

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146747.7 - phase: 0 /pseudo
(305 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
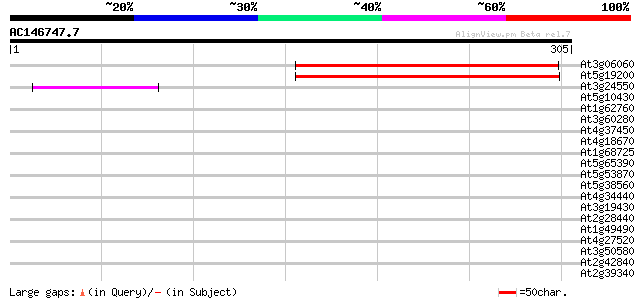
Score E
Sequences producing significant alignments: (bits) Value
At3g06060 unknown protein 199 2e-51
At5g19200 FVT1 - like protein 185 3e-47
At3g24550 protein kinase, putative 43 2e-04
At5g10430 AtAGP4 41 0.001
At1g62760 hypothetical protein 40 0.001
At3g60280 uclacyanin 3 40 0.002
At4g37450 arabinogalactan protein AGP18 39 0.005
At4g18670 extensin-like protein 39 0.005
At1g68725 putative arabinogalactan protein AGP19 39 0.005
At5g65390 arabinogalactan protein AGP7 38 0.006
At5g53870 predicted GPI-anchored protein 38 0.006
At5g38560 putative protein 38 0.006
At4g34440 serine/threonine protein kinase - like 38 0.006
At3g19430 putative late embryogenesis abundant protein 38 0.006
At2g28440 En/Spm-like transposon protein 38 0.006
At1g49490 hypothetical protein 38 0.008
At4g27520 early nodulin-like 2 predicted GPI-anchored protein 37 0.010
At3g50580 proline-rich protein 37 0.010
At2g42840 En/Spm-like transposon protein 37 0.010
At2g39340 unknown protein 37 0.010
>At3g06060 unknown protein
Length = 326
Score = 199 bits (506), Expect = 2e-51
Identities = 94/143 (65%), Positives = 120/143 (83%)
Query: 156 ITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTPGLVEENKRKPELTKIIA 215
+ ++AYSASKFGL+GLA+ALQQEVI D+IHV+LIFPPDT+TPG EE K +PE+T IIA
Sbjct: 182 VYGYAAYSASKFGLQGLAQALQQEVISDDIHVTLIFPPDTNTPGFEEEQKSRPEVTAIIA 241
Query: 216 ASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLSPQRSFLMAFVEVIAAGI 275
ASSG M+ +EVA+KA DGI++G+F +SCN EG LSLAT+G+SPQRSF +AF+EVI AG
Sbjct: 242 ASSGSMETEEVAKKAMDGIKAGNFTVSCNFEGFLLSLATTGMSPQRSFWLAFLEVITAGP 301
Query: 276 MRIAALCFQWNWYGSIEKWHKQR 298
+R+ AL FQW+WY +IEKW K +
Sbjct: 302 IRLIALFFQWDWYKAIEKWSKTK 324
>At5g19200 FVT1 - like protein
Length = 331
Score = 185 bits (469), Expect = 3e-47
Identities = 86/144 (59%), Positives = 115/144 (79%)
Query: 156 ITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTPGLVEENKRKPELTKIIA 215
I ++AYSASKFGL+GLA+ALQQEVI D IHV+L+FPPDTDTPG +E K++PELT IIA
Sbjct: 180 IYGYTAYSASKFGLQGLAQALQQEVISDGIHVTLLFPPDTDTPGFEQELKKRPELTSIIA 239
Query: 216 ASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLSPQRSFLMAFVEVIAAGI 275
ASSG MK +EVA+ FDGI++G F ++C+ G LS+A++G+SPQ SF +A EV+ G+
Sbjct: 240 ASSGSMKTNEVAKICFDGIKAGKFTVTCHFIGFLLSIASTGMSPQGSFWLALTEVMFGGL 299
Query: 276 MRIAALCFQWNWYGSIEKWHKQRK 299
+R+A+L FQW WY +IEKW ++ K
Sbjct: 300 IRLASLVFQWQWYKTIEKWSQRNK 323
>At3g24550 protein kinase, putative
Length = 652
Score = 43.1 bits (100), Expect = 2e-04
Identities = 27/71 (38%), Positives = 37/71 (52%), Gaps = 2/71 (2%)
Query: 13 SSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPP-LMAHVSPSWHDP-SPNSK 70
SSP S P+P S +P + SP+A + PSPT P + SP+ P +P+
Sbjct: 57 SSPLPPSLPPPSPPGSLTPPLPQPSPSAPITPSPPSPTTPSNPRSPPSPNQGPPNTPSGS 116
Query: 71 KLRTPSNTPPA 81
RTPSNT P+
Sbjct: 117 TPRTPSNTKPS 127
Score = 39.7 bits (91), Expect = 0.002
Identities = 23/70 (32%), Positives = 35/70 (49%), Gaps = 3/70 (4%)
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKK 71
S++PS ++ +P+P SP S++ A++SP PT P SPS + SP
Sbjct: 2 STAPSPGTTPSPSP---PSPPTNSTTTTPPPAASSPPPTTTPSSPPPSPSTNSTSPPPSS 58
Query: 72 LRTPSNTPPA 81
PS PP+
Sbjct: 59 PLPPSLPPPS 68
Score = 34.7 bits (78), Expect = 0.066
Identities = 25/69 (36%), Positives = 35/69 (50%), Gaps = 10/69 (14%)
Query: 13 SSPSSTSSSAPAPSRSQSP-TVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKK 71
S P++++++ P P+ S P T T SSP S S + T PP + + PS PSP
Sbjct: 17 SPPTNSTTTTPPPAASSPPPTTTPSSP---PPSPSTNSTSPPPSSPLPPSLPPPSPPG-- 71
Query: 72 LRTPSNTPP 80
S TPP
Sbjct: 72 ----SLTPP 76
>At5g10430 AtAGP4
Length = 135
Score = 40.8 bits (94), Expect = 0.001
Identities = 23/78 (29%), Positives = 37/78 (46%), Gaps = 1/78 (1%)
Query: 4 AFFIFSLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWH 63
A F S + +P+ T ++ P P+ + P T A +A+P+P PP A +P+
Sbjct: 13 ALFATSALAQAPAPTPTATPPPA-TPPPVATPPPVATPPPAATPAPATPPPAATPAPATT 71
Query: 64 DPSPNSKKLRTPSNTPPA 81
PS P+ +PPA
Sbjct: 72 PPSVAPSPADVPTASPPA 89
Score = 33.1 bits (74), Expect = 0.19
Identities = 20/72 (27%), Positives = 33/72 (45%), Gaps = 2/72 (2%)
Query: 10 LSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNS 69
+++ P +T A P+ + P + +PA S +PSP P + +P SP+S
Sbjct: 40 VATPPPVATPPPAATPAPATPPPAATPAPATTPPSVAPSPADVPTASPPAPEGPTVSPSS 99
Query: 70 KKLRTPSNTPPA 81
PS+ PA
Sbjct: 100 AP--GPSDASPA 109
>At1g62760 hypothetical protein
Length = 312
Score = 40.4 bits (93), Expect = 0.001
Identities = 32/90 (35%), Positives = 48/90 (52%), Gaps = 16/90 (17%)
Query: 4 AFFIFSLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASP-----SPTVPP----- 53
A +F +SSS S SSS+P+ S S + SS+P + + +SP SP+ PP
Sbjct: 15 AILLFITTSSSSLSPSSSSPSLSPSPPSSSPSSAPPSSLSPSSPPPLSLSPSSPPPPPPS 74
Query: 54 --LMAHVSPSWHDPSPNSKKLRTPSNTPPA 81
++ +SPS PSP S +PS+ PP+
Sbjct: 75 SSPLSSLSPSL-SPSPPSS---SPSSAPPS 100
Score = 39.3 bits (90), Expect = 0.003
Identities = 31/78 (39%), Positives = 42/78 (53%), Gaps = 9/78 (11%)
Query: 9 SLSSSSPSSTSSSAPAPSRSQSP----TVTSSSPAAQAASASPSPTVPPLMAHVSPSWHD 64
SLS S PSS+ SSAP S S S +++ SSP S+SP ++ P + SPS
Sbjct: 35 SLSPSPPSSSPSSAPPSSLSPSSPPPLSLSPSSPPPPPPSSSPLSSLSPSL---SPSPPS 91
Query: 65 PSPNS--KKLRTPSNTPP 80
SP+S +PS+ PP
Sbjct: 92 SSPSSAPPSSLSPSSPPP 109
Score = 37.0 bits (84), Expect = 0.013
Identities = 26/73 (35%), Positives = 37/73 (50%), Gaps = 2/73 (2%)
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAA--SASPSPTVPPLMAHVSPSWHDPS 66
S SS P S S S+P P S ++S SP+ + S+SPS P ++ SP S
Sbjct: 54 SPSSPPPLSLSPSSPPPPPPSSSPLSSLSPSLSPSPPSSSPSSAPPSSLSPSSPPPLSLS 113
Query: 67 PNSKKLRTPSNTP 79
P+S PS++P
Sbjct: 114 PSSPPPPPPSSSP 126
Score = 33.9 bits (76), Expect = 0.11
Identities = 28/75 (37%), Positives = 40/75 (53%), Gaps = 6/75 (8%)
Query: 9 SLSSSSP-----SSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWH 63
SLS SSP SS+ S+ +PS S SP +SS +A +S SPS P ++ SP
Sbjct: 62 SLSPSSPPPPPPSSSPLSSLSPSLSPSPP-SSSPSSAPPSSLSPSSPPPLSLSPSSPPPP 120
Query: 64 DPSPNSKKLRTPSNT 78
PS + +PS++
Sbjct: 121 PPSSSPLSSLSPSSS 135
Score = 32.7 bits (73), Expect = 0.25
Identities = 25/70 (35%), Positives = 32/70 (45%), Gaps = 1/70 (1%)
Query: 9 SLSSSSPSSTSSSAPAPSRS-QSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSP 67
SLS S PSS+ SSAP S S SP S SP++ S + L S S +
Sbjct: 84 SLSPSPPSSSPSSAPPSSLSPSSPPPLSLSPSSPPPPPPSSSPLSSLSPSSSSSTYSNQT 143
Query: 68 NSKKLRTPSN 77
N ++T N
Sbjct: 144 NLDYIKTSCN 153
Score = 30.4 bits (67), Expect = 1.2
Identities = 24/68 (35%), Positives = 31/68 (45%), Gaps = 1/68 (1%)
Query: 10 LSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNS 69
LSS SPS S S P+ S S +P + S + S SPS PP + S PS +S
Sbjct: 78 LSSLSPS-LSPSPPSSSPSSAPPSSLSPSSPPPLSLSPSSPPPPPPSSSPLSSLSPSSSS 136
Query: 70 KKLRTPSN 77
+N
Sbjct: 137 STYSNQTN 144
>At3g60280 uclacyanin 3
Length = 222
Score = 40.0 bits (92), Expect = 0.002
Identities = 28/77 (36%), Positives = 44/77 (56%), Gaps = 7/77 (9%)
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASP-SPTVPPLMAHVSPSWHDP---S 66
++ SPS+ SS PS SP T S+P++ + SP SP++PP + + PS P +
Sbjct: 121 AAPSPSTPSSPPSTPSTPSSPPSTPSTPSSPPSPPSPPSPSLPP--SSLPPSASPPTNGT 178
Query: 67 PNSKKLR-TPSNTPPAL 82
P+S+ L P+ PP+L
Sbjct: 179 PDSETLTPPPAPLPPSL 195
Score = 34.3 bits (77), Expect = 0.086
Identities = 24/62 (38%), Positives = 36/62 (57%), Gaps = 11/62 (17%)
Query: 20 SSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLRTPSNTP 79
++AP+PS SP T S+P++ ++ S +P+ PP SP PSP S L PS+ P
Sbjct: 120 AAAPSPSTPSSPPSTPSTPSSPPSTPS-TPSSPP-----SP----PSPPSPSL-PPSSLP 168
Query: 80 PA 81
P+
Sbjct: 169 PS 170
>At4g37450 arabinogalactan protein AGP18
Length = 209
Score = 38.5 bits (88), Expect = 0.005
Identities = 29/77 (37%), Positives = 39/77 (49%), Gaps = 15/77 (19%)
Query: 10 LSSSSPSSTSSSAPAPSRSQSPTVTS------SSPAAQAASASPSPTVPPLMAHVSPSWH 63
+SS + S T+ SAP S ++SP VTS +P A A+S SP P A VS S
Sbjct: 25 ISSPTKSPTTPSAPTTSPTKSPAVTSPTTAPAKTPTASASSPVESPKSP---APVSESSP 81
Query: 64 DPSPNSKKLRTPSNTPP 80
P+P P ++PP
Sbjct: 82 PPTP------VPESSPP 92
Score = 30.0 bits (66), Expect = 1.6
Identities = 21/73 (28%), Positives = 31/73 (41%), Gaps = 3/73 (4%)
Query: 10 LSSSSPSST---SSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPS 66
+S SSP T SS P P+ S V+S A A + P+P P+ +P+
Sbjct: 76 VSESSPPPTPVPESSPPVPAPMVSSPVSSPPVPAPVADSPPAPVAAPVADVPAPAPSKHK 135
Query: 67 PNSKKLRTPSNTP 79
+KK + P
Sbjct: 136 KTTKKSKKHQAAP 148
Score = 30.0 bits (66), Expect = 1.6
Identities = 24/59 (40%), Positives = 29/59 (48%), Gaps = 4/59 (6%)
Query: 11 SSSSP-SSTSSSAPAPSRSQSPT-VTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSP 67
S+SSP S S AP S PT V SSP A S + PP+ A V+ S P+P
Sbjct: 62 SASSPVESPKSPAPVSESSPPPTPVPESSPPVPAPMVSSPVSSPPVPAPVADS--PPAP 118
Score = 29.6 bits (65), Expect = 2.1
Identities = 20/53 (37%), Positives = 24/53 (44%), Gaps = 4/53 (7%)
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPS 61
S S +P S SS P P SP V PA +S SP VP +A P+
Sbjct: 69 SPKSPAPVSESSPPPTPVPESSPPV----PAPMVSSPVSSPPVPAPVADSPPA 117
Score = 29.3 bits (64), Expect = 2.8
Identities = 18/50 (36%), Positives = 24/50 (48%), Gaps = 4/50 (8%)
Query: 20 SSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNS 69
+ PAP+ S+ T S QAA P+P P L+ +P P PNS
Sbjct: 124 ADVPAPAPSKHKKTTKKSKKHQAA---PAP-APELLGPPAPPTESPGPNS 169
>At4g18670 extensin-like protein
Length = 839
Score = 38.5 bits (88), Expect = 0.005
Identities = 25/70 (35%), Positives = 34/70 (47%), Gaps = 2/70 (2%)
Query: 13 SSPSSTSSSAPAPSRSQSPTVTS--SSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSK 70
S PSS ++ P S SPT S SP + + S SP TVP + + PSP+S
Sbjct: 493 SPPSSPTTPTPGGSPPSSPTTPSPGGSPPSPSISPSPPITVPSPPSTPTSPGSPPSPSSP 552
Query: 71 KLRTPSNTPP 80
+P +PP
Sbjct: 553 TPSSPIPSPP 562
Score = 37.0 bits (84), Expect = 0.013
Identities = 25/74 (33%), Positives = 36/74 (47%), Gaps = 3/74 (4%)
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAAS--ASPSPTVPPLMAHVSPSWHDPSPN 68
SS + S S P+PS S SP +T SP + S + PSP+ P + + PS PS
Sbjct: 509 SSPTTPSPGGSPPSPSISPSPPITVPSPPSTPTSPGSPPSPSSPTPSSPI-PSPPTPSTP 567
Query: 69 SKKLRTPSNTPPAL 82
+ N+PP +
Sbjct: 568 PTPISPGQNSPPII 581
Score = 36.6 bits (83), Expect = 0.017
Identities = 27/77 (35%), Positives = 37/77 (47%), Gaps = 11/77 (14%)
Query: 9 SLSSSSPSSTSSSAPAPSR--------SQSPTVTSSSP-AAQAASASPSPTVPPLMAHVS 59
S SS +PSS S P PS SP + S P + +SPSP +PP++ S
Sbjct: 548 SPSSPTPSSPIPSPPTPSTPPTPISPGQNSPPIIPSPPFTGPSPPSSPSPPLPPVIP--S 605
Query: 60 PSWHDPSPNSKKLRTPS 76
P P+P+S TP+
Sbjct: 606 PPIVGPTPSSPPPSTPT 622
Score = 36.6 bits (83), Expect = 0.017
Identities = 24/72 (33%), Positives = 36/72 (49%), Gaps = 4/72 (5%)
Query: 14 SPSSTSS---SAPAPSRSQSPTVTSSSPAAQAAS-ASPSPTVPPLMAHVSPSWHDPSPNS 69
SP +T S S P+PS SP T+ SP + S +P+P P + +P+ P+S
Sbjct: 438 SPPTTPSPGGSPPSPSIVPSPPSTTPSPGSPPTSPTTPTPGGSPPSSPTTPTPGGSPPSS 497
Query: 70 KKLRTPSNTPPA 81
TP +PP+
Sbjct: 498 PTTPTPGGSPPS 509
Score = 35.0 bits (79), Expect = 0.051
Identities = 24/73 (32%), Positives = 34/73 (45%), Gaps = 3/73 (4%)
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
S+S S P + S PS SP S P+ S +PSP PP + +P+ P+
Sbjct: 427 SISPSPPITVPSPPTTPSPGGSPPSPSIVPS--PPSTTPSPGSPP-TSPTTPTPGGSPPS 483
Query: 69 SKKLRTPSNTPPA 81
S TP +PP+
Sbjct: 484 SPTTPTPGGSPPS 496
Score = 35.0 bits (79), Expect = 0.051
Identities = 20/69 (28%), Positives = 31/69 (43%)
Query: 13 SSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKL 72
S PSS ++ P S SPT + + ++ +PSP P +SPS P+
Sbjct: 480 SPPSSPTTPTPGGSPPSSPTTPTPGGSPPSSPTTPSPGGSPPSPSISPSPPITVPSPPST 539
Query: 73 RTPSNTPPA 81
T +PP+
Sbjct: 540 PTSPGSPPS 548
Score = 34.7 bits (78), Expect = 0.066
Identities = 27/87 (31%), Positives = 37/87 (42%), Gaps = 21/87 (24%)
Query: 14 SPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSP-----TVPPLMAHVSPSWHDPSPN 68
SP ST +S +P SPT +S P+ S P+P PP++ SP + PSP
Sbjct: 535 SPPSTPTSPGSPPSPSSPTPSSPIPSPPTPSTPPTPISPGQNSPPIIP--SPPFTGPSPP 592
Query: 69 SKKL--------------RTPSNTPPA 81
S TPS+ PP+
Sbjct: 593 SSPSPPLPPVIPSPPIVGPTPSSPPPS 619
Score = 34.3 bits (77), Expect = 0.086
Identities = 20/73 (27%), Positives = 35/73 (47%), Gaps = 3/73 (4%)
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
S+ S PS+T S P+ +PT S P++ +P+P P + +P+ P+
Sbjct: 453 SIVPSPPSTTPSPGSPPTSPTTPTPGGSPPSSPT---TPTPGGSPPSSPTTPTPGGSPPS 509
Query: 69 SKKLRTPSNTPPA 81
S +P +PP+
Sbjct: 510 SPTTPSPGGSPPS 522
Score = 31.2 bits (69), Expect = 0.73
Identities = 27/83 (32%), Positives = 36/83 (42%), Gaps = 11/83 (13%)
Query: 9 SLSSSSPSSTSSSAPA-PSRSQSPTVTSSSPAAQAASASP-----SPTVPPLMAHVSPSW 62
S ++ +P + S+P PS SP S SP+ SP SP PP + +PS
Sbjct: 497 SPTTPTPGGSPPSSPTTPSPGGSPPSPSISPSPPITVPSPPSTPTSPGSPPSPSSPTPSS 556
Query: 63 HDPSPNSKKLRTPSNTPPALK*G 85
PSP TPS P + G
Sbjct: 557 PIPSP-----PTPSTPPTPISPG 574
Score = 30.0 bits (66), Expect = 1.6
Identities = 23/82 (28%), Positives = 31/82 (37%), Gaps = 12/82 (14%)
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPA-----------AQAASASPSPTVPPLMAHVS 59
S+ P S PS SP S SP+ + PSP++ P +
Sbjct: 403 STRPPVVVPSPPTTPSPGGSPPSPSISPSPPITVPSPPTTPSPGGSPPSPSIVPSPPSTT 462
Query: 60 PSWHDPSPNSKKLRTPSNTPPA 81
PS P P S TP +PP+
Sbjct: 463 PSPGSP-PTSPTTPTPGGSPPS 483
Score = 29.6 bits (65), Expect = 2.1
Identities = 27/80 (33%), Positives = 34/80 (41%), Gaps = 19/80 (23%)
Query: 13 SSPSSTSSSAPAPSRSQSPTVTS------SSPAAQAASASP--SPTVPPLMAHVSPSWHD 64
S P+S ++ P S SPT + SSP SP SPT P SP
Sbjct: 467 SPPTSPTTPTPGGSPPSSPTTPTPGGSPPSSPTTPTPGGSPPSSPTTP------SPGGSP 520
Query: 65 PSPN---SKKLRTPSNTPPA 81
PSP+ S + PS PP+
Sbjct: 521 PSPSISPSPPITVPS--PPS 538
Score = 28.5 bits (62), Expect = 4.7
Identities = 26/81 (32%), Positives = 31/81 (38%), Gaps = 10/81 (12%)
Query: 8 FSLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTV------PPLMAHVSPS 61
+SL +P+ S P P S P S P PTV PP AH SP
Sbjct: 721 YSLPPPTPTYHYISPPPPPTPIHSPPPQSHPPCIEYSPPPPPTVHYNPPPPPSPAHYSP- 779
Query: 62 WHDPSPNSKKLRTPSNTPPAL 82
PSP +P PPA+
Sbjct: 780 --PPSPPVYYYNSPP-PPPAV 797
>At1g68725 putative arabinogalactan protein AGP19
Length = 248
Score = 38.5 bits (88), Expect = 0.005
Identities = 23/75 (30%), Positives = 40/75 (52%), Gaps = 10/75 (13%)
Query: 12 SSSPSSTSSSAPAPSRSQSPTV-----TSSSPAAQAASASPSP-TVPPLMAHVSPSWHDP 65
++SP +++++AP P+ + PT T+++P AA SP T PP + SP P
Sbjct: 28 AASPVTSTTTAPPPTTAAPPTTAAPPPTTTTPPVSAAQPPASPVTPPPAVTPTSP----P 83
Query: 66 SPNSKKLRTPSNTPP 80
+P + +P+ PP
Sbjct: 84 APKVAPVISPATPPP 98
Score = 34.7 bits (78), Expect = 0.066
Identities = 25/73 (34%), Positives = 34/73 (46%), Gaps = 11/73 (15%)
Query: 15 PSSTSSSAPAP------SRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
P+ T +S PAP S + P SP A A + SP P PP +P+ P+P
Sbjct: 75 PAVTPTSPPAPKVAPVISPATPPPQPPQSPPASAPTVSPPPVSPP----PAPTSPPPTPA 130
Query: 69 SKKLRTPSNTPPA 81
S P++ PPA
Sbjct: 131 SPP-PAPASPPPA 142
Score = 32.7 bits (73), Expect = 0.25
Identities = 21/81 (25%), Positives = 36/81 (43%), Gaps = 10/81 (12%)
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSK 70
++++P +++ PA + P VT +SP A + SP PP SP P+ +
Sbjct: 55 TTTTPPVSAAQPPASPVTPPPAVTPTSPPAPKVAPVISPATPPPQPPQSPPASAPTVSPP 114
Query: 71 KLR----------TPSNTPPA 81
+ TP++ PPA
Sbjct: 115 PVSPPPAPTSPPPTPASPPPA 135
Score = 32.7 bits (73), Expect = 0.25
Identities = 19/58 (32%), Positives = 25/58 (42%)
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNS 69
+S+P+ + P SP T +SP AS P+P PP P PSP S
Sbjct: 106 ASAPTVSPPPVSPPPAPTSPPPTPASPPPAPASPPPAPASPPPAPVSPPPVQAPSPIS 163
Score = 32.0 bits (71), Expect = 0.43
Identities = 19/55 (34%), Positives = 26/55 (46%), Gaps = 3/55 (5%)
Query: 13 SSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSP 67
S P + +S P P+ SP +SP AS P+P PP + SP P+P
Sbjct: 117 SPPPAPTSPPPTPA---SPPPAPASPPPAPASPPPAPVSPPPVQAPSPISLPPAP 168
Score = 31.2 bits (69), Expect = 0.73
Identities = 22/84 (26%), Positives = 42/84 (49%), Gaps = 11/84 (13%)
Query: 7 IFSLSSSSPSSTSS----SAPAPSRSQSPTVTSSSPAAQAASASP-----SPTVPPLMAH 57
+ S +++ P +T++ +AP P+ + +P V+++ P A + P SP P +
Sbjct: 32 VTSTTTAPPPTTAAPPTTAAPPPT-TTTPPVSAAQPPASPVTPPPAVTPTSPPAPKVAPV 90
Query: 58 VSPSWHDPS-PNSKKLRTPSNTPP 80
+SP+ P P S P+ +PP
Sbjct: 91 ISPATPPPQPPQSPPASAPTVSPP 114
Score = 30.0 bits (66), Expect = 1.6
Identities = 24/70 (34%), Positives = 33/70 (46%), Gaps = 6/70 (8%)
Query: 13 SSPSSTSSSAPAP-SRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKK 71
S P+S + +P P S +PT +P AS P+P PP A SP SP +
Sbjct: 103 SPPASAPTVSPPPVSPPPAPTSPPPTP----ASPPPAPASPP-PAPASPPPAPVSPPPVQ 157
Query: 72 LRTPSNTPPA 81
+P + PPA
Sbjct: 158 APSPISLPPA 167
>At5g65390 arabinogalactan protein AGP7
Length = 130
Score = 38.1 bits (87), Expect = 0.006
Identities = 25/87 (28%), Positives = 39/87 (44%), Gaps = 7/87 (8%)
Query: 3 DAFFIFSLSSSS-------PSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLM 55
+AFFI +L ++S PS T++ P P + P T + + SP+PT P
Sbjct: 7 EAFFIVALFTTSCLAQAPAPSPTTTVTPPPVATPPPAATPAPTTTPPPAVSPAPTSSPPS 66
Query: 56 AHVSPSWHDPSPNSKKLRTPSNTPPAL 82
+ SPS P+ + P +P L
Sbjct: 67 SAPSPSSDAPTASPPAPEGPGVSPGEL 93
Score = 27.7 bits (60), Expect = 8.1
Identities = 18/42 (42%), Positives = 22/42 (51%), Gaps = 6/42 (14%)
Query: 14 SPSSTS---SSAPAPSRSQSPTVTSSSPAAQAASASPSPTVP 52
SP+ TS SSAP+PS S T+S PA + SP P
Sbjct: 57 SPAPTSSPPSSAPSPS---SDAPTASPPAPEGPGVSPGELAP 95
>At5g53870 predicted GPI-anchored protein
Length = 370
Score = 38.1 bits (87), Expect = 0.006
Identities = 29/76 (38%), Positives = 37/76 (48%), Gaps = 11/76 (14%)
Query: 12 SSSPSSTSSSAPAPSRSQSPTVT--------SSSPAAQAASASPSPTVPPLMAHVSPSWH 63
S SP S + A APS+SQ P + SSSP + + SPS A SPS
Sbjct: 153 SHSPVSPVAPASAPSKSQPPRSSVSPAQPPKSSSPISHTPALSPSHATSHSPATPSPSPK 212
Query: 64 DPSPNSKKLRTPSNTP 79
PSP S +PS++P
Sbjct: 213 SPSPVS---HSPSHSP 225
Score = 37.4 bits (85), Expect = 0.010
Identities = 26/75 (34%), Positives = 35/75 (46%), Gaps = 8/75 (10%)
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASP------SPTVPPLMAHVSPSWHDP 65
S SP+ T S +PA + S SP S A A S SP SP P + +P P
Sbjct: 222 SHSPAHTPSHSPAHTPSHSPAHAPSHSPAHAPSHSPAHAPSHSPAHSPSHSPATPK--SP 279
Query: 66 SPNSKKLRTPSNTPP 80
SP+S ++P+ P
Sbjct: 280 SPSSSPAQSPATPSP 294
Score = 37.0 bits (84), Expect = 0.013
Identities = 28/73 (38%), Positives = 37/73 (50%), Gaps = 7/73 (9%)
Query: 12 SSSPSSTSSSAPAPSRSQSP-TVTSSSPAAQAAS--ASPSPTVPPLMAHVSPSWHDPSPN 68
S SP+ S +PA S S SP T S SP++ A A+PSP P + VS PSP+
Sbjct: 254 SHSPAHAPSHSPAHSPSHSPATPKSPSPSSSPAQSPATPSPMTPQSPSPVS----SPSPD 309
Query: 69 SKKLRTPSNTPPA 81
+ +TP A
Sbjct: 310 QSAAPSDQSTPLA 322
Score = 36.2 bits (82), Expect = 0.023
Identities = 30/81 (37%), Positives = 39/81 (48%), Gaps = 18/81 (22%)
Query: 11 SSSSPSSTSSSAPAPSRSQSPT----------VTSSSPAAQAASASPSPTVPPLMAHVSP 60
S S P +S S P +S SP TS SP A+ SPSP P ++H SP
Sbjct: 167 SKSQPPRSSVSPAQPPKSSSPISHTPALSPSHATSHSP----ATPSPSPKSPSPVSH-SP 221
Query: 61 SWHDP--SPNSKKLRTPSNTP 79
S H P +P+ TPS++P
Sbjct: 222 S-HSPAHTPSHSPAHTPSHSP 241
Score = 35.4 bits (80), Expect = 0.039
Identities = 30/78 (38%), Positives = 38/78 (48%), Gaps = 7/78 (8%)
Query: 9 SLSSSSPSSTSSS---APAPSRSQSPTVTSSSPAAQAASASP--SPTVPPLMAHVSPSWH 63
S S SPS S S +PA + S SP T S A A S SP +P+ P A H
Sbjct: 208 SPSPKSPSPVSHSPSHSPAHTPSHSPAHTPSHSPAHAPSHSPAHAPSHSPAHAPSHSPAH 267
Query: 64 DP--SPNSKKLRTPSNTP 79
P SP + K +PS++P
Sbjct: 268 SPSHSPATPKSPSPSSSP 285
Score = 34.7 bits (78), Expect = 0.066
Identities = 24/71 (33%), Positives = 36/71 (49%), Gaps = 6/71 (8%)
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
S S SS + S + P+P QSP+ SS Q SA+PS PL +PS + +P
Sbjct: 278 SPSPSSSPAQSPATPSPMTPQSPSPVSSPSPDQ--SAAPSDQSTPL----APSPSETTPT 331
Query: 69 SKKLRTPSNTP 79
+ + P+ +P
Sbjct: 332 ADNITAPAPSP 342
Score = 33.9 bits (76), Expect = 0.11
Identities = 25/68 (36%), Positives = 37/68 (53%), Gaps = 7/68 (10%)
Query: 14 SPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDP--SPNSKK 71
SPS +S +PA + SP+ S SP + + S SP+ T AH +PS H P +P+
Sbjct: 195 SPSHATSHSPA---TPSPSPKSPSPVSHSPSHSPAHTPSHSPAH-TPS-HSPAHAPSHSP 249
Query: 72 LRTPSNTP 79
PS++P
Sbjct: 250 AHAPSHSP 257
Score = 33.1 bits (74), Expect = 0.19
Identities = 28/76 (36%), Positives = 41/76 (53%), Gaps = 8/76 (10%)
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPA---AQAASASPSPTVPPLMAHVSPSWHDP 65
+LS S +S S + P+PS +SP+ S SP+ A S SP+ T AH +PS H P
Sbjct: 193 ALSPSHATSHSPATPSPS-PKSPSPVSHSPSHSPAHTPSHSPAHTPSHSPAH-APS-HSP 249
Query: 66 --SPNSKKLRTPSNTP 79
+P+ PS++P
Sbjct: 250 AHAPSHSPAHAPSHSP 265
Score = 32.7 bits (73), Expect = 0.25
Identities = 27/77 (35%), Positives = 37/77 (47%), Gaps = 4/77 (5%)
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDP--S 66
S ++ SPS S S + S S SP T S A S SP+ AH +PS H P +
Sbjct: 203 SPATPSPSPKSPSPVSHSPSHSPAHTPSHSPAHTPSHSPAHAPSHSPAH-APS-HSPAHA 260
Query: 67 PNSKKLRTPSNTPPALK 83
P+ +PS++P K
Sbjct: 261 PSHSPAHSPSHSPATPK 277
Score = 30.8 bits (68), Expect = 0.95
Identities = 24/71 (33%), Positives = 36/71 (49%), Gaps = 3/71 (4%)
Query: 10 LSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASAS-PSPTVPPLMAHVSPSWHDPSPN 68
L+ + S + +P PS S + S SP + A AS PS + PP + VSP+ P +
Sbjct: 128 LAVRNQPSAPAHSPVPSVSPTQPPKSHSPVSPVAPASAPSKSQPP-RSSVSPA-QPPKSS 185
Query: 69 SKKLRTPSNTP 79
S TP+ +P
Sbjct: 186 SPISHTPALSP 196
Score = 30.0 bits (66), Expect = 1.6
Identities = 18/60 (30%), Positives = 32/60 (53%), Gaps = 2/60 (3%)
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPT-VPPLMAHVSPSWHDPSPNS 69
S ++PS + +P+P S SP S++P+ Q+ +PSP+ P +++ P NS
Sbjct: 288 SPATPSPMTPQSPSPVSSPSPD-QSAAPSDQSTPLAPSPSETTPTADNITAPAPSPRTNS 346
>At5g38560 putative protein
Length = 681
Score = 38.1 bits (87), Expect = 0.006
Identities = 28/72 (38%), Positives = 33/72 (44%), Gaps = 14/72 (19%)
Query: 14 SPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTV----PPLMAHVSPSWHDPSPNS 69
SP SS+P P SP +SS P + SP PTV PP + SP PS
Sbjct: 47 SPPPVVSSSPPPPVVSSPPPSSSPPPSPPVITSPPPTVASSPPPPVVIASP---PPS--- 100
Query: 70 KKLRTPSNTPPA 81
TP+ TPPA
Sbjct: 101 ----TPATTPPA 108
Score = 36.6 bits (83), Expect = 0.017
Identities = 22/73 (30%), Positives = 34/73 (46%), Gaps = 4/73 (5%)
Query: 10 LSSSSPSSTSSSAPAPSRSQS----PTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDP 65
++S PS+ +++ PAP ++ S P + S PA + P P+ P SP P
Sbjct: 94 IASPPPSTPATTPPAPPQTVSPPPPPDASPSPPAPTTTNPPPKPSPSPPGETPSPPGETP 153
Query: 66 SPNSKKLRTPSNT 78
SP TP+ T
Sbjct: 154 SPPKPSPSTPTPT 166
Score = 36.2 bits (82), Expect = 0.023
Identities = 26/74 (35%), Positives = 31/74 (41%), Gaps = 6/74 (8%)
Query: 13 SSPSSTSSSAPAPSRSQSPTVT-SSSPAAQAASASPSPTVPPLMAHVSPSWHDPSP---- 67
SSP +SS P+P SP T +SSP ASP P+ P P P P
Sbjct: 62 SSPPPSSSPPPSPPVITSPPPTVASSPPPPVVIASPPPSTPATTPPAPPQTVSPPPPPDA 121
Query: 68 -NSKKLRTPSNTPP 80
S T +N PP
Sbjct: 122 SPSPPAPTTTNPPP 135
Score = 34.7 bits (78), Expect = 0.066
Identities = 21/67 (31%), Positives = 31/67 (45%), Gaps = 6/67 (8%)
Query: 14 SPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLR 73
SP S++SS AP Q+ T S+P SP + PP+++ P P
Sbjct: 11 SPPSSNSSTTAPPPLQTQPTTPSAPPPVTPPPSPPQSPPPVVS------SSPPPPVVSSP 64
Query: 74 TPSNTPP 80
PS++PP
Sbjct: 65 PPSSSPP 71
Score = 33.9 bits (76), Expect = 0.11
Identities = 23/79 (29%), Positives = 31/79 (39%), Gaps = 12/79 (15%)
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAAS--------ASPSPTVPPLMAHVSP 60
++S P S S PAP+ + P S SP + S PSP+ P SP
Sbjct: 112 TVSPPPPPDASPSPPAPTTTNPPPKPSPSPPGETPSPPGETPSPPKPSPSTPTPTTTTSP 171
Query: 61 SWHDPSPNSKKLRTPSNTP 79
P P + PS+ P
Sbjct: 172 ----PPPPATSASPPSSNP 186
Score = 33.1 bits (74), Expect = 0.19
Identities = 21/47 (44%), Positives = 26/47 (54%), Gaps = 4/47 (8%)
Query: 14 SPSSTSSSAPAPSRSQ-SPTVTSSSPAAQAASASP---SPTVPPLMA 56
SP + S P PS S +PT T+S P A SASP +PT P +A
Sbjct: 147 SPPGETPSPPKPSPSTPTPTTTTSPPPPPATSASPPSSNPTDPSTLA 193
Score = 32.7 bits (73), Expect = 0.25
Identities = 21/68 (30%), Positives = 33/68 (47%), Gaps = 3/68 (4%)
Query: 14 SPSSTSSSAPAPSRSQSPTV-TSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKL 72
+P T S P P S SP T+++P + + + P T P +PS PSP++
Sbjct: 108 APPQTVSPPPPPDASPSPPAPTTTNPPPKPSPSPPGETPSP--PGETPSPPKPSPSTPTP 165
Query: 73 RTPSNTPP 80
T ++ PP
Sbjct: 166 TTTTSPPP 173
Score = 32.7 bits (73), Expect = 0.25
Identities = 20/71 (28%), Positives = 38/71 (53%), Gaps = 3/71 (4%)
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSP--SWHDPSPN 68
S S+P+ T++++P P + S + SS+P ++ +P PT P++ P P+ N
Sbjct: 159 SPSTPTPTTTTSPPPPPATSASPPSSNP-TDPSTLAPPPTPLPVVPREKPIAKPTGPASN 217
Query: 69 SKKLRTPSNTP 79
+ PS++P
Sbjct: 218 NGNNTLPSSSP 228
Score = 30.8 bits (68), Expect = 0.95
Identities = 23/83 (27%), Positives = 33/83 (39%), Gaps = 15/83 (18%)
Query: 12 SSSPSSTSSSAPAP--SRSQSPTVTSSSPAAQAASASPSP-------------TVPPLMA 56
+S P + +SS P P S P+ +++P A + SP P T PP
Sbjct: 78 TSPPPTVASSPPPPVVIASPPPSTPATTPPAPPQTVSPPPPPDASPSPPAPTTTNPPPKP 137
Query: 57 HVSPSWHDPSPNSKKLRTPSNTP 79
SP PSP + P +P
Sbjct: 138 SPSPPGETPSPPGETPSPPKPSP 160
>At4g34440 serine/threonine protein kinase - like
Length = 670
Score = 38.1 bits (87), Expect = 0.006
Identities = 29/86 (33%), Positives = 41/86 (46%), Gaps = 16/86 (18%)
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSP--------------TVPPLMA 56
S+ + S SS P P S P+ S+ P +AS SP P + PP++A
Sbjct: 21 SNGTSPSNESSPPTPPSSPPPSSISAPPPDISASFSPPPAPPTQETSPPTSPSSSPPVVA 80
Query: 57 HVSPSW-HDPSPNSKKLRTPSNTPPA 81
+ SP +PSP + + TP TPPA
Sbjct: 81 NPSPQTPENPSPPAPEGSTPV-TPPA 105
Score = 38.1 bits (87), Expect = 0.006
Identities = 22/72 (30%), Positives = 36/72 (49%), Gaps = 2/72 (2%)
Query: 13 SSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPP--LMAHVSPSWHDPSPNSK 70
SSP+ +S+ PS SP+ SS P ++ S + PP + A SP P+ +
Sbjct: 8 SSPAPETSNGTPPSNGTSPSNESSPPTPPSSPPPSSISAPPPDISASFSPPPAPPTQETS 67
Query: 71 KLRTPSNTPPAL 82
+PS++PP +
Sbjct: 68 PPTSPSSSPPVV 79
Score = 36.2 bits (82), Expect = 0.023
Identities = 27/79 (34%), Positives = 36/79 (45%), Gaps = 11/79 (13%)
Query: 13 SSPSSTSSSAPAPSRSQS-------PTVTSSSPAAQAAS----ASPSPTVPPLMAHVSPS 61
SSP +S SAP P S S PT +S P + ++S A+PSP P + +P
Sbjct: 37 SSPPPSSISAPPPDISASFSPPPAPPTQETSPPTSPSSSPPVVANPSPQTPENPSPPAPE 96
Query: 62 WHDPSPNSKKLRTPSNTPP 80
P +TPSN P
Sbjct: 97 GSTPVTPPAPPQTPSNQSP 115
Score = 32.0 bits (71), Expect = 0.43
Identities = 24/82 (29%), Positives = 39/82 (47%), Gaps = 11/82 (13%)
Query: 9 SLSSSSPSSTSSSAPAPS----RSQSPTVTSSSPAAQAASA-----SPSPTVPPLMAHVS 59
S+S+ P ++S +P P+ + PT SSSP A + +PSP P V+
Sbjct: 43 SISAPPPDISASFSPPPAPPTQETSPPTSPSSSPPVVANPSPQTPENPSPPAPEGSTPVT 102
Query: 60 PSWHDPSPNSKKLRTPSNTPPA 81
P +P+++ P TPP+
Sbjct: 103 PPAPPQTPSNQSPERP--TPPS 122
Score = 30.4 bits (67), Expect = 1.2
Identities = 22/68 (32%), Positives = 31/68 (45%), Gaps = 2/68 (2%)
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPP-LMAHVSPSW-HDPSPNS 69
+S P+S SSS P + T + SP A S +P PP ++ SP PSP +
Sbjct: 66 TSPPTSPSSSPPVVANPSPQTPENPSPPAPEGSTPVTPPAPPQTPSNQSPERPTPPSPGA 125
Query: 70 KKLRTPSN 77
R +N
Sbjct: 126 NDDRNRTN 133
>At3g19430 putative late embryogenesis abundant protein
Length = 550
Score = 38.1 bits (87), Expect = 0.006
Identities = 24/68 (35%), Positives = 34/68 (49%), Gaps = 6/68 (8%)
Query: 13 SSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKL 72
S PS T +P P + +P+V S +P +P+P+VP VSP P+P+
Sbjct: 89 SVPSPTPPVSPPPP-TPTPSVPSPTPPVSPPPPTPTPSVPSPTPPVSPPPPTPTPS---- 143
Query: 73 RTPSNTPP 80
PS TPP
Sbjct: 144 -VPSPTPP 150
Score = 38.1 bits (87), Expect = 0.006
Identities = 24/68 (35%), Positives = 34/68 (49%), Gaps = 6/68 (8%)
Query: 13 SSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKL 72
S PS T +P P + +P+V S +P +P+P+VP VSP P+P+
Sbjct: 107 SVPSPTPPVSPPPP-TPTPSVPSPTPPVSPPPPTPTPSVPSPTPPVSPPPPTPTPS---- 161
Query: 73 RTPSNTPP 80
PS TPP
Sbjct: 162 -VPSPTPP 168
Score = 33.9 bits (76), Expect = 0.11
Identities = 22/62 (35%), Positives = 30/62 (47%), Gaps = 8/62 (12%)
Query: 22 APAPSRSQ---SPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLRTPSNT 78
AP P S +P+V S +P +P+P+VP VSP P+P+ PS T
Sbjct: 76 APVPPVSPPPPTPSVPSPTPPVSPPPPTPTPSVPSPTPPVSPPPPTPTPS-----VPSPT 130
Query: 79 PP 80
PP
Sbjct: 131 PP 132
Score = 33.5 bits (75), Expect = 0.15
Identities = 23/71 (32%), Positives = 30/71 (41%), Gaps = 1/71 (1%)
Query: 11 SSSSPSSTSSSAPAPS-RSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNS 69
S + P S P PS S +P V+ P + SP+P V P +PS P+P
Sbjct: 110 SPTPPVSPPPPTPTPSVPSPTPPVSPPPPTPTPSVPSPTPPVSPPPPTPTPSVPSPTPPV 169
Query: 70 KKLRTPSNTPP 80
PS PP
Sbjct: 170 PTDPMPSPPPP 180
Score = 33.5 bits (75), Expect = 0.15
Identities = 22/71 (30%), Positives = 27/71 (37%), Gaps = 1/71 (1%)
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPS-PNS 69
S P S P PS P VT + P S PP + SP P+ P
Sbjct: 176 SPPPPVSPPPPTPTPSVPSPPDVTPTPPTPSVPSPPDVTPTPPTPSVPSPPDVTPTPPTP 235
Query: 70 KKLRTPSNTPP 80
+ TPS +PP
Sbjct: 236 PSVPTPSGSPP 246
Score = 30.8 bits (68), Expect = 0.95
Identities = 25/71 (35%), Positives = 31/71 (43%), Gaps = 4/71 (5%)
Query: 11 SSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSK 70
S + P S P PS SPT S P + PSPT PP+ PS P P S
Sbjct: 128 SPTPPVSPPPPTPTPS-VPSPTPPVSPPPPTPTPSVPSPT-PPVPTDPMPS--PPPPVSP 183
Query: 71 KLRTPSNTPPA 81
TP+ + P+
Sbjct: 184 PPPTPTPSVPS 194
Score = 29.3 bits (64), Expect = 2.8
Identities = 20/73 (27%), Positives = 33/73 (44%), Gaps = 7/73 (9%)
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAAS----ASPSPTVPPLMAHVSPSWHDPSP 67
S P + + S P+P+ SP + +P+ + + P P+ PP VSP P+P
Sbjct: 134 SPPPPTPTPSVPSPTPPVSPPPPTPTPSVPSPTPPVPTDPMPSPPP---PVSPPPPTPTP 190
Query: 68 NSKKLRTPSNTPP 80
+ + TPP
Sbjct: 191 SVPSPPDVTPTPP 203
Score = 28.1 bits (61), Expect = 6.2
Identities = 16/47 (34%), Positives = 22/47 (46%), Gaps = 5/47 (10%)
Query: 34 TSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLRTPSNTPP 80
T +P + P+P+VP VSP P+P+ PS TPP
Sbjct: 73 TPPAPVPPVSPPPPTPSVPSPTPPVSPPPPTPTPS-----VPSPTPP 114
>At2g28440 En/Spm-like transposon protein
Length = 268
Score = 38.1 bits (87), Expect = 0.006
Identities = 23/59 (38%), Positives = 34/59 (56%), Gaps = 3/59 (5%)
Query: 12 SSSPSSTSSSAPAPS-RSQSPTVTSSSPAAQAASASPSPTVPPLM--AHVSPSWHDPSP 67
SSSP + S P+ S + SP +SSP ++ + SPSP PP + SPS+ +P+P
Sbjct: 108 SSSPEADSPLPPSSSPEANSPQSPASSPKPESLADSPSPPPPPPQPESPSSPSYPEPAP 166
Score = 35.0 bits (79), Expect = 0.051
Identities = 26/71 (36%), Positives = 33/71 (45%), Gaps = 2/71 (2%)
Query: 12 SSSPSSTSSSAPAPS-RSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSK 70
SSSP S P+ S + SP SSSP A + + S P +A SPS P P +
Sbjct: 96 SSSPEVDSPQPPSSSPEADSPLPPSSSPEANSPQSPASSPKPESLAD-SPSPPPPPPQPE 154
Query: 71 KLRTPSNTPPA 81
+PS PA
Sbjct: 155 SPSSPSYPEPA 165
Score = 34.7 bits (78), Expect = 0.066
Identities = 28/73 (38%), Positives = 33/73 (44%), Gaps = 3/73 (4%)
Query: 12 SSSPSSTSSSAPAPSRSQ-SPTVTSSSPAAQAASA-SPSPTVP-PLMAHVSPSWHDPSPN 68
SSSP S +P+ S + SP SSSP + A S SP V PL SP P P
Sbjct: 48 SSSPEEDSPLSPSSSPEEDSPLPPSSSPEEDSPLAPSSSPEVDSPLAPSSSPEVDSPQPP 107
Query: 69 SKKLRTPSNTPPA 81
S S PP+
Sbjct: 108 SSSPEADSPLPPS 120
Score = 30.4 bits (67), Expect = 1.2
Identities = 20/72 (27%), Positives = 34/72 (46%), Gaps = 1/72 (1%)
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMA-HVSPSWHDPSP 67
S ++SP S +SS S + SP+ P ++ S+ P P+ A S DP P
Sbjct: 122 SPEANSPQSPASSPKPESLADSPSPPPPPPQPESPSSPSYPEPAPVPAPSDDDSDDDPEP 181
Query: 68 NSKKLRTPSNTP 79
++ +P+ +P
Sbjct: 182 ETEYFPSPAPSP 193
Score = 27.7 bits (60), Expect = 8.1
Identities = 30/97 (30%), Positives = 41/97 (41%), Gaps = 22/97 (22%)
Query: 7 IFSLSSSSPSSTSSSAPAPSR---SQSPTVTSSSP---AAQAASASP---SPTVP----- 52
IF + + S S +A +P R + SP SSSP + + S+SP SP P
Sbjct: 17 IFDFAGAQEESPSPAAVSPGREPSTDSPLSPSSSPEEDSPLSPSSSPEEDSPLPPSSSPE 76
Query: 53 ---PLMAHVSPSWHDP-----SPNSKKLRTPSNTPPA 81
PL SP P SP + PS++P A
Sbjct: 77 EDSPLAPSSSPEVDSPLAPSSSPEVDSPQPPSSSPEA 113
>At1g49490 hypothetical protein
Length = 847
Score = 37.7 bits (86), Expect = 0.008
Identities = 30/97 (30%), Positives = 45/97 (45%), Gaps = 35/97 (36%)
Query: 10 LSSSSPSSTSSSAPAPSRSQS-----------------PTVTS--------SSPAAQAAS 44
+ SS+PSS + P PS S+S PT +S SS + Q+ S
Sbjct: 690 VQSSTPSSEPTQVPTPSSSESYQAPNLSPVQAPTPVQAPTTSSETSQVPTPSSESNQSPS 749
Query: 45 ASPSPTVPPLMAHVSPSWHDPSPNSKKLR--TPSNTP 79
+P+P + P+ H P+PNSK ++ TPS+ P
Sbjct: 750 QAPTPILEPV--------HAPTPNSKPVQSPTPSSEP 778
Score = 33.5 bits (75), Expect = 0.15
Identities = 26/75 (34%), Positives = 28/75 (36%), Gaps = 5/75 (6%)
Query: 10 LSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAAS-----ASPSPTVPPLMAHVSPSWHD 64
+ S SP S S P P S P V SS P S ASP P PP H P
Sbjct: 540 MPSPSPPSPIYSPPPPVHSPPPPVYSSPPPPHVYSPPPPVASPPPPSPPPPVHSPPPPPV 599
Query: 65 PSPNSKKLRTPSNTP 79
SP P +P
Sbjct: 600 FSPPPPVFSPPPPSP 614
Score = 32.7 bits (73), Expect = 0.25
Identities = 19/66 (28%), Positives = 23/66 (34%)
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKK 71
S + SS +P P R+ P + P S S P P SP P S K
Sbjct: 385 SGGSNGGSSPSPNPPRTSEPKPSKPEPVMPKPSDSSKPETPKTPEQPSPKPQPPKHESPK 444
Query: 72 LRTPSN 77
P N
Sbjct: 445 PEEPEN 450
Score = 32.0 bits (71), Expect = 0.43
Identities = 20/66 (30%), Positives = 28/66 (42%)
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKK 71
S P S S P PS S P V S P + + + PP+ A PS + +
Sbjct: 608 SPPPPSPVYSPPPPSHSPPPPVYSPPPPTFSPPPTHNTNQPPMGAPTPTQAPTPSSETTQ 667
Query: 72 LRTPSN 77
+ TPS+
Sbjct: 668 VPTPSS 673
Score = 30.4 bits (67), Expect = 1.2
Identities = 20/69 (28%), Positives = 27/69 (38%), Gaps = 1/69 (1%)
Query: 13 SSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNS-KK 71
S P SS P P P +S P SP PP+ + P + P P+
Sbjct: 558 SPPPPVYSSPPPPHVYSPPPPVASPPPPSPPPPVHSPPPPPVFSPPPPVFSPPPPSPVYS 617
Query: 72 LRTPSNTPP 80
PS++PP
Sbjct: 618 PPPPSHSPP 626
Score = 28.5 bits (62), Expect = 4.7
Identities = 26/93 (27%), Positives = 41/93 (43%), Gaps = 27/93 (29%)
Query: 14 SPSSTSSSAPAPSRS----------QSPT-VTSSSPAA--------------QAASASPS 48
+PSS ++ P PS Q+PT V SS+P++ QA + SP
Sbjct: 660 TPSSETTQVPTPSSESDQSQILSPVQAPTPVQSSTPSSEPTQVPTPSSSESYQAPNLSPV 719
Query: 49 PTVPPLMAHVSPSWHD--PSPNSKKLRTPSNTP 79
P+ A + S P+P+S+ ++PS P
Sbjct: 720 QAPTPVQAPTTSSETSQVPTPSSESNQSPSQAP 752
Score = 28.1 bits (61), Expect = 6.2
Identities = 20/65 (30%), Positives = 28/65 (42%), Gaps = 2/65 (3%)
Query: 18 TSSSAPAPS--RSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLRTP 75
T + AP PS +Q PT +S S +Q S +PT S P+P+S +
Sbjct: 654 TPTQAPTPSSETTQVPTPSSESDQSQILSPVQAPTPVQSSTPSSEPTQVPTPSSSESYQA 713
Query: 76 SNTPP 80
N P
Sbjct: 714 PNLSP 718
Score = 28.1 bits (61), Expect = 6.2
Identities = 20/68 (29%), Positives = 24/68 (34%), Gaps = 1/68 (1%)
Query: 13 SSPSSTSSSAPAPSRSQSPTVTSSSPAAQAAS-ASPSPTVPPLMAHVSPSWHDPSPNSKK 71
S P S P P S P V S P + S PS + PP + P P P
Sbjct: 586 SPPPPVHSPPPPPVFSPPPPVFSPPPPSPVYSPPPPSHSPPPPVYSPPPPTFSPPPTHNT 645
Query: 72 LRTPSNTP 79
+ P P
Sbjct: 646 NQPPMGAP 653
Score = 27.7 bits (60), Expect = 8.1
Identities = 19/66 (28%), Positives = 23/66 (34%)
Query: 15 PSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLRT 74
P S S P+P S P V S P ++ P PP P P P
Sbjct: 538 PPMPSPSPPSPIYSPPPPVHSPPPPVYSSPPPPHVYSPPPPVASPPPPSPPPPVHSPPPP 597
Query: 75 PSNTPP 80
P +PP
Sbjct: 598 PVFSPP 603
>At4g27520 early nodulin-like 2 predicted GPI-anchored protein
Length = 349
Score = 37.4 bits (85), Expect = 0.010
Identities = 28/72 (38%), Positives = 38/72 (51%), Gaps = 7/72 (9%)
Query: 12 SSSPSSTSSSAPAPSRSQSPTVTSSSP-AAQAASASP--SPTVPPLMAHVSPSWHDPSPN 68
S SP S ++S PAP +S SP SS+P + A +P S T+PP A P P
Sbjct: 207 SGSPVSPTTSPPAPPKSTSPVSPSSAPMTSPPAPMAPKSSSTIPPSSA---PMTSPPGSM 263
Query: 69 SKKLRTP-SNTP 79
+ K +P SN+P
Sbjct: 264 APKSSSPVSNSP 275
Score = 32.3 bits (72), Expect = 0.33
Identities = 24/65 (36%), Positives = 29/65 (43%), Gaps = 11/65 (16%)
Query: 16 SSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLRTP 75
SST S P + SP SSSP + S P T PP AH SP S +P
Sbjct: 147 SSTPGSMTPPGGAHSPK--SSSPVSPTTSP-PGSTTPPGGAH--------SPKSSSAVSP 195
Query: 76 SNTPP 80
+ +PP
Sbjct: 196 ATSPP 200
Score = 32.0 bits (71), Expect = 0.43
Identities = 19/71 (26%), Positives = 35/71 (48%), Gaps = 4/71 (5%)
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
S S+ SP+++ + AP + T+S PA +++ SP+ P+ + +P +P
Sbjct: 189 SSSAVSPATSPPGSMAPKSGSPVSPTTSPPAPPKSTSPVSPSSAPMTSPPAPM----APK 244
Query: 69 SKKLRTPSNTP 79
S PS+ P
Sbjct: 245 SSSTIPPSSAP 255
Score = 30.4 bits (67), Expect = 1.2
Identities = 28/83 (33%), Positives = 35/83 (41%), Gaps = 14/83 (16%)
Query: 9 SLSSSSPSSTSSSAPA----PSRSQSPTVTSS-----SPAAQAASASPSPTVPPLMAHVS 59
S SSSP S ++S P P + SP +S+ SP A S SP P
Sbjct: 161 SPKSSSPVSPTTSPPGSTTPPGGAHSPKSSSAVSPATSPPGSMAPKSGSPVSPTTSPPAP 220
Query: 60 PSWHDP-SPNSKKLRTPSNTPPA 81
P P SP+S P +PPA
Sbjct: 221 PKSTSPVSPSS----APMTSPPA 239
Score = 30.4 bits (67), Expect = 1.2
Identities = 24/72 (33%), Positives = 35/72 (48%), Gaps = 7/72 (9%)
Query: 12 SSSPSSTSSSAPAPSRSQ----SPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSP 67
SS+P ++ + AP S SPTV+ S A S S SP+ P + + PS PS
Sbjct: 252 SSAPMTSPPGSMAPKSSSPVSNSPTVSPS--LAPGGSTSSSPSDSPSGSAMGPSGDGPSA 309
Query: 68 NSKKLRTPSNTP 79
+ + TP+ P
Sbjct: 310 -AGDISTPAGAP 320
Score = 28.9 bits (63), Expect = 3.6
Identities = 28/86 (32%), Positives = 37/86 (42%), Gaps = 18/86 (20%)
Query: 13 SSPSSTSSSAPAPSRSQSPT-------VTSSSPAAQAASASPSPTVPPLMAH-------V 58
S+ S ++AP S S T SSSP + S P T PP AH V
Sbjct: 135 STAQSPHAAAPGSSTPGSMTPPGGAHSPKSSSPVSPTTS-PPGSTTPPGGAHSPKSSSAV 193
Query: 59 SPSWHDP---SPNSKKLRTPSNTPPA 81
SP+ P +P S +P+ +PPA
Sbjct: 194 SPATSPPGSMAPKSGSPVSPTTSPPA 219
>At3g50580 proline-rich protein
Length = 265
Score = 37.4 bits (85), Expect = 0.010
Identities = 25/75 (33%), Positives = 34/75 (45%), Gaps = 6/75 (8%)
Query: 9 SLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPN 68
S S P + S P P+ +SP+ S +P + SP+ PP +PS P+P
Sbjct: 86 STPSPPPPAPKKSPPPPTPKKSPSPPSLTPFVPHPTPKKSPSPPP-----TPSLPPPAP- 139
Query: 69 SKKLRTPSNTPPALK 83
K TPS PP K
Sbjct: 140 KKSPSTPSLPPPTPK 154
Score = 37.0 bits (84), Expect = 0.013
Identities = 25/71 (35%), Positives = 33/71 (46%), Gaps = 13/71 (18%)
Query: 13 SSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKL 72
S+PS+ S PAP +S P SP+ P++ P + H +P PSP
Sbjct: 83 STPSTPSPPPPAPKKSPPPPTPKKSPS--------PPSLTPFVPHPTPK-KSPSPPP--- 130
Query: 73 RTPSNTPPALK 83
TPS PPA K
Sbjct: 131 -TPSLPPPAPK 140
Score = 36.2 bits (82), Expect = 0.023
Identities = 25/70 (35%), Positives = 31/70 (43%), Gaps = 5/70 (7%)
Query: 11 SSSSPSSTSSSA-PAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNS 69
S S PS T P P +S SP T S P + +P++PP SP P P S
Sbjct: 107 SPSPPSLTPFVPHPTPKKSPSPPPTPSLPPPAPKKSPSTPSLPPPTPKKSP----PPPPS 162
Query: 70 KKLRTPSNTP 79
+PSN P
Sbjct: 163 HHSSSPSNPP 172
>At2g42840 En/Spm-like transposon protein
Length = 306
Score = 37.4 bits (85), Expect = 0.010
Identities = 23/68 (33%), Positives = 35/68 (50%), Gaps = 4/68 (5%)
Query: 14 SPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLR 73
SPS+ S +P PS + +P+ S +P S +P+P PP PS H +P+
Sbjct: 66 SPSTPSHPSP-PSHTPTPSTPSHTPTPHTPSHTPTPHTPPCNCGSPPS-HPSTPSHPS-- 121
Query: 74 TPSNTPPA 81
TPS+ P+
Sbjct: 122 TPSHPTPS 129
Score = 34.3 bits (77), Expect = 0.086
Identities = 21/59 (35%), Positives = 29/59 (48%), Gaps = 8/59 (13%)
Query: 21 SAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLRTPSNTP 79
S P+ S P+ T S + PSP+ P +H SP H P+P+ TPS+TP
Sbjct: 39 SPPSGSHGTPPSHTPPSSNCGSPPYDPSPSTP---SHPSPPSHTPTPS-----TPSHTP 89
Score = 30.0 bits (66), Expect = 1.6
Identities = 22/72 (30%), Positives = 33/72 (45%), Gaps = 7/72 (9%)
Query: 11 SSSSPSSTSSSAP---APSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSP 67
S + PSS S P +PS P+ S +P S +P+P P +H +P+ H P
Sbjct: 50 SHTPPSSNCGSPPYDPSPSTPSHPSPPSHTPTPSTPSHTPTPHTP---SH-TPTPHTPPC 105
Query: 68 NSKKLRTPSNTP 79
N + +TP
Sbjct: 106 NCGSPPSHPSTP 117
Score = 28.5 bits (62), Expect = 4.7
Identities = 17/62 (27%), Positives = 28/62 (44%), Gaps = 4/62 (6%)
Query: 14 SPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLR 73
+PS T + P SP S+P+ + + P+P+ PP + S P P + +
Sbjct: 93 TPSHTPTPHTPPCNCGSPPSHPSTPSHPSTPSHPTPSHPPSGGYYS----SPPPRTPVVV 148
Query: 74 TP 75
TP
Sbjct: 149 TP 150
Score = 27.7 bits (60), Expect = 8.1
Identities = 21/66 (31%), Positives = 28/66 (41%), Gaps = 6/66 (9%)
Query: 14 SPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPSWHDPSPNSKKLR 73
SP S S P PS + + S P + S P+ P S H P+P+
Sbjct: 39 SPPSGSHGTP-PSHTPPSSNCGSPPYDPSPSTPSHPSPPSHTPTPSTPSHTPTPH----- 92
Query: 74 TPSNTP 79
TPS+TP
Sbjct: 93 TPSHTP 98
>At2g39340 unknown protein
Length = 989
Score = 37.4 bits (85), Expect = 0.010
Identities = 23/81 (28%), Positives = 37/81 (45%), Gaps = 2/81 (2%)
Query: 2 ADAFFIFSLSSSSPSSTSSSAPAPSRSQSPTVTSSSPAAQAASASPSPTVPPLMAHVSPS 61
+ +F + ++S P+S S AP+ + P SS P +Q A+ S + H +PS
Sbjct: 262 SQSFPVPGVTSEMPASNSQPAPSYVQPWRPETDSSHPPSQQPGAAVSTSNDTYWMHQAPS 321
Query: 62 W--HDPSPNSKKLRTPSNTPP 80
H P P ++P T P
Sbjct: 322 LQAHHPVPPQNNYQSPLETKP 342
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.337 0.143 0.471
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,517,620
Number of Sequences: 26719
Number of extensions: 273950
Number of successful extensions: 6573
Number of sequences better than 10.0: 261
Number of HSP's better than 10.0 without gapping: 127
Number of HSP's successfully gapped in prelim test: 137
Number of HSP's that attempted gapping in prelim test: 3563
Number of HSP's gapped (non-prelim): 1730
length of query: 305
length of database: 11,318,596
effective HSP length: 99
effective length of query: 206
effective length of database: 8,673,415
effective search space: 1786723490
effective search space used: 1786723490
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146747.7


