

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146747.6 + phase: 0
(498 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
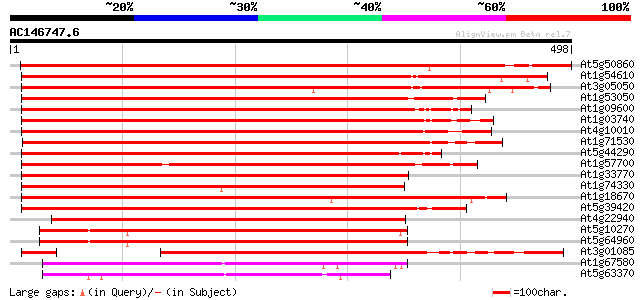
Score E
Sequences producing significant alignments: (bits) Value
At5g50860 cyclin-dependent protein kinase-like 714 0.0
At1g54610 Unknown protein (T22H22.5) 660 0.0
At3g05050 putative cyclin-dependent protein kinase 590 e-169
At1g53050 cell division-related protein, putative 587 e-168
At1g09600 putative protein kinase 556 e-159
At1g03740 putative protein kinase 540 e-154
At4g10010 protein kinase like protein 538 e-153
At1g71530 protein kinase-like 528 e-150
At5g44290 cyclin-dependent protein kinase-like protein 525 e-149
At1g57700 523 e-149
At1g33770 protein kinase, putative 523 e-149
At1g74330 putative protein kinase 515 e-146
At1g18670 hypothetical protein 503 e-142
At5g39420 cyclin-dependent kinase CDC2C (CDC2CAt) 493 e-140
At4g22940 putative cdc2 kinase homolog 414 e-116
At5g10270 cdc2-like protein kinase 348 3e-96
At5g64960 cdc2-like protein kinase 343 1e-94
At3g01085 putative protein 306 2e-83
At1g67580 putative protein kinase 241 5e-64
At5g63370 protein kinase 241 7e-64
>At5g50860 cyclin-dependent protein kinase-like
Length = 580
Score = 714 bits (1844), Expect = 0.0
Identities = 353/493 (71%), Positives = 415/493 (83%), Gaps = 13/493 (2%)
Query: 10 QGWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFD 69
+GWP WL+A G++I D TPRRA ++EKL KIGQGTYSNVYKAKDL++GKIVALKKVRFD
Sbjct: 89 EGWPPWLIAACGDSIKDLTPRRATTYEKLEKIGQGTYSNVYKAKDLLSGKIVALKKVRFD 148
Query: 70 NLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGV 129
NLE ESVKFMAREILVLR+L+HPNV+KL+GLVTSR+SCSLYLVFEYMEHDL+GL+A QG+
Sbjct: 149 NLEAESVKFMAREILVLRRLNHPNVIKLQGLVTSRVSCSLYLVFEYMEHDLSGLAATQGL 208
Query: 130 KFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKK 189
KF PQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDN+GILKIADFGLATFY+P +K
Sbjct: 209 KFDLPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNDGILKIADFGLATFYDPKQK 268
Query: 190 QSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIF 249
Q+MTSRVVTLWYRPPELLLGAT YG G+DLWSAGCI+AELLAGKP+MPGRTEVEQLHKIF
Sbjct: 269 QTMTSRVVTLWYRPPELLLGATSYGTGVDLWSAGCIMAELLAGKPVMPGRTEVEQLHKIF 328
Query: 250 KLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGT 309
KLCGSP++ YW+K++LPNAT+FKPQ PYKRC++E F F PSS+ L+++LL IDP RGT
Sbjct: 329 KLCGSPSDSYWKKYRLPNATLFKPQHPYKRCVAEAFNGFTPSSVHLVETLLTIDPADRGT 388
Query: 310 ASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVR 369
+++ALN EFFTTEP C+PSSLPKYPPSKEL+VK+RDEE RRQK L GK + +DGA+R+R
Sbjct: 389 STSALNSEFFTTEPLPCDPSSLPKYPPSKELNVKLRDEELRRQKGLAGKGSGIDGARRIR 448
Query: 370 AR--ERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKG 427
R GRAIPAPEANAE Q NLDRWR ++ N KSKSEKFPPPHQDGAVGYP ++ SK
Sbjct: 449 YRGDRTGRAIPAPEANAESQANLDRWRSISQTNGKSKSEKFPPPHQDGAVGYPLEDLSKK 508
Query: 428 PVSFGA-SDTSFSSGTFNVKPSGPTRSHDGTGLHKGTKTKKEESQMASSWKFMRPFK-PS 485
FGA ++TSF S +S +GT + K + K+ ++ ASS K++ K P
Sbjct: 509 TSVFGAKTETSFGL-------SRSLKSGEGTSMRK--ISNKDGARGASSRKYIWGLKPPP 559
Query: 486 TIGLSMDLLFRSK 498
+GLSMDLLFRS+
Sbjct: 560 ALGLSMDLLFRSR 572
>At1g54610 Unknown protein (T22H22.5)
Length = 572
Score = 660 bits (1702), Expect = 0.0
Identities = 334/476 (70%), Positives = 392/476 (82%), Gaps = 11/476 (2%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL GEA+ W PR+A++FEK+ KIGQGTYSNVYKAKD++TGKIVALKKVRFDN
Sbjct: 94 GWPSWLSDACGEALNGWVPRKADTFEKIDKIGQGTYSNVYKAKDMLTGKIVALKKVRFDN 153
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
LEPESVKFMAREILVLR+LDHPNVVKLEGLVTSRMSCSLYLVF+YM+HDLAGL++ VK
Sbjct: 154 LEPESVKFMAREILVLRRLDHPNVVKLEGLVTSRMSCSLYLVFQYMDHDLAGLASSPVVK 213
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
F+E +VKC M+QL+SGLEHCHSRGVLHRDIKGSNLLID+ G+LKIADFGLAT ++PN K+
Sbjct: 214 FSESEVKCLMRQLISGLEHCHSRGVLHRDIKGSNLLIDDGGVLKIADFGLATIFDPNHKR 273
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
MTSRVVTLWYR PELLLGAT YGVGIDLWSAGCILAELLAG+PIMPGRTEVEQLHKI+K
Sbjct: 274 PMTSRVVTLWYRAPELLLGATDYGVGIDLWSAGCILAELLAGRPIMPGRTEVEQLHKIYK 333
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP+E+YW+K K + I+KP++PYKR I ETFKDFPPSSLPLID+LL+I+P+ R TA
Sbjct: 334 LCGSPSEDYWKKGKFTHGAIYKPREPYKRSIRETFKDFPPSSLPLIDALLSIEPEDRQTA 393
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
SAAL EFFT+EPYACEP+ LPKYPPSKE+D K RDEE RRQ+A + KA DGA++ R
Sbjct: 394 SAALKSEFFTSEPYACEPADLPKYPPSKEIDAKRRDEETRRQRAAS-KAQG-DGARKNRH 451
Query: 371 RER-GRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQD-GAVGYPQDESSKGP 428
R+R RA+PAPEANAE+Q+N+DR R++THANAKSKSEKFPPPHQD GA+G P S
Sbjct: 452 RDRSNRALPAPEANAELQSNVDRRRLITHANAKSKSEKFPPPHQDGGAMGVPLGASQHID 511
Query: 429 VSFGASD--TSFSSGTFNV-KPSGPTRSHDGTG----LHKGTKTKKEESQMASSWK 477
+F D SF+S +FN K PT+ +G G KK+++ +S K
Sbjct: 512 PTFIPRDMVPSFTSSSFNFSKDEPPTQVQTWSGPLGHPITGVSRKKKDNTKSSKGK 567
>At3g05050 putative cyclin-dependent protein kinase
Length = 593
Score = 590 bits (1521), Expect = e-169
Identities = 306/481 (63%), Positives = 363/481 (74%), Gaps = 15/481 (3%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL V GEA+ W PR+A+SFEK+ KIG GTYSNVYKAKD +TG IVALKKVR D
Sbjct: 114 GWPSWLSEVCGEALSGWLPRKADSFEKIDKIGSGTYSNVYKAKDSLTGNIVALKKVRCDV 173
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
E ES+KFMAREIL+LR+LDHPNV+KLEGLVTSRMS SLYLVF YM+HDLAGL+A +K
Sbjct: 174 NERESLKFMAREILILRRLDHPNVIKLEGLVTSRMSSSLYLVFRYMDHDLAGLAASPEIK 233
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FTE QVKC+MKQLLSGLEHCH+RGVLHRDIKGSNLLID+ G+L+I DFGLATF++ +K+Q
Sbjct: 234 FTEQQVKCYMKQLLSGLEHCHNRGVLHRDIKGSNLLIDDGGVLRIGDFGLATFFDASKRQ 293
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
MT+RVVTLWYR PELL G Y VG+DLWSAGCILAELLAG+ IMPGR EVEQLH+I+K
Sbjct: 294 EMTNRVVTLWYRSPELLHGVVEYSVGVDLWSAGCILAELLAGRAIMPGRNEVEQLHRIYK 353
Query: 251 LCGSPAEEYWRKHKLPNA---TIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRR 307
LCGSP+EEYW+K +LP+ KP YKR I E +KDF P +L L+D+LLA+DP R
Sbjct: 354 LCGSPSEEYWKKIRLPSTHKHAHHKPLPQYKRRIREVYKDFSPEALSLLDTLLALDPAER 413
Query: 308 GTASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKR 367
TA+ L +FFTTEP AC+PS LPKYPPSKE+D K RDEE RRQ+ KA G +R
Sbjct: 414 QTATDVLMSDFFTTEPLACQPSDLPKYPPSKEIDAKRRDEEYRRQREAR-KAQGESG-RR 471
Query: 368 VRARERG-RAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDES-- 424
+R RER RA+PAPEANAE Q+N+DR R++THANAKSKSEKFPPPHQDG++G+ S
Sbjct: 472 MRPRERAPRAMPAPEANAENQSNIDRMRMITHANAKSKSEKFPPPHQDGSLGFQVGSSRR 531
Query: 425 ---SKGPVSFGASDTSFSSGTFNV--KPSGPTRSHDGTGLHKGTKTKKEESQMASSWKFM 479
S+ P S + +S+S F P P + D T + K E +MAS K
Sbjct: 532 LDPSEIPYSTNSFTSSYSKEPFQTWSGPLAPIGAPDSTTRRRNDINK--ERRMASKVKGK 589
Query: 480 R 480
R
Sbjct: 590 R 590
>At1g53050 cell division-related protein, putative
Length = 694
Score = 587 bits (1513), Expect = e-168
Identities = 284/412 (68%), Positives = 340/412 (81%), Gaps = 6/412 (1%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP WL +VAGEAI W PRRA+SFEKL KIGQGTYSNVY+A+DL KIVALKKVRFDN
Sbjct: 110 GWPPWLASVAGEAIRGWVPRRADSFEKLDKIGQGTYSNVYRARDLDQKKIVALKKVRFDN 169
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
LEPESV+FMAREI +LR+LDHPN++KLEGLVTSRMSCSLYLVFEYMEHDLAGL++ +K
Sbjct: 170 LEPESVRFMAREIQILRRLDHPNIIKLEGLVTSRMSCSLYLVFEYMEHDLAGLASHPAIK 229
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
F+E QVKC+++QLL GL+HCHSRGVLHRDIKGSNLLIDN G+LKIADFGLA+F++P + Q
Sbjct: 230 FSESQVKCYLQQLLHGLDHCHSRGVLHRDIKGSNLLIDNSGVLKIADFGLASFFDPRQTQ 289
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYRPPELLLGAT YG +DLWSAGCILAEL AGKPIMPGRTEVEQLHKIFK
Sbjct: 290 PLTSRVVTLWYRPPELLLGATRYGAAVDLWSAGCILAELYAGKPIMPGRTEVEQLHKIFK 349
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP E+YW K +LP+ATIFKP QPYKR + ETFK+FP +L L+++LL+++PD RGTA
Sbjct: 350 LCGSPTEDYWVKSRLPHATIFKPTQPYKRLVGETFKEFPQPALALLETLLSVNPDDRGTA 409
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
+AAL EFF+T P C+PSSLPKYPPSKELD +MRDEE+RRQ N + R
Sbjct: 410 TAALKSEFFSTRPLPCDPSSLPKYPPSKELDARMRDEESRRQVG----GNRDQRHQERRG 465
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQD 422
+ RAIPAP+ANAE+ ++ + + + + +S+SEKF P ++ A G+P D
Sbjct: 466 TKESRAIPAPDANAELVASMQKRQ--SQSTNRSRSEKFNPHPEEVASGFPID 515
>At1g09600 putative protein kinase
Length = 714
Score = 556 bits (1433), Expect = e-159
Identities = 271/400 (67%), Positives = 333/400 (82%), Gaps = 7/400 (1%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL +VAGEAI W PR+A+SFEKL KIGQGTYS+VYKA+DL T ++VALKKVRF N
Sbjct: 139 GWPSWLASVAGEAINGWIPRKADSFEKLEKIGQGTYSSVYKARDLETNQLVALKKVRFAN 198
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
++P+SV+FMAREI++LR+LDHPNV+KLEGL+TSR+S S+YL+FEYMEHDLAGL++ G+
Sbjct: 199 MDPDSVRFMAREIIILRRLDHPNVMKLEGLITSRVSGSMYLIFEYMEHDLAGLASTPGIN 258
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
F+E Q+KC+MKQLL GLEHCHSRGVLHRDIKGSNLL+D+ LKI DFGLA FY ++KQ
Sbjct: 259 FSEAQIKCYMKQLLHGLEHCHSRGVLHRDIKGSNLLLDHNNNLKIGDFGLANFYQGHQKQ 318
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYRPPELLLG+T YGV +DLWS GCILAEL GKPIMPGRTEVEQLHKIFK
Sbjct: 319 PLTSRVVTLWYRPPELLLGSTDYGVTVDLWSTGCILAELFTGKPIMPGRTEVEQLHKIFK 378
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP+EEYW+ KLP+ATIFKPQQPYKRC++ETFK P S+L L++ LLA++PD RGT
Sbjct: 379 LCGSPSEEYWKISKLPHATIFKPQQPYKRCVAETFKSLPSSALALVEVLLAVEPDARGTT 438
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
++AL EFFTT P A +PSSLPKY P KE+DVK ++EEA+R+K + K N +K+V +
Sbjct: 439 ASALESEFFTTSPLASDPSSLPKYQPRKEIDVKAQEEEAKRKKDTSSKQN---DSKQV-S 494
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPP 410
RE +A+PAP++NAE T++ + R H N S S+KF P
Sbjct: 495 RE-SKAVPAPDSNAESLTSIQK-RQGQH-NQVSNSDKFNP 531
>At1g03740 putative protein kinase
Length = 740
Score = 540 bits (1392), Expect = e-154
Identities = 264/419 (63%), Positives = 330/419 (78%), Gaps = 14/419 (3%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL++VAGE++ DW PRRAN+FEKL KIGQGTYS+VY+A+DL+ KIVALKKVRFD
Sbjct: 189 GWPSWLVSVAGESLVDWAPRRANTFEKLEKIGQGTYSSVYRARDLLHNKIVALKKVRFDL 248
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
+ ESVKFMAREI+V+R+LDHPNV+KLEGL+T+ +S SLYLVFEYM+HDL GLS+ GVK
Sbjct: 249 NDMESVKFMAREIIVMRRLDHPNVLKLEGLITAPVSSSLYLVFEYMDHDLLGLSSLPGVK 308
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FTEPQVKC+M+QLLSGLEHCHSRGVLHRDIKGSNLLID++G+LKIADFGLATF++P K
Sbjct: 309 FTEPQVKCYMRQLLSGLEHCHSRGVLHRDIKGSNLLIDSKGVLKIADFGLATFFDPAKSV 368
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
S+TS VVTLWYRPPELLLGA+ YGVG+DLWS GCIL EL AGKPI+PG+TEVEQLHKIFK
Sbjct: 369 SLTSHVVTLWYRPPELLLGASHYGVGVDLWSTGCILGELYAGKPILPGKTEVEQLHKIFK 428
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP E YWRK KLP++ FK PY+R +SE FKDFP S L L+++LL+IDPD R +A
Sbjct: 429 LCGSPTENYWRKQKLPSSAGFKTAIPYRRKVSEMFKDFPASVLSLLETLLSIDPDHRSSA 488
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
AL E+F T+P+AC+PS+LPKYPPSKE+D KMRDE R+Q K D R+ +
Sbjct: 489 DRALESEYFKTKPFACDPSNLPKYPPSKEIDAKMRDEAKRQQPMRAEKQERQDSMTRI-S 547
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPV 429
ER + +P +AN + +++ + + +S+++ F + ++ + GPV
Sbjct: 548 HER-KFVPPVKANNSLSMTMEK----QYKDLRSRNDSFK--------SFKEERTPHGPV 593
>At4g10010 protein kinase like protein
Length = 649
Score = 538 bits (1386), Expect = e-153
Identities = 267/417 (64%), Positives = 319/417 (76%), Gaps = 13/417 (3%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL +VAGEAI W PRRA SFEKL KIGQGTYS+VY+A+DL TGK+VA+KKVRF N
Sbjct: 132 GWPSWLTSVAGEAIKGWVPRRAESFEKLDKIGQGTYSSVYRARDLETGKMVAMKKVRFVN 191
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
++PESV+FMAREI +LRKLDHPNV+KLE LVTS++S SLYLVFEYMEHDL+GL+ GVK
Sbjct: 192 MDPESVRFMAREINILRKLDHPNVMKLECLVTSKLSGSLYLVFEYMEHDLSGLALRPGVK 251
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FTE Q+KC+MKQLLSGLEHCHSRG+LHRDIKG NLL++N+G+LKI DFGLA Y+P + Q
Sbjct: 252 FTESQIKCYMKQLLSGLEHCHSRGILHRDIKGPNLLVNNDGVLKIGDFGLANIYHPEQDQ 311
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYR PELLLGAT YG GIDLWS GCIL EL GKPIMPGRTEVEQ+HKIFK
Sbjct: 312 PLTSRVVTLWYRAPELLLGATEYGPGIDLWSVGCILTELFLGKPIMPGRTEVEQMHKIFK 371
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
CGSP+++YW+K KLP AT FKPQQPYKR + ETFK+ PPS+L L+D LL+++P +RGTA
Sbjct: 372 FCGSPSDDYWQKTKLPLATSFKPQQPYKRVLLETFKNLPPSALALVDKLLSLEPAKRGTA 431
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
S+ L+ +FFT EP C SSLPKYPPSKELD K+RDEEARR+K+ K + +R +
Sbjct: 432 SSTLSSKFFTMEPLPCNVSSLPKYPPSKELDAKVRDEEARRKKSETVKGRGPESVRR-GS 490
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKG 427
R+ PE A Q+ + + K P +D G D KG
Sbjct: 491 RDFKSTATTPEFVASGQSK------------DTITTKRFNPQEDSRTGLRGDRDQKG 535
>At1g71530 protein kinase-like
Length = 655
Score = 528 bits (1361), Expect = e-150
Identities = 267/426 (62%), Positives = 322/426 (74%), Gaps = 13/426 (3%)
Query: 12 WPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNL 71
WPSWL +VAGEAI W PR A SFEKL KIGQGTYS+VYKA+DL TGKIVA+KKVRF N+
Sbjct: 124 WPSWLASVAGEAIKGWVPRCAESFEKLDKIGQGTYSSVYKARDLETGKIVAMKKVRFVNM 183
Query: 72 EPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKF 131
+PESV+FMAREIL+LRKLDHPNV+KLEGLVTSR+S SLYLVFEYMEHDLAGL+A G+KF
Sbjct: 184 DPESVRFMAREILILRKLDHPNVMKLEGLVTSRLSGSLYLVFEYMEHDLAGLAATPGIKF 243
Query: 132 TEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQS 191
+EPQ+KC+M+QL GLEHCH RG+LHRDIKGSNLLI+NEG+LKI DFGLA FY +
Sbjct: 244 SEPQIKCYMQQLFRGLEHCHRRGILHRDIKGSNLLINNEGVLKIGDFGLANFYRGDGDLQ 303
Query: 192 MTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKL 251
+TSRVVTLWYR PELLLGAT YG IDLWSAGCIL EL AGKPIMPGRTEVEQ+HKIFKL
Sbjct: 304 LTSRVVTLWYRAPELLLGATEYGPAIDLWSAGCILTELFAGKPIMPGRTEVEQMHKIFKL 363
Query: 252 CGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTAS 311
CGSP+E+YWR+ LP AT FKP PYK ++ETF FP S+L LI+ LLAI+P++RG+A+
Sbjct: 364 CGSPSEDYWRRATLPLATSFKPSHPYKPVLAETFNHFPSSALMLINKLLAIEPEKRGSAA 423
Query: 312 AALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRAR 371
+ L EFFTTEP PS+LP+YPPSKELD K+R+EEAR+ +A K + R R +
Sbjct: 424 STLRSEFFTTEPLPANPSNLPRYPPSKELDAKLRNEEARKLRAEGNKRRGGETVTRGRPK 483
Query: 372 ERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPVSF 431
+ + PE A Q+ VT + K K++ ++G G+ + +G
Sbjct: 484 DL-KTAQTPEFMAAGQSK------VTCISHKFKTD------EEGGTGFRIEPPRRGIQQN 530
Query: 432 GASDTS 437
G + S
Sbjct: 531 GKAHAS 536
>At5g44290 cyclin-dependent protein kinase-like protein
Length = 644
Score = 525 bits (1352), Expect = e-149
Identities = 253/373 (67%), Positives = 310/373 (82%), Gaps = 3/373 (0%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP+WL++VAGEA+ +WTPRRA++FEKL KIGQGTYS+VYKA+DL KIVALK+VRFD
Sbjct: 113 GWPAWLVSVAGEALVNWTPRRASTFEKLEKIGQGTYSSVYKARDLTNNKIVALKRVRFDL 172
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
+ ESVKFMAREI+V+R+LDHPNV+KLEGL+T+ +S SLYLVFEYM+HDL GL++ G+K
Sbjct: 173 SDLESVKFMAREIIVMRRLDHPNVLKLEGLITASVSSSLYLVFEYMDHDLVGLASIPGIK 232
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
F+EPQVKC+M+QLLSGL HCHSRGVLHRDIKGSNLLID+ G+LKIADFGLATF++P
Sbjct: 233 FSEPQVKCYMQQLLSGLHHCHSRGVLHRDIKGSNLLIDSNGVLKIADFGLATFFDPQNCV 292
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYRPPELLLGA YGVG+DLWS GCIL EL +GKPI+ G+TEVEQLHKIFK
Sbjct: 293 PLTSRVVTLWYRPPELLLGACHYGVGVDLWSTGCILGELYSGKPILAGKTEVEQLHKIFK 352
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP E+YWRK KLP + F+P PY R ++E FKD P + L L+++LL+IDPDRRG+A
Sbjct: 353 LCGSPTEDYWRKLKLPPSAAFRPALPYGRRVAEMFKDLPTNVLSLLEALLSIDPDRRGSA 412
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
+ AL E+F TEP+AC+PSSLPKYPPSKE+D K+RD +A+RQ+ K D R R+
Sbjct: 413 ARALESEYFRTEPFACDPSSLPKYPPSKEIDAKIRD-DAKRQRPTQEKHERQDSQTR-RS 470
Query: 371 RERGRAIPAPEAN 383
ER + IP +AN
Sbjct: 471 HER-KLIPPVKAN 482
>At1g57700
Length = 686
Score = 523 bits (1348), Expect = e-149
Identities = 254/405 (62%), Positives = 317/405 (77%), Gaps = 13/405 (3%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL++VAGEAI W PR A+SFEKL IGQGTYS+VY+A+DL T +IVALKKVRF N
Sbjct: 122 GWPSWLVSVAGEAINGWIPRSADSFEKLEMIGQGTYSSVYRARDLETNQIVALKKVRFAN 181
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
++PESV+FMAREI++LR+L+HPNV+KLEGL+ S+ S S+YL+FEYM+HDLAGL++ G+K
Sbjct: 182 MDPESVRFMAREIIILRRLNHPNVMKLEGLIISKASGSMYLIFEYMDHDLAGLASTPGIK 241
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
F++ Q QLL GLEHCHS GVLHRDIK SNLL+D LKI DFGL+ FY +KQ
Sbjct: 242 FSQAQ------QLLLGLEHCHSCGVLHRDIKCSNLLLDRNNNLKIGDFGLSNFYRGQRKQ 295
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYRPPELLLG+T YGV +DLWS GCILAEL GKP++PGRTEVEQ+HKIFK
Sbjct: 296 PLTSRVVTLWYRPPELLLGSTDYGVTVDLWSTGCILAELFTGKPLLPGRTEVEQMHKIFK 355
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP+EEYWR+ +L +ATIFKPQ PYKRC+++TFKD P S+L L++ LLA++PD RGTA
Sbjct: 356 LCGSPSEEYWRRSRLRHATIFKPQHPYKRCVADTFKDLPSSALALLEVLLAVEPDARGTA 415
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
S+AL EFFTT+P+ EPSSLP+Y P KE D K+R+EEARR+K + K N ++ R
Sbjct: 416 SSALQSEFFTTKPFPSEPSSLPRYQPRKEFDAKLREEEARRRKGSSSKQN-----EQKRL 470
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDG 415
+A+PAP ANAE+ ++ + + N S SEKF P G
Sbjct: 471 ARESKAVPAPSANAELLASIQ--KRLGETNRTSISEKFNPEGDSG 513
>At1g33770 protein kinase, putative
Length = 614
Score = 523 bits (1347), Expect = e-149
Identities = 250/344 (72%), Positives = 293/344 (84%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP WL +VAGEAI W PRRA+SFEKL KIGQGTYS VYKA+DL TGKIVA+KKVRF N
Sbjct: 117 GWPYWLTSVAGEAIKGWVPRRADSFEKLDKIGQGTYSIVYKARDLETGKIVAMKKVRFAN 176
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
++PESV+FMAREI +LRKLDHPNV+KL+ LVTS++S SL+LVFEYMEHDL+GL+ GVK
Sbjct: 177 MDPESVRFMAREINILRKLDHPNVMKLQCLVTSKLSGSLHLVFEYMEHDLSGLALRPGVK 236
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FTEPQ+KCFMKQLL GLEHCHSRG+LHRDIKGSNLL++N+G+LKI DFGLA+FY P++ Q
Sbjct: 237 FTEPQIKCFMKQLLCGLEHCHSRGILHRDIKGSNLLVNNDGVLKIGDFGLASFYKPDQDQ 296
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYR PELLLG+T YG IDLWS GCILAEL KPIMPGRTEVEQ+HKIFK
Sbjct: 297 PLTSRVVTLWYRAPELLLGSTEYGPAIDLWSVGCILAELFVCKPIMPGRTEVEQMHKIFK 356
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP+EE+W K P AT +KPQ PYKR + ETFK+ SSL L+D LL+++P++R +A
Sbjct: 357 LCGSPSEEFWNTTKFPQATSYKPQHPYKRVLLETFKNLSSSSLDLLDKLLSVEPEKRCSA 416
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKA 354
S+ L EFFTTEP C SSLPKYPPSKELD K+RDEEA+R+KA
Sbjct: 417 SSTLLSEFFTTEPLPCHISSLPKYPPSKELDAKVRDEEAKRKKA 460
>At1g74330 putative protein kinase
Length = 445
Score = 515 bits (1326), Expect = e-146
Identities = 240/342 (70%), Positives = 290/342 (84%), Gaps = 2/342 (0%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP+WL VAGEAI W P R+++FEKL KIGQGTYSNV++A + TG+IVALKKVRFDN
Sbjct: 97 GWPAWLSNVAGEAIHGWVPLRSDAFEKLEKIGQGTYSNVFRAVETETGRIVALKKVRFDN 156
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
EPESVKFMAREIL+LR+L+HPN++KLEGL+TS++SC++ LVFEYMEHDL GL + +K
Sbjct: 157 FEPESVKFMAREILILRRLNHPNIIKLEGLITSKLSCNIQLVFEYMEHDLTGLLSSPDIK 216
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNP--NK 188
FT PQ+KC+MKQLLSGL+HCHSRGV+HRDIKGSNLL+ NEGILK+ADFGLA F N +K
Sbjct: 217 FTTPQIKCYMKQLLSGLDHCHSRGVMHRDIKGSNLLLSNEGILKVADFGLANFSNSSGHK 276
Query: 189 KQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKI 248
K+ +TSRVVTLWYRPPELLLGAT YG +DLWS GC+ AELL GKPI+ GRTEVEQLHKI
Sbjct: 277 KKPLTSRVVTLWYRPPELLLGATDYGASVDLWSVGCVFAELLLGKPILRGRTEVEQLHKI 336
Query: 249 FKLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRG 308
FKLCGSP E+YW+K KLP+A +FKPQQ Y C+ ET KD + + LI++LL+IDP +RG
Sbjct: 337 FKLCGSPPEDYWKKSKLPHAMLFKPQQTYDSCLRETLKDLSETEINLIETLLSIDPHKRG 396
Query: 309 TASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEAR 350
TAS+AL ++FTT+P+AC+PSSLP YPPSKE+D K RDE AR
Sbjct: 397 TASSALVSQYFTTKPFACDPSSLPIYPPSKEIDTKHRDEAAR 438
>At1g18670 hypothetical protein
Length = 662
Score = 503 bits (1294), Expect = e-142
Identities = 251/440 (57%), Positives = 321/440 (72%), Gaps = 10/440 (2%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP+WL VAGEAI W P R+++FEKL KIGQGTYS+V++A++ TG+IVALKKVRFDN
Sbjct: 107 GWPAWLSNVAGEAIHGWVPFRSDAFEKLEKIGQGTYSSVFRARETETGRIVALKKVRFDN 166
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
EPESV+FMAREIL+LRKL+HPN++KLEG+VTS++SCS++LVFEYMEHDL GL + +
Sbjct: 167 FEPESVRFMAREILILRKLNHPNIIKLEGIVTSKLSCSIHLVFEYMEHDLTGLLSSPDID 226
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPN-KK 189
FT PQ+KC+MKQLLSGL+HCH+RGV+HRDIKGSNLL++NEGILK+ADFGLA F N + K
Sbjct: 227 FTTPQIKCYMKQLLSGLDHCHARGVMHRDIKGSNLLVNNEGILKVADFGLANFCNASGNK 286
Query: 190 QSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIF 249
Q +TSRVVTLWYRPPELLLGAT YG +DLWS GC+ AELL GKP++ GRTEVEQLHKIF
Sbjct: 287 QPLTSRVVTLWYRPPELLLGATEYGASVDLWSVGCVFAELLIGKPVLQGRTEVEQLHKIF 346
Query: 250 KLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISET--FKDFPPSSLPLIDSLLAIDPDRR 307
KLCGSP E+YW+K KLP+A +FKPQQ Y C+ ET K + + LI++LL+I P +R
Sbjct: 347 KLCGSPPEDYWKKSKLPHAMLFKPQQHYDGCLRETLKLKGLSDADINLIETLLSIQPHKR 406
Query: 308 GTASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKR 367
GTAS AL ++FT++P+AC+PSSLP Y PSKE+D K R++ R++ + NG+
Sbjct: 407 GTASTALVSQYFTSKPFACDPSSLPVYSPSKEIDAKHREDTTRKKISGNGRRGTESRKPT 466
Query: 368 VRARERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKF-----PPPHQDGAVGYPQD 422
+ + PA + Q R H + S S F P H+ + ++
Sbjct: 467 RKPPAFAKLAPAEDVRHHSQKFQKRNGHSVHNSIDSDSTLFEKMQKPSNHEKDEASHVKN 526
Query: 423 ESSKGPVSF-GASDTSFSSG 441
+S+G V F G S SSG
Sbjct: 527 -ASQGDVPFSGPLQVSVSSG 545
>At5g39420 cyclin-dependent kinase CDC2C (CDC2CAt)
Length = 644
Score = 493 bits (1270), Expect = e-140
Identities = 230/395 (58%), Positives = 306/395 (77%), Gaps = 4/395 (1%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP+WL + A EA+ W P +A +F+KL KIGQGTYS+V++A+++ TGK+VALKKV+FDN
Sbjct: 81 GWPAWLCSAAAEAVHGWVPLKAEAFQKLEKIGQGTYSSVFRAREVETGKMVALKKVKFDN 140
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
L+PES++FMAREIL+LRKL+HPN++KLEG+VTSR S S+YLVFEYMEHDLAGLS+ ++
Sbjct: 141 LQPESIRFMAREILILRKLNHPNIMKLEGIVTSRASSSIYLVFEYMEHDLAGLSSNPDIR 200
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FTEPQ+KC+MKQLL GLEHCH RGV+HRDIK SN+L++N+G+LK+ DFGLA P+ K
Sbjct: 201 FTEPQIKCYMKQLLWGLEHCHMRGVIHRDIKASNILVNNKGVLKLGDFGLANVVTPSNKN 260
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYR PELL+G+T YGV +DLWS GC+ AE+L GKPI+ GRTE+EQLHKI+K
Sbjct: 261 QLTSRVVTLWYRAPELLMGSTSYGVSVDLWSVGCVFAEILMGKPILKGRTEIEQLHKIYK 320
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP + +W++ KLP+AT FKPQ Y+ + E KD + + L+++LL+++PD+RGTA
Sbjct: 321 LCGSPQDSFWKRTKLPHATSFKPQHTYEATLRERCKDLSATGVYLLETLLSMEPDKRGTA 380
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
S+ALN E+F T PYAC+PSSLPKYPP+KE+D K RD+ R++ L + + V G K R
Sbjct: 381 SSALNSEYFLTRPYACDPSSLPKYPPNKEMDAKYRDDMRRKRANLKLRDSGV-GRKHKRP 439
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKS 405
RA P+ A++ D V N S++
Sbjct: 440 H---RAEYDPKNYAKLPIRKDTLEVKNIPNEASRA 471
>At4g22940 putative cdc2 kinase homolog
Length = 353
Score = 414 bits (1064), Expect = e-116
Identities = 193/315 (61%), Positives = 252/315 (79%), Gaps = 1/315 (0%)
Query: 38 LAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKLDHPNVVKL 97
L +IG GT+S V+KA+DL+ K VALK++RFD ES+K +AREI++LRKLDHPNV+KL
Sbjct: 1 LQQIGGGTFSKVFKARDLLRNKTVALKRIRFDINNSESIKCIAREIIILRKLDHPNVIKL 60
Query: 98 EGLV-TSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVL 156
EGL+ S +LYL+FEYMEHDL GLS+ GV F+EPQVKC+M+QLL GL+HCH+ VL
Sbjct: 61 EGLMLVDHDSSTLYLIFEYMEHDLLGLSSLLGVHFSEPQVKCYMRQLLRGLDHCHTNHVL 120
Query: 157 HRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGVG 216
HRD+K SNLLI+ +G+LKIADFGLATF++P+ +T+ V TLWYRPPELLLGA+ YG+G
Sbjct: 121 HRDMKSSNLLINGDGVLKIADFGLATFFDPHNSVPLTTHVATLWYRPPELLLGASHYGIG 180
Query: 217 IDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQP 276
+DLWS GC++ EL AGKPI+PG+ E +QLHKIFKLCGSP+++YW K KL +T +P P
Sbjct: 181 VDLWSTGCVIGELYAGKPILPGKNETDQLHKIFKLCGSPSDDYWTKLKLQLSTPLRPIYP 240
Query: 277 YKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFFTTEPYACEPSSLPKYPP 336
Y I+ETFK FP S + L+++LL+IDPD RGTA++AL ++F TEP AC+PS LPKYP
Sbjct: 241 YGSHIAETFKQFPASVISLLETLLSIDPDFRGTAASALKSKYFKTEPLACDPSCLPKYPS 300
Query: 337 SKELDVKMRDEEARR 351
SKE+++KMRD ++
Sbjct: 301 SKEINIKMRDNTRKQ 315
>At5g10270 cdc2-like protein kinase
Length = 505
Score = 348 bits (894), Expect = 3e-96
Identities = 182/350 (52%), Positives = 237/350 (67%), Gaps = 24/350 (6%)
Query: 27 WTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILV 85
W R + FEKL +IG+GTY VY AK++ TG+IVALKK+R DN E E A REI +
Sbjct: 18 WGSRSVDCFEKLEQIGEGTYGQVYMAKEIKTGEIVALKKIRMDN-EREGFPITAIREIKI 76
Query: 86 LRKLDHPNVVKLEGLVTS--------------RMSCSLYLVFEYMEHDLAGLSAGQGVKF 131
L+KL H NV++L+ +VTS + +Y+VFEYM+HDL GL+ G++F
Sbjct: 77 LKKLHHENVIQLKEIVTSPGRDRDDQGKPDNNKYKGGIYMVFEYMDHDLTGLADRPGLRF 136
Query: 132 TEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQS 191
T PQ+KC+MKQLL+GL +CH VLHRDIKGSNLLIDNEG LK+ADFGLA Y+ + +
Sbjct: 137 TVPQIKCYMKQLLTGLHYCHVNQVLHRDIKGSNLLIDNEGNLKLADFGLARSYSHDHTGN 196
Query: 192 MTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKL 251
+T+RV+TLWYRPPELLLGAT YG ID+WS GCI AELL KPI+PG+ E EQL+KIF+L
Sbjct: 197 LTNRVITLWYRPPELLLGATKYGPAIDMWSVGCIFAELLHAKPILPGKNEQEQLNKIFEL 256
Query: 252 CGSPAEEYW-RKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
CGSP E+ W K+P FKP +P KR + E F+ F +L L++ +L +DP +R +A
Sbjct: 257 CGSPDEKLWPGVSKMPWFNNFKPARPLKRRVREFFRHFDRHALELLEKMLVLDPAQRISA 316
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMR-------DEEARRQK 353
AL+ E+F T+P C+P SLP Y S E K + +E A+RQK
Sbjct: 317 KDALDAEYFWTDPLPCDPKSLPTYESSHEFQTKKKRQQQRQNEEAAKRQK 366
>At5g64960 cdc2-like protein kinase
Length = 513
Score = 343 bits (880), Expect = 1e-94
Identities = 176/343 (51%), Positives = 231/343 (67%), Gaps = 17/343 (4%)
Query: 27 WTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILV 85
W R + FEKL +IG+GTY VY AK++ TG+IVALKK+R DN E E A REI +
Sbjct: 18 WGSRSVDCFEKLEQIGEGTYGQVYMAKEIKTGEIVALKKIRMDN-EREGFPITAIREIKI 76
Query: 86 LRKLDHPNVVKLEGLVTS--------------RMSCSLYLVFEYMEHDLAGLSAGQGVKF 131
L+KL H NV+ L+ +VTS + +Y+VFEYM+HDL GL+ G++F
Sbjct: 77 LKKLHHENVIHLKEIVTSPGRDRDDQGKPDNNKYKGGIYMVFEYMDHDLTGLADRPGLRF 136
Query: 132 TEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQS 191
T PQ+KC+MKQLL+GL +CH VLHRDIKGSNLLIDNEG LK+ADFGLA Y+ + +
Sbjct: 137 TVPQIKCYMKQLLTGLHYCHVNQVLHRDIKGSNLLIDNEGNLKLADFGLARSYSHDHTGN 196
Query: 192 MTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKL 251
+T+RV+TLWYRPPELLLGAT YG ID+WS GCI AELL GKPI+PG+TE EQL+KI++L
Sbjct: 197 LTNRVITLWYRPPELLLGATKYGPAIDMWSVGCIFAELLNGKPILPGKTENEQLNKIYEL 256
Query: 252 CGSPAEEYW-RKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
CGSP E W K+P K +P KR + E ++ F +L L++ +L +DP +R A
Sbjct: 257 CGSPDESNWPGVSKMPWYNQMKSSRPLKRRVREIYRHFDRHALELLEKMLVLDPSQRICA 316
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQK 353
AL+ E+F T+P C+P SLP Y S E K + ++ R +
Sbjct: 317 KDALDAEYFWTDPLPCDPKSLPTYESSHEFQTKKKRQQMRHNE 359
>At3g01085 putative protein
Length = 536
Score = 306 bits (784), Expect = 2e-83
Identities = 157/359 (43%), Positives = 235/359 (64%), Gaps = 31/359 (8%)
Query: 135 QVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTS 194
++KC+MKQLL G+EHCH RG++HRDIK +N+L++N+G+LK+ADFGLA P K +TS
Sbjct: 120 KIKCYMKQLLWGVEHCHLRGIMHRDIKAANILVNNKGVLKLADFGLANIVTPRNKNQLTS 179
Query: 195 RVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGS 254
RVVTLWYR PELL+G+T Y V +DLWS GC+ AE+L G+P++ GRTE+EQLHKI+KL GS
Sbjct: 180 RVVTLWYRAPELLMGSTSYSVSVDLWSVGCVFAEILTGRPLLKGRTEIEQLHKIYKLSGS 239
Query: 255 PAEEYWRKHKL-PNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAA 313
P EE+W K+KL P +F+PQ Y+ C+ E F +FP +++ L+++LL+IDP++RGTAS+A
Sbjct: 240 PDEEFWEKNKLHPQTKMFRPQHQYEGCLRERFDEFPKTAINLLENLLSIDPEKRGTASSA 299
Query: 314 LNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRARER 373
L E+F T+PYAC+PS+LPKYPP+KE+D K R+E RR++ K + + K ++R
Sbjct: 300 LMSEYFNTQPYACDPSTLPKYPPNKEMDAKYREELQRRRRVSIKKRDNLATKKLGKSR-- 357
Query: 374 GRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPVSFGA 433
A + TNL+ R+ TH K ++E + V P + S +
Sbjct: 358 -------RATVKEPTNLN--RLPTHQETKKEAE------TEIVVQTPSETSQ----ATTR 398
Query: 434 SDTSFSSGTFNVKPSGPTRSHDGTGLHKGTKTKKEESQMASSWKFMRPFKPSTI-GLSM 491
S+ ++S + P+ +G K++E+ +AS+ +++P S + G+SM
Sbjct: 399 SEFPYNSLSQTTAPA--------SGFAWAGTKKRKENDVASTLTYIQPGSASHVSGMSM 449
Score = 43.5 bits (101), Expect = 3e-04
Identities = 18/31 (58%), Positives = 20/31 (64%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKI 41
GWPSWL + A EA+ W P RA FEK KI
Sbjct: 91 GWPSWLSSAAPEAVHGWVPLRAEDFEKREKI 121
>At1g67580 putative protein kinase
Length = 752
Score = 241 bits (616), Expect = 5e-64
Identities = 143/340 (42%), Positives = 193/340 (56%), Gaps = 17/340 (5%)
Query: 30 RRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKL 89
R + FE+L KI +GTY VY+AKD TG+IVALKKV+ + REI +L
Sbjct: 401 RSVDEFERLNKIDEGTYGVVYRAKDKKTGEIVALKKVKMEKEREGFPLTSLREINILLSF 460
Query: 90 DHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEH 149
HP++V ++ +V S+++V EYMEHDL L +F++ +VKC M QLL G+++
Sbjct: 461 HHPSIVDVKEVVVGSSLDSIFMVMEYMEHDLKALMETMKQRFSQSEVKCLMLQLLEGVKY 520
Query: 150 CHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLG 209
H VLHRD+K SNLL++N G LKI DFGLA Y K T VVTLWYR PELLLG
Sbjct: 521 LHDNWVLHRDLKTSNLLLNNRGELKICDFGLARQYGSPLK-PYTHLVVTLWYRAPELLLG 579
Query: 210 ATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRK-HKLPNA 268
A Y ID+WS GCI+AELL P+ G+TE +QL KIF++ G+P E W KLP
Sbjct: 580 AKQYSTAIDMWSLGCIMAELLMKAPLFNGKTEFDQLDKIFRILGTPNESIWPGFSKLPGV 639
Query: 269 TIFKPQQPY----KRCISETFKDFP---PSSLPLIDSLLAIDPDRRGTASAALNHEFFTT 321
+ + Y K+ + +F P + L++ LL DP+RR T + AL H++F
Sbjct: 640 KVNFVKHQYNLLRKKFPATSFTGAPVLSDAGFDLLNKLLTYDPERRITVNEALKHDWFRE 699
Query: 322 EPYACEPSSLPKYPPSKELD------VKMRD--EEARRQK 353
P +P +P D VK D EE RR++
Sbjct: 700 VPLPKSKDFMPTFPAQHAQDRRGRRMVKSPDPLEEQRRKE 739
>At5g63370 protein kinase
Length = 612
Score = 241 bits (615), Expect = 7e-64
Identities = 140/326 (42%), Positives = 191/326 (57%), Gaps = 22/326 (6%)
Query: 30 RRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRF--DNLEPESVKFMA--REILV 85
R N F+KL KI +GTY VYKA+D T +IVALKK++ D E E + REI +
Sbjct: 292 RSVNEFQKLNKINEGTYGIVYKARDEKTKEIVALKKIKMKEDRFEEEYGFPLTSLREINI 351
Query: 86 LRKLDHPNVVKL-EGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLL 144
L +HP +V + E +V + +Y+V E++EHDL G+ + F+ +VKC M QLL
Sbjct: 352 LLSCNHPAIVNVKEVVVGGKNDNDVYMVMEHLEHDLRGVMDRRKEPFSTSEVKCLMMQLL 411
Query: 145 SGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPP 204
GL++ H+ ++HRD+K SNLL++N G LKI DFG+A Y K T V+T WYRPP
Sbjct: 412 DGLKYLHTNWIIHRDLKPSNLLMNNCGELKICDFGMARQYGSPIKP-YTQMVITQWYRPP 470
Query: 205 ELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRK-H 263
ELLLGA Y +D+WS GCI+AELL+ KP+ PG++E++QL KIF + G+P E W
Sbjct: 471 ELLLGAKEYSTAVDMWSVGCIMAELLSQKPLFPGKSELDQLQKIFAVLGTPNEAIWPGFS 530
Query: 264 KLPNATIFKPQQPYKRCISETFKDFPPSS-----------LPLIDSLLAIDPDRRGTASA 312
PNA P QPY + K FP S L++SLL +DP++R T
Sbjct: 531 SFPNAKAKFPTQPY----NMLRKKFPAISFVGGQILSERGFDLLNSLLTLDPEKRLTVED 586
Query: 313 ALNHEFFTTEPYACEPSSLPKYPPSK 338
ALNH +F P +P YPP +
Sbjct: 587 ALNHGWFHEVPLPKSKDFMPTYPPKR 612
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.134 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,002,311
Number of Sequences: 26719
Number of extensions: 541599
Number of successful extensions: 3675
Number of sequences better than 10.0: 989
Number of HSP's better than 10.0 without gapping: 889
Number of HSP's successfully gapped in prelim test: 100
Number of HSP's that attempted gapping in prelim test: 1315
Number of HSP's gapped (non-prelim): 1162
length of query: 498
length of database: 11,318,596
effective HSP length: 103
effective length of query: 395
effective length of database: 8,566,539
effective search space: 3383782905
effective search space used: 3383782905
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146747.6


