

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146722.4 - phase: 1
(88 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
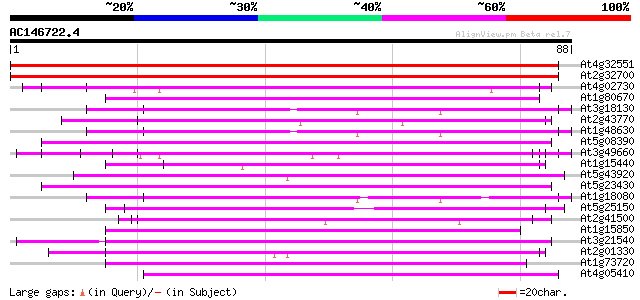
Score E
Sequences producing significant alignments: (bits) Value
At4g32551 Leunig protein 144 1e-35
At2g32700 unknown protein 106 2e-24
At4g02730 putative WD-repeat protein 52 4e-08
At1g80670 mRNA export protein, putative 52 5e-08
At3g18130 protein kinase C-receptor/G-protein, putative 49 3e-07
At2g43770 putative splicing factor 49 3e-07
At1g48630 unknown protein 49 3e-07
At5g08390 katanin p80 subunit - like protein 48 7e-07
At3g49660 putative WD-40 repeat - protein 48 7e-07
At1g15440 unknown protein 48 1e-06
At5g43920 WD-repeat protein-like 47 2e-06
At5g23430 unknown protein 47 2e-06
At1g18080 putative guanine nucleotide-binding protein beta sub... 47 2e-06
At5g25150 transcription initiation factor IID-associated factor-... 46 3e-06
At2g41500 putative U4/U6 small nuclear ribonucleoprotein 45 6e-06
At1g15850 hypothetical protein 45 6e-06
At3g21540 WD repeat like protein 45 8e-06
At2g01330 putative WD-40-repeat protein 45 8e-06
At1g73720 unknown protein 45 8e-06
At4g05410 U3 snoRNP-associated -like protein 44 1e-05
>At4g32551 Leunig protein
Length = 931
Score = 144 bits (362), Expect = 1e-35
Identities = 66/86 (76%), Positives = 75/86 (86%)
Query: 1 FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD 60
FTF+E+NSVRAST KV C HFSSDGK+LAS GHDKK VLWYTD++K K TLEEH+ +ITD
Sbjct: 639 FTFTEVNSVRASTTKVTCCHFSSDGKMLASAGHDKKAVLWYTDTMKPKTTLEEHTAMITD 698
Query: 61 VRFSPSMPRLATSSYDKTVRVWDVDN 86
+RFSPS RLATSS+DKTVRVWD DN
Sbjct: 699 IRFSPSQLRLATSSFDKTVRVWDADN 724
Score = 28.9 bits (63), Expect = 0.47
Identities = 13/33 (39%), Positives = 21/33 (63%)
Query: 50 TLEEHSLLITDVRFSPSMPRLATSSYDKTVRVW 82
TL H LIT + S + +A++S+DK V++W
Sbjct: 898 TLPAHEGLITSLAVSTATGLVASASHDKLVKLW 930
>At2g32700 unknown protein
Length = 787
Score = 106 bits (265), Expect = 2e-24
Identities = 46/86 (53%), Positives = 68/86 (78%)
Query: 1 FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD 60
F+F+E++ +R S +KV+C FS DGKLLAS GHDKKV +W ++L+ ++T EEH+ +ITD
Sbjct: 498 FSFNEVSCIRKSASKVICCSFSYDGKLLASAGHDKKVFIWNMETLQVESTPEEHAHIITD 557
Query: 61 VRFSPSMPRLATSSYDKTVRVWDVDN 86
VRF P+ +LATSS+DKT+++WD +
Sbjct: 558 VRFRPNSTQLATSSFDKTIKIWDASD 583
Score = 33.5 bits (75), Expect = 0.019
Identities = 20/56 (35%), Positives = 34/56 (60%), Gaps = 2/56 (3%)
Query: 27 LLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVW 82
LL GG+ + + LW T K T+ H +I+ + SPS +A++S+DK+V++W
Sbjct: 733 LLVIGGY-QAIELWNTMENKCM-TVAGHECVISALAQSPSTGVVASASHDKSVKIW 786
Score = 25.4 bits (54), Expect = 5.2
Identities = 9/28 (32%), Positives = 17/28 (60%)
Query: 59 TDVRFSPSMPRLATSSYDKTVRVWDVDN 86
T VRF P + ++ + TV ++D++N
Sbjct: 638 TQVRFQPRTGQFLAAASENTVSIFDIEN 665
>At4g02730 putative WD-repeat protein
Length = 333
Score = 52.4 bits (124), Expect = 4e-08
Identities = 27/81 (33%), Positives = 42/81 (51%)
Query: 3 FSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVR 62
+ L ++ T + C FS+DG LLAS DK ++LW + E HS I+D+
Sbjct: 33 YRHLKTLEGHTAAISCVKFSNDGNLLASASVDKTMILWSATNYSLIHRYEGHSSGISDLA 92
Query: 63 FSPSMPRLATSSYDKTVRVWD 83
+S ++S D T+R+WD
Sbjct: 93 WSSDSHYTCSASDDCTLRIWD 113
Score = 48.5 bits (114), Expect = 6e-07
Identities = 22/78 (28%), Positives = 42/78 (53%)
Query: 6 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSP 65
L +R TN V C +F+ L+ SG D+ + +W + K ++ HS+ I+ V F+
Sbjct: 121 LKVLRGHTNFVFCVNFNPPSNLIVSGSFDETIRIWEVKTGKCVRMIKAHSMPISSVHFNR 180
Query: 66 SMPRLATSSYDKTVRVWD 83
+ ++S+D + ++WD
Sbjct: 181 DGSLIVSASHDGSCKIWD 198
Score = 42.7 bits (99), Expect = 3e-05
Identities = 28/78 (35%), Positives = 39/78 (49%), Gaps = 5/78 (6%)
Query: 13 TNKVVC--SHFS-SDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPR 69
TNKV C S FS ++GK + SG D V LW + LE H+ + V P
Sbjct: 255 TNKVFCITSAFSVTNGKYIVSGSEDNCVYLWDLQARNILQRLEGHTDAVISVSCHPVQNE 314
Query: 70 LATSS--YDKTVRVWDVD 85
+++S DKT+R+W D
Sbjct: 315 ISSSGNHLDKTIRIWKQD 332
Score = 33.5 bits (75), Expect = 0.019
Identities = 17/39 (43%), Positives = 24/39 (60%)
Query: 50 TLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDVDNVS 88
TLE H+ I+ V+FS LA++S DKT+ +W N S
Sbjct: 38 TLEGHTAAISCVKFSNDGNLLASASVDKTMILWSATNYS 76
>At1g80670 mRNA export protein, putative
Length = 349
Score = 52.0 bits (123), Expect = 5e-08
Identities = 27/68 (39%), Positives = 38/68 (55%)
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
V+CS + DG + SGG DK+ +W S Q T+ H I + + P M LAT S+
Sbjct: 75 VLCSAWKDDGTTVFSGGCDKQAKMWPLLSGGQPVTVAMHEGPIAAMAWIPGMNLLATGSW 134
Query: 76 DKTVRVWD 83
DKT++ WD
Sbjct: 135 DKTLKYWD 142
Score = 26.2 bits (56), Expect = 3.0
Identities = 10/27 (37%), Positives = 16/27 (59%)
Query: 58 ITDVRFSPSMPRLATSSYDKTVRVWDV 84
I+ + FSP L +S+D VR W++
Sbjct: 28 ISSLSFSPRADILVATSWDNQVRCWEI 54
>At3g18130 protein kinase C-receptor/G-protein, putative
Length = 326
Score = 49.3 bits (116), Expect = 3e-07
Identities = 28/79 (35%), Positives = 47/79 (59%), Gaps = 6/79 (7%)
Query: 13 TNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEE---HSLLITDVRFSPS--M 67
T V+ FS+D + + S D+ + LW T + K T+ E H ++ VRFSP+ +
Sbjct: 105 TKDVLSVAFSTDNRQIVSASRDRTIKLWNTLG-ECKYTISEGDGHKEWVSCVRFSPNTLV 163
Query: 68 PRLATSSYDKTVRVWDVDN 86
P + ++S+DKTV+VW++ N
Sbjct: 164 PTIVSASWDKTVKVWNLQN 182
Score = 40.4 bits (93), Expect = 2e-04
Identities = 24/67 (35%), Positives = 40/67 (58%), Gaps = 2/67 (2%)
Query: 22 SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRV 81
S DG L ASGG D ++LW K+ +LE S +I + FSP+ L ++ + ++R+
Sbjct: 202 SPDGSLCASGGKDGVILLWDLAEGKKLYSLEAGS-IIHSLCFSPNRYWLCAAT-ENSIRI 259
Query: 82 WDVDNVS 88
WD+++ S
Sbjct: 260 WDLESKS 266
Score = 35.8 bits (81), Expect = 0.004
Identities = 20/71 (28%), Positives = 36/71 (50%), Gaps = 6/71 (8%)
Query: 19 SHF------SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLAT 72
SHF SSDG+ SG D ++ LW + + H+ + V FS ++ +
Sbjct: 63 SHFVEDVVLSSDGQFALSGSWDGELRLWDLATGETTRRFVGHTKDVLSVAFSTDNRQIVS 122
Query: 73 SSYDKTVRVWD 83
+S D+T+++W+
Sbjct: 123 ASRDRTIKLWN 133
Score = 32.7 bits (73), Expect = 0.032
Identities = 20/71 (28%), Positives = 32/71 (44%), Gaps = 2/71 (2%)
Query: 16 VVCSHFSSDGKL--LASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATS 73
V C FS + + + S DK V +W + K + +L HS + V SP A+
Sbjct: 152 VSCVRFSPNTLVPTIVSASWDKTVKVWNLQNCKLRNSLVGHSGYLNTVAVSPDGSLCASG 211
Query: 74 SYDKTVRVWDV 84
D + +WD+
Sbjct: 212 GKDGVILLWDL 222
Score = 28.5 bits (62), Expect = 0.61
Identities = 22/82 (26%), Positives = 36/82 (43%), Gaps = 6/82 (7%)
Query: 9 VRASTNKVVCSHFSSDGK-LLASGGHDKKVVLW-YTDSLKQ----KATLEEHSLLITDVR 62
+RA T+ V D ++ + DK ++LW T K + L HS + DV
Sbjct: 11 MRAHTDIVTAIATPIDNSDIIVTASRDKSIILWKLTKDDKSYGVAQRRLTGHSHFVEDVV 70
Query: 63 FSPSMPRLATSSYDKTVRVWDV 84
S + S+D +R+WD+
Sbjct: 71 LSSDGQFALSGSWDGELRLWDL 92
>At2g43770 putative splicing factor
Length = 343
Score = 49.3 bits (116), Expect = 3e-07
Identities = 24/66 (36%), Positives = 38/66 (57%), Gaps = 1/66 (1%)
Query: 21 FSSDGKLLASGGHDKKVVLWYTDS-LKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTV 79
F+ G L+ASG HD+++ LW K L+ H I D+ ++ ++ ++S DKTV
Sbjct: 61 FNPAGTLIASGSHDREIFLWRVHGDCKNFMVLKGHKNAILDLHWTSDGSQIVSASPDKTV 120
Query: 80 RVWDVD 85
R WDV+
Sbjct: 121 RAWDVE 126
Score = 40.8 bits (94), Expect = 1e-04
Identities = 23/79 (29%), Positives = 39/79 (49%), Gaps = 5/79 (6%)
Query: 9 VRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD---VRFSP 65
++ N ++ H++SDG + S DK V W ++ KQ + EHS + R P
Sbjct: 92 LKGHKNAILDLHWTSDGSQIVSASPDKTVRAWDVETGKQIKKMAEHSSFVNSCCPTRRGP 151
Query: 66 SMPRLATSSYDKTVRVWDV 84
P + + S D T ++WD+
Sbjct: 152 --PLIISGSDDGTAKLWDM 168
Score = 37.0 bits (84), Expect = 0.002
Identities = 21/70 (30%), Positives = 31/70 (44%)
Query: 15 KVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSS 74
++ FS + +GG D V +W + TLE H IT + SP L T+
Sbjct: 182 QITAVSFSDAADKIFTGGVDNDVKVWDLRKGEATMTLEGHQDTITGMSLSPDGSYLLTNG 241
Query: 75 YDKTVRVWDV 84
D + VWD+
Sbjct: 242 MDNKLCVWDM 251
Score = 36.6 bits (83), Expect = 0.002
Identities = 20/71 (28%), Positives = 37/71 (51%), Gaps = 1/71 (1%)
Query: 14 NKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATS 73
N + CS +S DG + +G D+ V +W T S + L H+ + + F P+ P + +
Sbjct: 274 NLLKCS-WSPDGTKVTAGSSDRMVHIWDTTSRRTIYKLPGHTGSVNECVFHPTEPIIGSC 332
Query: 74 SYDKTVRVWDV 84
S DK + + ++
Sbjct: 333 SSDKNIYLGEI 343
Score = 24.6 bits (52), Expect = 8.8
Identities = 9/34 (26%), Positives = 20/34 (58%)
Query: 51 LEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDV 84
L H + ++F+P+ +A+ S+D+ + +W V
Sbjct: 49 LSGHPSAVYTMKFNPAGTLIASGSHDREIFLWRV 82
>At1g48630 unknown protein
Length = 326
Score = 49.3 bits (116), Expect = 3e-07
Identities = 28/79 (35%), Positives = 47/79 (59%), Gaps = 6/79 (7%)
Query: 13 TNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEE---HSLLITDVRFSPS--M 67
T V+ FS+D + + S D+ + LW T + K T+ E H ++ VRFSP+ +
Sbjct: 105 TKDVLSVAFSTDNRQIVSASRDRTIKLWNTLG-ECKYTISEADGHKEWVSCVRFSPNTLV 163
Query: 68 PRLATSSYDKTVRVWDVDN 86
P + ++S+DKTV+VW++ N
Sbjct: 164 PTIVSASWDKTVKVWNLQN 182
Score = 40.4 bits (93), Expect = 2e-04
Identities = 24/67 (35%), Positives = 40/67 (58%), Gaps = 2/67 (2%)
Query: 22 SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRV 81
S DG L ASGG D ++LW K+ +LE S +I + FSP+ L ++ + ++R+
Sbjct: 202 SPDGSLCASGGKDGVILLWDLAEGKKLYSLEAGS-IIHSLCFSPNRYWLCAAT-ENSIRI 259
Query: 82 WDVDNVS 88
WD+++ S
Sbjct: 260 WDLESKS 266
Score = 36.2 bits (82), Expect = 0.003
Identities = 20/71 (28%), Positives = 36/71 (50%), Gaps = 6/71 (8%)
Query: 19 SHF------SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLAT 72
SHF SSDG+ SG D ++ LW + + H+ + V FS ++ +
Sbjct: 63 SHFVQDVVLSSDGQFALSGSWDGELRLWDLATGESTRRFVGHTKDVLSVAFSTDNRQIVS 122
Query: 73 SSYDKTVRVWD 83
+S D+T+++W+
Sbjct: 123 ASRDRTIKLWN 133
Score = 34.7 bits (78), Expect = 0.009
Identities = 21/71 (29%), Positives = 32/71 (44%), Gaps = 2/71 (2%)
Query: 16 VVCSHFSSDGKL--LASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATS 73
V C FS + + + S DK V +W + K + TL HS + V SP A+
Sbjct: 152 VSCVRFSPNTLVPTIVSASWDKTVKVWNLQNCKLRNTLAGHSGYLNTVAVSPDGSLCASG 211
Query: 74 SYDKTVRVWDV 84
D + +WD+
Sbjct: 212 GKDGVILLWDL 222
Score = 27.3 bits (59), Expect = 1.4
Identities = 16/63 (25%), Positives = 28/63 (44%), Gaps = 5/63 (7%)
Query: 27 LLASGGHDKKVVLW-YTDSLKQKATLEE----HSLLITDVRFSPSMPRLATSSYDKTVRV 81
++ + DK ++LW T K + HS + DV S + S+D +R+
Sbjct: 30 VIVTSSRDKSIILWKLTKEDKSYGVAQRRMTGHSHFVQDVVLSSDGQFALSGSWDGELRL 89
Query: 82 WDV 84
WD+
Sbjct: 90 WDL 92
>At5g08390 katanin p80 subunit - like protein
Length = 823
Score = 48.1 bits (113), Expect = 7e-07
Identities = 26/80 (32%), Positives = 40/80 (49%)
Query: 6 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSP 65
+++ + T V F+ DG+ + SGG D V +W + K + H I + F P
Sbjct: 229 IHTYKGHTRGVNVLRFTPDGRWIVSGGEDNVVKVWDLTAGKLLHEFKSHEGKIQSLDFHP 288
Query: 66 SMPRLATSSYDKTVRVWDVD 85
LAT S DKTV+ WD++
Sbjct: 289 HEFLLATGSADKTVKFWDLE 308
Score = 37.7 bits (86), Expect = 0.001
Identities = 25/84 (29%), Positives = 40/84 (46%), Gaps = 2/84 (2%)
Query: 2 TFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDV 61
+F E + A+ N + SS ++L +GG D KV LW +L HS I V
Sbjct: 101 SFREFVAHSAAVNCLKIGRKSS--RVLVTGGEDHKVNLWAIGKPNAILSLYGHSSGIDSV 158
Query: 62 RFSPSMPRLATSSYDKTVRVWDVD 85
F S +A + T+++WD++
Sbjct: 159 TFDASEGLVAAGAASGTIKLWDLE 182
Score = 35.4 bits (80), Expect = 0.005
Identities = 18/64 (28%), Positives = 28/64 (43%)
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVR 80
F + L+A+G + LW + K TL H V F P A+ S D ++
Sbjct: 160 FDASEGLVAAGAASGTIKLWDLEEAKVVRTLTGHRSNCVSVNFHPFGEFFASGSLDTNLK 219
Query: 81 VWDV 84
+WD+
Sbjct: 220 IWDI 223
>At3g49660 putative WD-40 repeat - protein
Length = 317
Score = 48.1 bits (113), Expect = 7e-07
Identities = 23/64 (35%), Positives = 35/64 (53%)
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVR 80
FSSD + + S DK + LW ++ TL H+ V F+P + + S+D+TVR
Sbjct: 79 FSSDARFIVSASDDKTLKLWDVETGSLIKTLIGHTNYAFCVNFNPQSNMIVSGSFDETVR 138
Query: 81 VWDV 84
+WDV
Sbjct: 139 IWDV 142
Score = 46.6 bits (109), Expect = 2e-06
Identities = 25/82 (30%), Positives = 40/82 (48%)
Query: 2 TFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDV 61
T S + ++ TN C +F+ ++ SG D+ V +W + K L HS +T V
Sbjct: 102 TGSLIKTLIGHTNYAFCVNFNPQSNMIVSGSFDETVRIWDVTTGKCLKVLPAHSDPVTAV 161
Query: 62 RFSPSMPRLATSSYDKTVRVWD 83
F+ + +SSYD R+WD
Sbjct: 162 DFNRDGSLIVSSSYDGLCRIWD 183
Score = 45.8 bits (107), Expect = 4e-06
Identities = 25/67 (37%), Positives = 38/67 (56%), Gaps = 1/67 (1%)
Query: 17 VCSHFS-SDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
+ S FS ++GK + SG D V +W +S K LE H+ + +V P+ +A+ S
Sbjct: 246 ISSAFSVTNGKRIVSGSEDNCVHMWELNSKKLLQKLEGHTETVMNVACHPTENLIASGSL 305
Query: 76 DKTVRVW 82
DKTVR+W
Sbjct: 306 DKTVRIW 312
Score = 44.7 bits (104), Expect = 8e-06
Identities = 29/83 (34%), Positives = 44/83 (52%), Gaps = 6/83 (7%)
Query: 12 STNKVVCS-HFSSDGKLLASGGHDKKVVLWYTDSLKQKAT-----LEEHSLLITDVRFSP 65
S N+ V S FSSDG+LLAS DK + + +++ H I+DV FS
Sbjct: 22 SHNRAVSSVKFSSDGRLLASASADKTIRTYTINTINDPIAEPVQEFTGHENGISDVAFSS 81
Query: 66 SMPRLATSSYDKTVRVWDVDNVS 88
+ ++S DKT+++WDV+ S
Sbjct: 82 DARFIVSASDDKTLKLWDVETGS 104
Score = 38.1 bits (87), Expect = 8e-04
Identities = 20/82 (24%), Positives = 43/82 (52%), Gaps = 1/82 (1%)
Query: 6 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLK-QKATLEEHSLLITDVRFS 64
L + A ++ V F+ DG L+ S +D +W + + K +++ + ++ VRFS
Sbjct: 148 LKVLPAHSDPVTAVDFNRDGSLIVSSSYDGLCRIWDSGTGHCVKTLIDDENPPVSFVRFS 207
Query: 65 PSMPRLATSSYDKTVRVWDVDN 86
P+ + + D T+R+W++ +
Sbjct: 208 PNGKFILVGTLDNTLRLWNISS 229
Score = 32.0 bits (71), Expect = 0.055
Identities = 13/39 (33%), Positives = 25/39 (63%)
Query: 50 TLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDVDNVS 88
TL H+ ++ V+FS LA++S DKT+R + ++ ++
Sbjct: 19 TLTSHNRAVSSVKFSSDGRLLASASADKTIRTYTINTIN 57
Score = 28.1 bits (61), Expect = 0.80
Identities = 18/69 (26%), Positives = 30/69 (43%), Gaps = 3/69 (4%)
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEH---SLLITDVRFSPSMPRLATSSYDK 77
FS +GK + G D + LW S K T H I+ + R+ + S D
Sbjct: 206 FSPNGKFILVGTLDNTLRLWNISSAKFLKTYTGHVNAQYCISSAFSVTNGKRIVSGSEDN 265
Query: 78 TVRVWDVDN 86
V +W++++
Sbjct: 266 CVHMWELNS 274
>At1g15440 unknown protein
Length = 900
Score = 47.8 bits (112), Expect = 1e-06
Identities = 26/68 (38%), Positives = 35/68 (51%)
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
V C +S D +LLA+G D KV +W S T EH+ +T + F L ++S
Sbjct: 392 VNCVTYSPDSQLLATGADDNKVKVWNVMSGTCFITFTEHTNAVTALHFMADNHSLLSASL 451
Query: 76 DKTVRVWD 83
D TVR WD
Sbjct: 452 DGTVRAWD 459
Score = 38.1 bits (87), Expect = 8e-04
Identities = 21/61 (34%), Positives = 34/61 (55%), Gaps = 1/61 (1%)
Query: 25 GKLLASGGHDK-KVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWD 83
G ++ +G D ++ +W + + K L H + + FSP LA+SS+D TVR+WD
Sbjct: 486 GDVVCAGTLDSFEIFVWSKKTGQIKDILSGHEAPVHGLMFSPLTQLLASSSWDYTVRLWD 545
Query: 84 V 84
V
Sbjct: 546 V 546
Score = 35.4 bits (80), Expect = 0.005
Identities = 24/65 (36%), Positives = 32/65 (48%), Gaps = 4/65 (6%)
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLE--EHSLLITDVRFSPSMPRLATSSYDKT 78
FS +LLAS D V LW D K T+E H+ + V F P +LA+S+ D
Sbjct: 525 FSPLTQLLASSSWDYTVRLW--DVFASKGTVETFRHNHDVLTVAFRPDGKQLASSTLDGQ 582
Query: 79 VRVWD 83
+ WD
Sbjct: 583 INFWD 587
Score = 28.9 bits (63), Expect = 0.47
Identities = 24/78 (30%), Positives = 36/78 (45%), Gaps = 9/78 (11%)
Query: 12 STNKVVCSHFSSDGKLLASGGHDKKVVL---WYTDS--LKQKATLEEHSLLITDVRFSPS 66
S K+ + F+ G L G +L W T++ LKQ+ H + V +SP
Sbjct: 345 SRQKLTTAVFNERGNWLTFGCAKLGQLLVWDWRTETYILKQQG----HYFDVNCVTYSPD 400
Query: 67 MPRLATSSYDKTVRVWDV 84
LAT + D V+VW+V
Sbjct: 401 SQLLATGADDNKVKVWNV 418
>At5g43920 WD-repeat protein-like
Length = 308
Score = 47.0 bits (110), Expect = 2e-06
Identities = 26/80 (32%), Positives = 42/80 (52%), Gaps = 3/80 (3%)
Query: 11 ASTNKVVCSHFSSDGKLLASGGHDKKVVLWYT---DSLKQKATLEEHSLLITDVRFSPSM 67
A N+V FS+ GK LA+ D ++W + ++ K TLE H ++ V +SP
Sbjct: 222 AHKNEVWFVQFSNSGKYLATASSDCTAIIWKVLDDNKVELKHTLESHQNPVSFVSWSPDD 281
Query: 68 PRLATSSYDKTVRVWDVDNV 87
+L T + +++WDVD V
Sbjct: 282 TKLLTCGNAEVLKLWDVDTV 301
>At5g23430 unknown protein
Length = 837
Score = 47.0 bits (110), Expect = 2e-06
Identities = 25/80 (31%), Positives = 40/80 (49%)
Query: 6 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSP 65
+++ + T V F+ DG+ + SGG D V +W + K + H I + F P
Sbjct: 136 IHTYKGHTRGVNVLRFTPDGRWVVSGGEDNIVKVWDLTAGKLLTEFKSHEGQIQSLDFHP 195
Query: 66 SMPRLATSSYDKTVRVWDVD 85
LAT S D+TV+ WD++
Sbjct: 196 HEFLLATGSADRTVKFWDLE 215
Score = 36.6 bits (83), Expect = 0.002
Identities = 19/60 (31%), Positives = 30/60 (49%)
Query: 26 KLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDVD 85
++L +GG D KV LW +L HS I V F S +A + T+++WD++
Sbjct: 30 RVLVTGGEDHKVNLWAIGKPNAILSLYGHSSGIDSVTFDASEVLVAAGAASGTIKLWDLE 89
Score = 33.9 bits (76), Expect = 0.015
Identities = 18/64 (28%), Positives = 28/64 (43%)
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVR 80
F + L+A+G + LW + K TL H V F P A+ S D ++
Sbjct: 67 FDASEVLVAAGAASGTIKLWDLEEAKIVRTLTGHRSNCISVDFHPFGEFFASGSLDTNLK 126
Query: 81 VWDV 84
+WD+
Sbjct: 127 IWDI 130
>At1g18080 putative guanine nucleotide-binding protein beta
subunit
Length = 327
Score = 46.6 bits (109), Expect = 2e-06
Identities = 28/80 (35%), Positives = 45/80 (56%), Gaps = 7/80 (8%)
Query: 13 TNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEE----HSLLITDVRFSPS-- 66
T V+ FS D + + S D+ + LW T + K T+ E H ++ VRFSP+
Sbjct: 105 TKDVLSVAFSLDNRQIVSASRDRTIKLWNTLG-ECKYTISEGGEGHRDWVSCVRFSPNTL 163
Query: 67 MPRLATSSYDKTVRVWDVDN 86
P + ++S+DKTV+VW++ N
Sbjct: 164 QPTIVSASWDKTVKVWNLSN 183
Score = 38.1 bits (87), Expect = 8e-04
Identities = 24/67 (35%), Positives = 40/67 (58%), Gaps = 2/67 (2%)
Query: 22 SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRV 81
S DG L ASGG D V+LW K+ +LE +S +I + FSP+ L ++ + +++
Sbjct: 203 SPDGSLCASGGKDGVVLLWDLAEGKKLYSLEANS-VIHALCFSPNRYWLCAAT-EHGIKI 260
Query: 82 WDVDNVS 88
WD+++ S
Sbjct: 261 WDLESKS 267
Score = 35.4 bits (80), Expect = 0.005
Identities = 21/71 (29%), Positives = 33/71 (45%), Gaps = 2/71 (2%)
Query: 16 VVCSHFSSDG--KLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATS 73
V C FS + + S DK V +W + K ++TL H+ ++ V SP A+
Sbjct: 153 VSCVRFSPNTLQPTIVSASWDKTVKVWNLSNCKLRSTLAGHTGYVSTVAVSPDGSLCASG 212
Query: 74 SYDKTVRVWDV 84
D V +WD+
Sbjct: 213 GKDGVVLLWDL 223
Score = 34.3 bits (77), Expect = 0.011
Identities = 20/71 (28%), Positives = 35/71 (49%), Gaps = 6/71 (8%)
Query: 19 SHF------SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLAT 72
SHF SSDG+ SG D ++ LW + H+ + V FS ++ +
Sbjct: 63 SHFVEDVVLSSDGQFALSGSWDGELRLWDLAAGVSTRRFVGHTKDVLSVAFSLDNRQIVS 122
Query: 73 SSYDKTVRVWD 83
+S D+T+++W+
Sbjct: 123 ASRDRTIKLWN 133
Score = 30.0 bits (66), Expect = 0.21
Identities = 21/83 (25%), Positives = 36/83 (43%), Gaps = 6/83 (7%)
Query: 8 SVRASTNKVVCSHFSSDGK-LLASGGHDKKVVLWYTDSLKQ-----KATLEEHSLLITDV 61
++RA T+ V D ++ S DK ++LW + + L HS + DV
Sbjct: 10 TMRAHTDMVTAIATPIDNADIIVSASRDKSIILWKLTKDDKAYGVAQRRLTGHSHFVEDV 69
Query: 62 RFSPSMPRLATSSYDKTVRVWDV 84
S + S+D +R+WD+
Sbjct: 70 VLSSDGQFALSGSWDGELRLWDL 92
>At5g25150 transcription initiation factor IID-associated
factor-like protein
Length = 700
Score = 46.2 bits (108), Expect = 3e-06
Identities = 24/66 (36%), Positives = 36/66 (54%), Gaps = 3/66 (4%)
Query: 19 SHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKT 78
+ FS G AS HD+ +W D ++ + H ++DV + P+ +AT S DKT
Sbjct: 466 AQFSPFGHYFASCSHDRTARIWSMDRIQPLRIMAGH---LSDVDWHPNCNYIATGSSDKT 522
Query: 79 VRVWDV 84
VR+WDV
Sbjct: 523 VRLWDV 528
Score = 42.7 bits (99), Expect = 3e-05
Identities = 21/72 (29%), Positives = 34/72 (47%)
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
V + FS G + S D + LW T + H+ + D +FSP A+ S+
Sbjct: 421 VYSATFSPPGDFVLSSSADTTIRLWSTKLNANLVCYKGHNYPVWDAQFSPFGHYFASCSH 480
Query: 76 DKTVRVWDVDNV 87
D+T R+W +D +
Sbjct: 481 DRTARIWSMDRI 492
Score = 32.7 bits (73), Expect = 0.032
Identities = 24/93 (25%), Positives = 35/93 (36%), Gaps = 24/93 (25%)
Query: 14 NKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKAT----------------------- 50
N + CS S DG L+A G D + +W + Q +
Sbjct: 353 NGLNCSSISHDGSLVAGGFSDSSIKVWDMAKIGQAGSGALQAENDSSDQSIGPNGRRSYT 412
Query: 51 -LEEHSLLITDVRFSPSMPRLATSSYDKTVRVW 82
L HS + FSP + +SS D T+R+W
Sbjct: 413 LLLGHSGPVYSATFSPPGDFVLSSSADTTIRLW 445
Score = 31.2 bits (69), Expect = 0.094
Identities = 15/57 (26%), Positives = 27/57 (47%)
Query: 28 LASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDV 84
+A+G DK V LW + + H ++ + SP +A+ D T+ +WD+
Sbjct: 514 IATGSSDKTVRLWDVQTGECVRIFIGHRSMVLSLAMSPDGRYMASGDEDGTIMMWDL 570
Score = 24.6 bits (52), Expect = 8.8
Identities = 10/42 (23%), Positives = 21/42 (49%)
Query: 22 SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRF 63
S DG+ +ASG D +++W + + L H+ + + +
Sbjct: 550 SPDGRYMASGDEDGTIMMWDLSTARCITPLMGHNSCVWSLSY 591
>At2g41500 putative U4/U6 small nuclear ribonucleoprotein
Length = 554
Score = 45.1 bits (105), Expect = 6e-06
Identities = 26/65 (40%), Positives = 34/65 (52%), Gaps = 1/65 (1%)
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVR 80
FS LA+ D+ LW TD + T E H + V F PS L T+SYDKT R
Sbjct: 306 FSPVDDCLATASADRTAKLWKTDGTLLQ-TFEGHLDRLARVAFHPSGKYLGTTSYDKTWR 364
Query: 81 VWDVD 85
+WD++
Sbjct: 365 LWDIN 369
Score = 45.1 bits (105), Expect = 6e-06
Identities = 28/69 (40%), Positives = 37/69 (53%), Gaps = 2/69 (2%)
Query: 18 CSHFSSDGKLLASGGHDKKVVLWYTDSLKQK-ATLEEHSLLITDVRFSPSMPRLATSSYD 76
CS FS DGK+LA+ LW + A L++H TDV FSP LAT+S D
Sbjct: 261 CS-FSRDGKILATCSLSGVTKLWEMPQVTNTIAVLKDHKERATDVVFSPVDDCLATASAD 319
Query: 77 KTVRVWDVD 85
+T ++W D
Sbjct: 320 RTAKLWKTD 328
Score = 43.9 bits (102), Expect = 1e-05
Identities = 23/64 (35%), Positives = 35/64 (53%), Gaps = 1/64 (1%)
Query: 20 HFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPR-LATSSYDKT 78
+FS +G LASGG D + +W K + H+ L++ V++ P LAT+SYD
Sbjct: 430 NFSPNGYHLASGGEDNQCRIWDLRMRKSLYIIPAHANLVSQVKYEPQEGYFLATASYDMK 489
Query: 79 VRVW 82
V +W
Sbjct: 490 VNIW 493
Score = 37.7 bits (86), Expect = 0.001
Identities = 24/83 (28%), Positives = 35/83 (41%)
Query: 2 TFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDV 61
T +EL + V F DG L AS G D +W + + + H + V
Sbjct: 370 TGAELLLQEGHSRSVYGIAFQQDGALAASCGLDSLARVWDLRTGRSILVFQGHIKPVFSV 429
Query: 62 RFSPSMPRLATSSYDKTVRVWDV 84
FSP+ LA+ D R+WD+
Sbjct: 430 NFSPNGYHLASGGEDNQCRIWDL 452
Score = 34.7 bits (78), Expect = 0.009
Identities = 15/59 (25%), Positives = 29/59 (48%)
Query: 24 DGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVW 82
+G LA+ +D KV +W +L H + + + +AT S+D+T+++W
Sbjct: 477 EGYFLATASYDMKVNIWSGRDFSLVKSLAGHESKVASLDITADSSCIATVSHDRTIKLW 535
Score = 32.7 bits (73), Expect = 0.032
Identities = 18/64 (28%), Positives = 27/64 (42%)
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVR 80
F GK L + +DK LW ++ + E HS + + F A+ D R
Sbjct: 347 FHPSGKYLGTTSYDKTWRLWDINTGAELLLQEGHSRSVYGIAFQQDGALAASCGLDSLAR 406
Query: 81 VWDV 84
VWD+
Sbjct: 407 VWDL 410
Score = 27.7 bits (60), Expect = 1.0
Identities = 14/54 (25%), Positives = 28/54 (50%), Gaps = 4/54 (7%)
Query: 3 FSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYT----DSLKQKATLE 52
FS + S+ +KV ++D +A+ HD+ + LW + D ++K T++
Sbjct: 498 FSLVKSLAGHESKVASLDITADSSCIATVSHDRTIKLWTSSGNDDEDEEKETMD 551
>At1g15850 hypothetical protein
Length = 140
Score = 45.1 bits (105), Expect = 6e-06
Identities = 22/65 (33%), Positives = 35/65 (53%)
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
V+CS + DG + +GG DK+ +W S Q +T+ H + + P M L T S+
Sbjct: 76 VLCSAWKDDGTTVFTGGCDKQAKMWPLLSGAQPSTVAMHDAPFNQIAWIPGMNLLVTGSW 135
Query: 76 DKTVR 80
DKT++
Sbjct: 136 DKTLK 140
Score = 25.4 bits (54), Expect = 5.2
Identities = 10/27 (37%), Positives = 16/27 (59%)
Query: 58 ITDVRFSPSMPRLATSSYDKTVRVWDV 84
I+ + FSP L +S+D VR W++
Sbjct: 29 ISSLSFSPKADILVATSWDCQVRCWEI 55
>At3g21540 WD repeat like protein
Length = 949
Score = 44.7 bits (104), Expect = 8e-06
Identities = 23/70 (32%), Positives = 35/70 (49%)
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
V ++ G +LASG D ++LW L H +TD+ F +L +SS
Sbjct: 109 VTALRYNKVGSMLASGSKDNDIILWDVVGESGLFRLRGHRDQVTDLVFLDGGKKLVSSSK 168
Query: 76 DKTVRVWDVD 85
DK +RVWD++
Sbjct: 169 DKFLRVWDLE 178
Score = 42.4 bits (98), Expect = 4e-05
Identities = 21/70 (30%), Positives = 35/70 (50%)
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
V+C SSDG+L+ +G DK + +W D ++ H + V+F + L +
Sbjct: 578 VMCIDISSDGELIVTGSQDKNLKIWGLDFGDCHKSIFAHGDSVMGVKFVRNTHYLFSIGK 637
Query: 76 DKTVRVWDVD 85
D+ V+ WD D
Sbjct: 638 DRLVKYWDAD 647
Score = 40.0 bits (92), Expect = 2e-04
Identities = 25/84 (29%), Positives = 42/84 (49%), Gaps = 1/84 (1%)
Query: 2 TFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDV 61
T S + S++ + + V+ S D K +A D V ++Y DSLK +L H L + +
Sbjct: 523 TVSNVKSMKMNDD-VLAVAISPDAKHIAVALLDSTVKVFYMDSLKFYLSLYGHKLPVMCI 581
Query: 62 RFSPSMPRLATSSYDKTVRVWDVD 85
S + T S DK +++W +D
Sbjct: 582 DISSDGELIVTGSQDKNLKIWGLD 605
Score = 36.6 bits (83), Expect = 0.002
Identities = 24/76 (31%), Positives = 35/76 (45%)
Query: 8 SVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSM 67
S+ A + V+ F + L S G D+ V W D + TLE H I + S
Sbjct: 612 SIFAHGDSVMGVKFVRNTHYLFSIGKDRLVKYWDADKFEHLLTLEGHHAEIWCLAISNRG 671
Query: 68 PRLATSSYDKTVRVWD 83
L T S+D+++R WD
Sbjct: 672 DFLVTGSHDRSMRRWD 687
Score = 25.4 bits (54), Expect = 5.2
Identities = 15/58 (25%), Positives = 27/58 (45%), Gaps = 4/58 (6%)
Query: 34 DKKVVLWYTDSLKQKATLEEHSLLITD----VRFSPSMPRLATSSYDKTVRVWDVDNV 87
D +V W + K+ S+ + D V SP +A + D TV+V+ +D++
Sbjct: 508 DHEVKFWEYQATKKLTVSNVKSMKMNDDVLAVAISPDAKHIAVALLDSTVKVFYMDSL 565
>At2g01330 putative WD-40-repeat protein
Length = 611
Score = 44.7 bits (104), Expect = 8e-06
Identities = 24/71 (33%), Positives = 39/71 (54%), Gaps = 3/71 (4%)
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLWYT---DSLKQKATLEEHSLLITDVRFSPSMPRLAT 72
V S S DGK GG D K+ ++ ++LK++A LE+H +T +R+SP + A+
Sbjct: 451 VAASVISPDGKEAIVGGQDGKLHIYSVSGDNNLKEEAVLEKHRGALTVIRYSPDLTMFAS 510
Query: 73 SSYDKTVRVWD 83
++ VWD
Sbjct: 511 GDANREAVVWD 521
Score = 41.2 bits (95), Expect = 9e-05
Identities = 26/81 (32%), Positives = 42/81 (51%), Gaps = 3/81 (3%)
Query: 7 NSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLW---YTDSLKQKATLEEHSLLITDVRF 63
+S R +N V C +S DG + DKK +++ D + + A+ + H I V +
Sbjct: 182 SSHREHSNFVNCIRYSPDGTKFITVSSDKKGMIYDGKTGDKVGELASEDGHKGSIYAVSW 241
Query: 64 SPSMPRLATSSYDKTVRVWDV 84
SP R+ T S DK+ +VW+V
Sbjct: 242 SPDSKRVLTVSADKSAKVWEV 262
Score = 35.0 bits (79), Expect = 0.007
Identities = 20/66 (30%), Positives = 37/66 (55%), Gaps = 1/66 (1%)
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKAT-LEEHSLLITDVRFSPSMPRLATSSYDKTV 79
+S D + ASG +++ V+W ++ + K + H+ I + +SP+ +AT S D V
Sbjct: 501 YSPDLTMFASGDANREAVVWDRETKQVKLNNMLFHTARINSLAWSPNNKMVATGSIDTCV 560
Query: 80 RVWDVD 85
V++VD
Sbjct: 561 IVYEVD 566
Score = 31.6 bits (70), Expect = 0.072
Identities = 20/69 (28%), Positives = 33/69 (46%), Gaps = 9/69 (13%)
Query: 27 LLASGGHDKKVVLW------YTDSLKQKATLE---EHSLLITDVRFSPSMPRLATSSYDK 77
+L SG +L+ + SL+Q ++ EH +T R+SP+ +A++
Sbjct: 20 ILISGDSKSDTILYCNGRSVFIRSLRQLQDVQVYGEHGYAVTVARYSPNGEWIASADVSG 79
Query: 78 TVRVWDVDN 86
TVRVW N
Sbjct: 80 TVRVWGTHN 88
Score = 28.5 bits (62), Expect = 0.61
Identities = 16/56 (28%), Positives = 27/56 (47%)
Query: 28 LASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWD 83
+A+ G D V + K ++ EHS + +R+SP + T S DK ++D
Sbjct: 161 IATCGEDFLVNFYDGPPFKFHSSHREHSNFVNCIRYSPDGTKFITVSSDKKGMIYD 216
Score = 27.3 bits (59), Expect = 1.4
Identities = 19/83 (22%), Positives = 41/83 (48%), Gaps = 3/83 (3%)
Query: 5 ELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLL--ITDVR 62
+LN++ T ++ +S + K++A+G D V+++ D ++ L + V
Sbjct: 528 KLNNMLFHTARINSLAWSPNNKMVATGSIDTCVIVYEVDKPASSRITARNAHLGGVNAVA 587
Query: 63 FSPSMPRLATSSYDKTVRVWDVD 85
F +A+S D +VR+W ++
Sbjct: 588 FIDDC-TVASSGEDASVRLWHIE 609
>At1g73720 unknown protein
Length = 511
Score = 44.7 bits (104), Expect = 8e-06
Identities = 22/66 (33%), Positives = 37/66 (55%)
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
V+C FS D ++LASG D K+ +W + + HS +T + FS +L ++S+
Sbjct: 266 VLCIDFSRDSEMLASGSQDGKIKIWRIRTGVCIRRFDAHSQGVTSLSFSRDGSQLLSTSF 325
Query: 76 DKTVRV 81
D+T R+
Sbjct: 326 DQTARI 331
Score = 36.6 bits (83), Expect = 0.002
Identities = 23/75 (30%), Positives = 37/75 (48%), Gaps = 8/75 (10%)
Query: 18 CSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLE---EHSLLITD-----VRFSPSMPR 69
C+ FS DG+ LAS D + +W S K K L+ + S ++ D + FS
Sbjct: 218 CARFSPDGQFLASSSVDGFIEVWDYISGKLKKDLQYQADESFMMHDDPVLCIDFSRDSEM 277
Query: 70 LATSSYDKTVRVWDV 84
LA+ S D +++W +
Sbjct: 278 LASGSQDGKIKIWRI 292
Score = 33.9 bits (76), Expect = 0.015
Identities = 22/73 (30%), Positives = 31/73 (42%)
Query: 11 ASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRL 70
A + V FS DG L S D+ + S K H+ + F+ R+
Sbjct: 303 AHSQGVTSLSFSRDGSQLLSTSFDQTARIHGLKSGKLLKEFRGHTSYVNHAIFTSDGSRI 362
Query: 71 ATSSYDKTVRVWD 83
T+S D TV+VWD
Sbjct: 363 ITASSDCTVKVWD 375
Score = 24.6 bits (52), Expect = 8.8
Identities = 17/66 (25%), Positives = 29/66 (43%)
Query: 17 VCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYD 76
V + S+ G + G DKK+ + S + + H + + P LAT S D
Sbjct: 444 VAACVSTKGDWIYCIGEDKKLYCFNYQSGGLEHFMMVHEKDVIGITHHPHRNLLATYSED 503
Query: 77 KTVRVW 82
T+++W
Sbjct: 504 CTMKLW 509
>At4g05410 U3 snoRNP-associated -like protein
Length = 504
Score = 44.3 bits (103), Expect = 1e-05
Identities = 21/65 (32%), Positives = 37/65 (56%)
Query: 22 SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRV 81
SSDG+ LA+GG D+ V +W + + H ++ + F L + S+D+TV+V
Sbjct: 231 SSDGRYLATGGVDRHVHIWDVRTREHVQAFPGHRNTVSCLCFRYGTSELYSGSFDRTVKV 290
Query: 82 WDVDN 86
W+V++
Sbjct: 291 WNVED 295
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.128 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,893,060
Number of Sequences: 26719
Number of extensions: 62241
Number of successful extensions: 795
Number of sequences better than 10.0: 178
Number of HSP's better than 10.0 without gapping: 149
Number of HSP's successfully gapped in prelim test: 29
Number of HSP's that attempted gapping in prelim test: 242
Number of HSP's gapped (non-prelim): 518
length of query: 88
length of database: 11,318,596
effective HSP length: 64
effective length of query: 24
effective length of database: 9,608,580
effective search space: 230605920
effective search space used: 230605920
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146722.4


