 221
G +P L QFNVS NQL G+IP+ LS + +++F+ N LCG PL C
Sbjct: 173 QLSGPIPEITLPLQQFNVSSNQLNGSIPSSLSSWPRTAFEGN-TLCGKPLDTC 224
>At1g33600 unknown protein
Length = 478
Score = 95.9 bits (237), Expect = 1e-20
Identities = 61/173 (35%), Positives = 87/173 (50%), Gaps = 10/173 (5%)
Query: 34 SCNANDKATLLKIRDHFG-GPNGRLSDWDNGTDCCSDWSFVGCGKPLPGRITVVTIS--- 89
+C+ +D+A LL + P G LS W GTDCCS W VGC L R+T +TI+
Sbjct: 27 TCHPDDEAGLLAFKSGITQDPTGILSSWKKGTDCCS-WKGVGC---LTNRVTGLTINGQS 82
Query: 90 --RGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSI 147
G LSGT+ L +L + + +TG P +L ++++ ++ LSG +
Sbjct: 83 DVTGSFLSGTISPSLAKLQHLVGIYFTNLRNITGSFPQFLFQLPNVKQVYFTNSRLSGPL 142
Query: 148 PTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLS 200
P +G L L E L N TG IP+S +L L N+ N L G IP GL+
Sbjct: 143 PANIGALSELGELSLDGNLFTGPIPSSISNLTRLYLLNLGDNLLTGTIPLGLA 195
Score = 76.3 bits (186), Expect = 1e-14
Identities = 44/104 (42%), Positives = 61/104 (58%), Gaps = 2/104 (1%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQR-LQKLDLGSNSLSGSIPTFLG 152
LS T+P F + L L+L+ K +G +P S + L+ L LDL N+LSG+IPTFL
Sbjct: 210 LSETIPDIFKSMQKLQSLTLSRN-KFSGNLPPSIASLKPILNYLDLSQNNLSGTIPTFLS 268
Query: 153 QLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIP 196
K L DLS N+ +GV+P S ++P L N+S N L G +P
Sbjct: 269 NFKVLDSLDLSRNRFSGVVPKSLANMPKLFHLNLSHNFLTGPLP 312
Score = 63.9 bits (154), Expect = 6e-11
Identities = 36/112 (32%), Positives = 58/112 (51%), Gaps = 2/112 (1%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
L+GT+P +L L L+ +++ IP+ F +Q+LQ L L N SG++P +
Sbjct: 186 LTGTIPLGLANLKILLSLNFGNN-RLSETIPDIFKSMQKLQSLTLSRNKFSGNLPPSIAS 244
Query: 154 LKGLQEF-DLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAK 204
LK + + DLS N L+G IP + L ++S N+ G +P L+ K
Sbjct: 245 LKPILNYLDLSQNNLSGTIPTFLSNFKVLDSLDLSRNRFSGVVPKSLANMPK 296
Score = 60.8 bits (146), Expect = 5e-10
Identities = 39/106 (36%), Positives = 52/106 (48%), Gaps = 22/106 (20%)
Query: 136 LDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLT---GVIPAS-------------FGSLP 179
+DL N +SGS+ F L EF S NKL G + S FG +P
Sbjct: 373 IDLSENEISGSLTWFFNLAHNLYEFQASGNKLRFDMGKLNLSERLESLDLSRNLIFGKVP 432
Query: 180 ----SLSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 221
L + N+S N LCG +P ++KF S+F N CLCG+PL+ C
Sbjct: 433 MTVAKLQKLNLSHNHLCGKLP--VTKFPASAFVGNDCLCGSPLSPC 476
Score = 58.9 bits (141), Expect = 2e-09
Identities = 39/110 (35%), Positives = 60/110 (54%), Gaps = 4/110 (3%)
Query: 82 RITVVTISRGWGLSGTLPAEFGDL-PYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGS 140
++ +T+SR SG LP L P LN+L L++ ++G IP S + L LDL
Sbjct: 223 KLQSLTLSRN-KFSGNLPPSIASLKPILNYLDLSQN-NLSGTIPTFLSNFKVLDSLDLSR 280
Query: 141 NSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQ 190
N SG +P L + L +LS+N LTG +PA ++ L+ ++S+NQ
Sbjct: 281 NRFSGVVPKSLANMPKLFHLNLSHNFLTGPLPA-MKNVDGLATLDLSYNQ 329
>At1g33590 unknown protein
Length = 477
Score = 95.5 bits (236), Expect = 2e-20
Identities = 58/180 (32%), Positives = 94/180 (52%), Gaps = 9/180 (5%)
Query: 34 SCNANDKATLLKIRDHFG-GPNGRLSDWDNGTDCCSDWSFVGCGKPLPGRITVVTIS--- 89
+C+ +D+A LL + P+G LS W GT CCS W+ V C R++ ++++
Sbjct: 26 TCHPDDEAGLLAFKAGITRDPSGILSSWKKGTACCS-WNGVTC--LTTDRVSALSVAGQA 82
Query: 90 --RGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSI 147
G LSGTL L +L+ + ++ +TG P +L L+ + + +N LSG++
Sbjct: 83 DVAGSFLSGTLSPSLAKLKHLDGIYFTDLKNITGSFPQFLFQLPNLKYVYIENNRLSGTL 142
Query: 148 PTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSF 207
P +G L L+ F L N+ TG IP+S +L L+Q + N L G IP G++ S+
Sbjct: 143 PANIGALSQLEAFSLEGNRFTGPIPSSISNLTLLTQLKLGNNLLTGTIPLGVANLKLMSY 202
Score = 83.2 bits (204), Expect = 1e-16
Identities = 48/110 (43%), Positives = 63/110 (56%), Gaps = 1/110 (0%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
LSGTLPA G L L SL E + TGPIP+S S L L +L LG+N L+G+IP +
Sbjct: 138 LSGTLPANIGALSQLEAFSL-EGNRFTGPIPSSISNLTLLTQLKLGNNLLTGTIPLGVAN 196
Query: 154 LKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFA 203
LK + +L N+LTG IP F S+P L +S N G +P ++ A
Sbjct: 197 LKLMSYLNLGGNRLTGTIPDIFKSMPELRSLTLSRNGFSGNLPPSIASLA 246
Score = 71.2 bits (173), Expect = 4e-13
Identities = 41/104 (39%), Positives = 59/104 (56%), Gaps = 2/104 (1%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQK-LDLGSNSLSGSIPTFLG 152
L+GT+P F +P L L+L+ +G +P S + L + + L+LG N LSG+IP FL
Sbjct: 210 LTGTIPDIFKSMPELRSLTLSRNG-FSGNLPPSIASLAPILRFLELGHNKLSGTIPNFLS 268
Query: 153 QLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIP 196
K L DLS N+ +GVIP SF +L + ++S N L P
Sbjct: 269 NFKALDTLDLSKNRFSGVIPKSFANLTKIFNLDLSHNLLTDPFP 312
Score = 64.7 bits (156), Expect = 4e-11
Identities = 38/121 (31%), Positives = 63/121 (51%), Gaps = 5/121 (4%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQ 153
L+GT+P +L +++L+L ++TG IP+ F + L+ L L N SG++P +
Sbjct: 186 LTGTIPLGVANLKLMSYLNLGGN-RLTGTIPDIFKSMPELRSLTLSRNGFSGNLPPSIAS 244
Query: 154 LKGLQEF-DLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIP---TGLSKFAKSSFDH 209
L + F +L +NKL+G IP + +L ++S N+ G IP L+K H
Sbjct: 245 LAPILRFLELGHNKLSGTIPNFLSNFKALDTLDLSKNRFSGVIPKSFANLTKIFNLDLSH 304
Query: 210 N 210
N
Sbjct: 305 N 305
Score = 60.8 bits (146), Expect = 5e-10
Identities = 39/109 (35%), Positives = 53/109 (47%), Gaps = 28/109 (25%)
Query: 136 LDLGSNSLSGSIPTFLGQLKGLQEF-----------------------DLSNNKLTGVIP 172
+DL N ++GS FL Q + L EF D+S N + G +P
Sbjct: 372 IDLSENEITGSPARFLNQTEYLVEFKAAGNKLRFDMGKLTFAKTLTTLDISRNLVFGKVP 431
Query: 173 ASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 221
A L +L NVS N LCG +P ++KF S+F N CLCG+PL+ C
Sbjct: 432 AMVAGLKTL---NVSHNHLCGKLP--VTKFPASAFVGNDCLCGSPLSPC 475
>At5g21090 leucine-rich repeat protein
Length = 218
Score = 95.1 bits (235), Expect = 3e-20
Identities = 64/178 (35%), Positives = 91/178 (50%), Gaps = 6/178 (3%)
Query: 43 LLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEF 102
L +R P+ L WD W V C + R+T V + LSG L E
Sbjct: 34 LYALRRSLTDPDHVLQSWDPTLVNPCTWFHVTCNQD--NRVTRVDLGNS-NLSGHLAPEL 90
Query: 103 GDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDL 162
G L +L +L L + + G IP+ L+ L LDL +N+L+G +PT LG+LK L L
Sbjct: 91 GKLEHLQYLELYKN-NIQGTIPSELGNLKNLISLDLYNNNLTGIVPTSLGKLKSLVFLRL 149
Query: 163 SNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPL 218
++N+LTG IP + ++PSL +VS N LCG IPT + +F++N L G L
Sbjct: 150 NDNRLTGPIPRALTAIPSLKVVDVSSNDLCGTIPTNGPFAHIPLQNFENNPRLEGPEL 207
>At1g68400 receptor kinase like protein
Length = 670
Score = 95.1 bits (235), Expect = 3e-20
Identities = 77/235 (32%), Positives = 106/235 (44%), Gaps = 35/235 (14%)
Query: 6 NKFLLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTD 65
NK LLL L +L S + S+S+ D TLL + G+L+ W+ T+
Sbjct: 8 NKHLLLSLLILLQSCLLSSSSSTDS----------ETLLNFK-LTADSTGKLNSWNTTTN 56
Query: 66 CCSDWSFVGCGKPLPGRITVVTIS-------------------RGWGLSGTLPAEFGDLP 106
C W+ V C + R+ + I+ + LSG +P +L
Sbjct: 57 PCQ-WTGVSCNRNRVTRLVLEDINLTGSISSLTSLTSLRVLSLKHNNLSGPIP-NLSNLT 114
Query: 107 YLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNK 166
L L L+ + +G P S + L RL +LDL N+ SG IP L L L L +N+
Sbjct: 115 ALKLLFLSNN-QFSGNFPTSITSLTRLYRLDLSFNNFSGQIPPDLTDLTHLLTLRLESNR 173
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKFAKSSFDHNKCLCGAPLAAC 221
+G IP +L L FNVS N G IP LS+F +S F N LCGAPL C
Sbjct: 174 FSGQIPNI--NLSDLQDFNVSGNNFNGQIPNSLSQFPESVFTQNPSLCGAPLLKC 226
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.138 0.429
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,365,828
Number of Sequences: 26719
Number of extensions: 244095
Number of successful extensions: 5644
Number of sequences better than 10.0: 419
Number of HSP's better than 10.0 without gapping: 370
Number of HSP's successfully gapped in prelim test: 49
Number of HSP's that attempted gapping in prelim test: 572
Number of HSP's gapped (non-prelim): 2613
length of query: 221
length of database: 11,318,596
effective HSP length: 95
effective length of query: 126
effective length of database: 8,780,291
effective search space: 1106316666
effective search space used: 1106316666
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146750.10


