

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146712.3 - phase: 0 /pseudo
(276 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
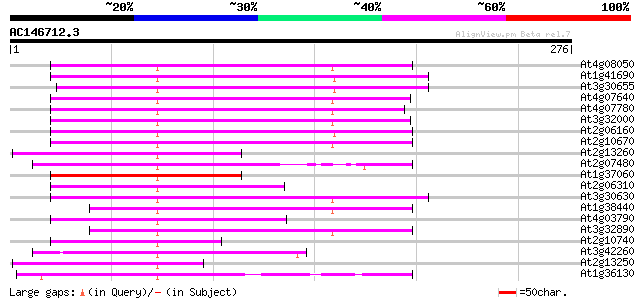
Score E
Sequences producing significant alignments: (bits) Value
At4g08050 104 5e-23
At1g41690 hypothetical protein 104 5e-23
At3g30655 hypothetical protein, 3' partial 100 1e-21
At4g07640 putative athila transposon protein 98 6e-21
At4g07780 putative athila transposon protein 96 3e-20
At3g32000 unknown protein 95 5e-20
At2g06160 putative Athila retroelement ORF1 protein 95 5e-20
At2g10670 pseudogene 92 3e-19
At2g13260 putative Athila retroelement ORF1 protein 91 5e-19
At2g07480 F9A16.15 91 5e-19
At1g37060 Athila retroelment ORF 1, putative 91 5e-19
At2g06310 putative Athila retroelement ORF1 protein 89 3e-18
At3g30630 hypothetical protein 87 1e-17
At1g38440 hypothetical protein 82 4e-16
At4g03790 putative athila-like protein 81 7e-16
At3g32890 Athila ORF 1, putative 79 2e-15
At2g10740 pseudogene 75 5e-14
At3g42260 putative protein 73 2e-13
At2g13250 putative Athila retroelement ORF1 protein 70 2e-12
At1g36130 hypothetical protein 69 3e-12
>At4g08050
Length = 1428
Score = 104 bits (260), Expect = 5e-23
Identities = 61/182 (33%), Positives = 99/182 (53%), Gaps = 4/182 (2%)
Query: 21 IVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*L 77
I P ++ NNFEIK L++++Q N+F G P EDP H+ F RL K N ++ L
Sbjct: 29 IAPPAIQNNNFEIKSGLISMIQGNKFHGLPMEDPLDHLDEFDRLCNLTKINGVSEDGFKL 88
Query: 78 HLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMGSL 137
LFPFSL D+A W +L SIT+WD ++AFL++F + + L + +G +K G
Sbjct: 89 RLFPFSLGDKAHIWEKNLSHDSITTWDDYKKAFLSKFFSNARTARLRNEIYGFSQKTGES 148
Query: 138 SMKRGRVSRKSLDFVPTMAW-KSGL*STPFNGLSYTTKMSVDAAAGGSLMNKNYTEAYAL 196
+ + + P ++ K+ L ST + G+ +M +D A+ G+ NK+ E + L
Sbjct: 149 FCEAWERFKGYTNQCPHHSFTKASLLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWEL 208
Query: 197 MK 198
++
Sbjct: 209 VE 210
>At1g41690 hypothetical protein
Length = 371
Score = 104 bits (260), Expect = 5e-23
Identities = 65/190 (34%), Positives = 100/190 (52%), Gaps = 4/190 (2%)
Query: 21 IVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*L 77
IV P V+ NNFEIK +L+ +VQ N+F G P EDP H+ F RL K N ++ L
Sbjct: 29 IVPPPVQNNNFEIKSSLIAMVQGNKFHGLPMEDPLDHLDEFERLCSLTKINGVSEDGFKL 88
Query: 78 HLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMGSL 137
LFPFSL D+A W +L GSIT+WD ++AFLA+F + + L + G +K
Sbjct: 89 RLFPFSLGDKAHLWEKTLPQGSITTWDDCKKAFLAKFFSYSRTARLRNEISGFTQKQSES 148
Query: 138 SMKRGRVSRKSLDFVPTMAWK-SGL*STPFNGLSYTTKMSVDAAAGGSLMNKNYTEAYAL 196
+ + P +K + L ST + G+ +M +D A+ G+ + K+ E + L
Sbjct: 149 FCEAWERFKGYQTKCPHHGFKQASLLSTLYRGVLPKIRMLLDTASNGNFLKKDVEEGWEL 208
Query: 197 MKTWLRITTN 206
++ + + N
Sbjct: 209 VEKFAQSDGN 218
>At3g30655 hypothetical protein, 3' partial
Length = 660
Score = 99.8 bits (247), Expect = 1e-21
Identities = 62/187 (33%), Positives = 97/187 (51%), Gaps = 4/187 (2%)
Query: 24 PTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*LHLF 80
P V+ NNFEIK L+ +VQ N+F G P EDP H+ F RL G K N ++ L LF
Sbjct: 32 PPVQNNNFEIKSGLIAMVQGNKFHGLPMEDPLDHLDEFERLYGLTKINGVSEDGFKLRLF 91
Query: 81 PFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMGSLSMK 140
PFSL D+A W +L SIT+WD ++AFLA+F + + L + G +K +
Sbjct: 92 PFSLGDKAHLWEKTLPQNSITAWDDCKKAFLAKFFSNSRTARLRNEISGFTQKQNESFGE 151
Query: 141 RGRVSRKSLDFVPTMAWK-SGL*STPFNGLSYTTKMSVDAAAGGSLMNKNYTEAYALMKT 199
+ P +K + L ST + G+ ++ +D A+ G+ + K+ E + L++
Sbjct: 152 AWERFKGYQTKCPHHGFKQASLLSTLYRGVLPKIRILLDTASNGNFLKKDVEEGWELVEN 211
Query: 200 WLRITTN 206
+ + N
Sbjct: 212 FAQSDGN 218
>At4g07640 putative athila transposon protein
Length = 866
Score = 97.8 bits (242), Expect = 6e-21
Identities = 59/181 (32%), Positives = 94/181 (51%), Gaps = 4/181 (2%)
Query: 21 IVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*L 77
I P ++ NNFEIK L++++Q N+F G P EDP H+ F RL K N + L
Sbjct: 29 IAPPAIQNNNFEIKSGLISMIQGNKFYGLPMEDPLDHLDEFDRLCNLTKINGVSADGFKL 88
Query: 78 HLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMGSL 137
LFPFSL D+A W +L SI +WD ++AFL++F + + L + G +K G
Sbjct: 89 RLFPFSLGDKAHIWEKNLPHDSIITWDDCKKAFLSKFFSNARTARLRNEISGFSQKTGES 148
Query: 138 SMKRGRVSRKSLDFVPTMAW-KSGL*STPFNGLSYTTKMSVDAAAGGSLMNKNYTEAYAL 196
+ + + P + K+ + ST + G+ +M +D A+ G+ NK+ E + L
Sbjct: 149 FCEAWERFKGYTNQCPHHGFTKASMLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWEL 208
Query: 197 M 197
+
Sbjct: 209 V 209
>At4g07780 putative athila transposon protein
Length = 446
Score = 95.5 bits (236), Expect = 3e-20
Identities = 59/178 (33%), Positives = 91/178 (50%), Gaps = 4/178 (2%)
Query: 21 IVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*L 77
IV P V+ NNFEI L++++Q N+F G P EDP ++ SF RL G K N ++ L
Sbjct: 37 IVPPPVQINNFEIMSGLISMIQGNKFHGLPKEDPLDNLDSFDRLCGLTKINGVTKDMFKL 96
Query: 78 HLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMGSL 137
LFPFSL D+A W +L SITSWD ++ FLA+F + + L + G +K
Sbjct: 97 RLFPFSLGDKAHHWKKTLPPDSITSWDDCKKDFLAKFFSNARTARLRNEISGFTQKNNET 156
Query: 138 SMKRGRVSRKSLDFVPTMAW-KSGL*STPFNGLSYTTKMSVDAAAGGSLMNKNYTEAY 194
+ + + P + K+ L T + G +M +D + G+ +N + E +
Sbjct: 157 FFEASERFKSYTTYCPHHGFKKASLLRTLYRGALPKIRMLLDTTSNGNFLNNDVAEGW 214
>At3g32000 unknown protein
Length = 839
Score = 94.7 bits (234), Expect = 5e-20
Identities = 60/181 (33%), Positives = 95/181 (52%), Gaps = 4/181 (2%)
Query: 21 IVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*L 77
I P ++ NNFEIK L++++Q N+F G P EDP H+ F RL K N + L
Sbjct: 29 IAPPAIQKNNFEIKSGLISMIQGNKFHGLPMEDPLDHLDEFDRLCNLTKINGVSENGFKL 88
Query: 78 HLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMGSL 137
LFPFSL D+A W +L SIT+ D ++AFL++F + + L + G +K+G
Sbjct: 89 RLFPFSLGDKALIWEMNLPHDSITTRDDCKKAFLSKFFSNDRTARLRNEISGFSQKIGES 148
Query: 138 SMKRGRVSRKSLDFVPTMAW-KSGL*STPFNGLSYTTKMSVDAAAGGSLMNKNYTEAYAL 196
+ + + P + K+ L ST + G+ +M +D A+ G+ NK+ E + L
Sbjct: 149 FCEAWERFKDYTNQCPHHGFTKASLLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWEL 208
Query: 197 M 197
+
Sbjct: 209 V 209
>At2g06160 putative Athila retroelement ORF1 protein
Length = 750
Score = 94.7 bits (234), Expect = 5e-20
Identities = 59/182 (32%), Positives = 92/182 (50%), Gaps = 4/182 (2%)
Query: 21 IVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*L 77
IV P V+ NNFEIK L+ +VQ N+F G P ED H+ F RL K N ++
Sbjct: 29 IVLPLVQNNNFEIKSGLIAMVQGNKFHGLPMEDSLDHLDEFERLCDLTKINGVSEDGFKF 88
Query: 78 HLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMGSL 137
LFPFSL D+ W +L SIT+WD ++AF A+F + + L + G +K
Sbjct: 89 RLFPFSLGDKTHLWEKTLPQNSITTWDDCKKAFFAKFFSNSRTARLRNEISGFTQKQNES 148
Query: 138 SMKRGRVSRKSLDFVPTMAWK-SGL*STPFNGLSYTTKMSVDAAAGGSLMNKNYTEAYAL 196
+ + P +K + L ST + G+ +M +D A+ G+ +NK+ + + L
Sbjct: 149 FCEAWERFKGYQTKCPHPGFKQASLLSTLYRGVLPKLRMLLDTASNGNFLNKDVEKGWEL 208
Query: 197 MK 198
++
Sbjct: 209 VE 210
>At2g10670 pseudogene
Length = 929
Score = 92.0 bits (227), Expect = 3e-19
Identities = 59/182 (32%), Positives = 92/182 (50%), Gaps = 4/182 (2%)
Query: 21 IVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*L 77
IV V+ NNF+IK L+ +VQ N+F G P EDP H+ RL G K N ++ L
Sbjct: 61 IVLRPVQNNNFKIKSGLIAMVQGNKFHGLPMEDPLDHLDELERLCGLTKINGVSEDGFKL 120
Query: 78 HLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMGSL 137
LFPFSL D+A W +L SIT+WD + AFLA+F + + L + G K
Sbjct: 121 RLFPFSLGDKAHLWEKTLPKNSITTWDDCKMAFLAKFFSNSRTARLRNEISGFTLKQNES 180
Query: 138 SMKRGRVSRKSLDFVPTMAW-KSGL*STPFNGLSYTTKMSVDAAAGGSLMNKNYTEAYAL 196
+ + P + ++ L +T + G+ +M +D A+ G+ + K+ E + L
Sbjct: 181 FCEAWERFKGYQTKCPHHGFSQASLLNTLYRGVLPKIRMLLDTASNGNFLKKDIEEGWEL 240
Query: 197 MK 198
++
Sbjct: 241 VE 242
>At2g13260 putative Athila retroelement ORF1 protein
Length = 622
Score = 91.3 bits (225), Expect = 5e-19
Identities = 51/116 (43%), Positives = 70/116 (59%), Gaps = 3/116 (2%)
Query: 2 AERTLKEYATPSMEEPQAIIVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSF 61
A + + + P++ + I P VE NNFEIK +L+N+VQ ++F G EDP H+ F
Sbjct: 28 AHQPIGAFDEPNIRGNRNGIQAPPVENNNFEIKSSLINMVQTSKFHGLSMEDPLDHLEQF 87
Query: 62 LRLSGTIKEN---QEAV*LHLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARF 114
L T+K N ++A L LFPFSL DRA W +L SITSWD +RAFL++F
Sbjct: 88 DMLCSTVKINGISEDAFKLRLFPFSLGDRARIWEKNLPQRSITSWDQCKRAFLSKF 143
>At2g07480 F9A16.15
Length = 1012
Score = 91.3 bits (225), Expect = 5e-19
Identities = 67/211 (31%), Positives = 102/211 (47%), Gaps = 47/211 (22%)
Query: 12 PSMEEPQAIIVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN 71
P +A IV P ++ NNF+IK L++++Q N+F G P EDP H+ +F RL K N
Sbjct: 15 PRNHHQRAGIVPPPIQNNNFKIKSDLISMIQGNKFHGLPMEDPLDHLDNFDRLCSLTKIN 74
Query: 72 ---QEAV*LHLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSH 128
+E+ L LFPFSL D+A W +L V SI +WD ++AFLA+F + + L +
Sbjct: 75 GVSEESFKLRLFPFSLGDKAHLWEKTLPVKSIDTWDDCKKAFLAKFFSNSRKARLRSEIS 134
Query: 129 GLIKKMGSLSMKRGRVSRKSLDFVPTMAWKSGL*STPFNGLSYTT--------------- 173
G +K S F + AW+ F G YTT
Sbjct: 135 GFNQK-------------NSESF--SEAWER------FKG--YTTQCPHHESLPPQYSIL 171
Query: 174 ------KMSVDAAAGGSLMNKNYTEAYALMK 198
+M +D A+ G+++NK+ E + L++
Sbjct: 172 RCLPKIRMLLDTASNGNILNKDVAEGWELVE 202
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 91.3 bits (225), Expect = 5e-19
Identities = 49/97 (50%), Positives = 61/97 (62%), Gaps = 3/97 (3%)
Query: 21 IVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*L 77
IV P V+ NNFEIK L+ +VQ N+F G P EDP H+ F RL K N ++ L
Sbjct: 29 IVPPPVQNNNFEIKSGLIAMVQSNKFHGLPMEDPLDHLDEFDRLCSLTKINRVSEDGFKL 88
Query: 78 HLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARF 114
LFPFSL D+A W SL GSITSW+ ++AFLA+F
Sbjct: 89 RLFPFSLGDKAHQWEKSLPQGSITSWNDCKKAFLAKF 125
>At2g06310 putative Athila retroelement ORF1 protein
Length = 733
Score = 88.6 bits (218), Expect = 3e-18
Identities = 46/118 (38%), Positives = 68/118 (56%), Gaps = 3/118 (2%)
Query: 21 IVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*L 77
I P ++ NNFEIK L++++Q N+F G P EDP H+ F RL K N ++ L
Sbjct: 29 IAPPVIQNNNFEIKSGLISMIQGNKFHGLPMEDPLNHLDEFDRLCNLTKINGVIEDGFNL 88
Query: 78 HLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMG 135
LFPFSL +A W +L SIT+WD ++AFL++F + + L + G ++K G
Sbjct: 89 RLFPFSLGGKAHIWEKNLPHDSITTWDDCKKAFLSKFFSNARTARLRNEISGFLQKAG 146
>At3g30630 hypothetical protein
Length = 785
Score = 87.0 bits (214), Expect = 1e-17
Identities = 57/190 (30%), Positives = 94/190 (49%), Gaps = 4/190 (2%)
Query: 21 IVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*L 77
I PT++ NNF+IK L++++Q N+F G P ED H+ F RL K N ++ L
Sbjct: 29 IAPPTIQNNNFKIKSGLISMIQGNKFHGLPMEDLLDHLDEFDRLCNLTKVNGVSEDGFKL 88
Query: 78 HLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMGSL 137
LFPF L D+A W ++ SI +W ++AFLA+F + K L + +K G
Sbjct: 89 RLFPFFLGDKAHIWEKNMPHDSIITWVDCKKAFLAKFFSNAKTARLINEISSFSQKTGES 148
Query: 138 SMKRGRVSRKSLDFVPTMAW-KSGL*STPFNGLSYTTKMSVDAAAGGSLMNKNYTEAYAL 196
+ + + P + K+ L ST + G+ +M +D A+ + NK+ E + L
Sbjct: 149 FCEAWERFKGYTNQCPHHGFKKASLLSTLYRGVLPRIRMLLDTASNENFQNKDVEEGWEL 208
Query: 197 MKTWLRITTN 206
++ + N
Sbjct: 209 IENLAQSNDN 218
>At1g38440 hypothetical protein
Length = 263
Score = 81.6 bits (200), Expect = 4e-16
Identities = 52/163 (31%), Positives = 84/163 (50%), Gaps = 4/163 (2%)
Query: 40 LVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*LHLFPFSLRDRASAWFHSLE 96
++Q+N+F G P EDP H+ F RL K N ++ L LFPFSL D+A W +L
Sbjct: 1 MIQENKFHGLPMEDPLDHLDEFDRLYNLTKINGVSEDGFKLRLFPFSLGDKAHIWEKNLP 60
Query: 97 VGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMGSLSMKRGRVSRKSLDFVPTMA 156
SIT+WD ++AFL++F + + L + G +K + + + P
Sbjct: 61 HDSITTWDDCKKAFLSKFFSNARTARLRNEISGFSQKTSESFCEAWERFKGYTNQCPHHG 120
Query: 157 W-KSGL*STPFNGLSYTTKMSVDAAAGGSLMNKNYTEAYALMK 198
+ K+ L ST + G+ M +DAA+ G+ NK+ E + L++
Sbjct: 121 FTKASLLSTLYRGVLPCIIMLLDAASNGNFQNKDVEEGWELVE 163
>At4g03790 putative athila-like protein
Length = 1064
Score = 80.9 bits (198), Expect = 7e-16
Identities = 45/119 (37%), Positives = 66/119 (54%), Gaps = 3/119 (2%)
Query: 21 IVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*L 77
I P ++ NNFEIK L++++Q N+F G P EDP H+ F RL K N ++ L
Sbjct: 29 IAPPAIQNNNFEIKSGLISIIQGNKFHGLPMEDPLDHLDEFDRLCNLTKINGVSEDGFKL 88
Query: 78 HLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMGS 136
LFPFSL D+A W +L SIT+W RR F SP + + E++ +++G+
Sbjct: 89 RLFPFSLGDKAHIWEKNLSHDSITTWMIARRLFYQSSSPMPELQDSEMRCLAFYRRLGN 147
>At3g32890 Athila ORF 1, putative
Length = 755
Score = 79.3 bits (194), Expect = 2e-15
Identities = 50/163 (30%), Positives = 83/163 (50%), Gaps = 4/163 (2%)
Query: 40 LVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*LHLFPFSLRDRASAWFHSLE 96
++Q N+F G P EDP H+ F RL K N ++ L LFPFSL D+A W +L
Sbjct: 1 MIQGNKFHGLPMEDPLDHLDEFDRLYNLTKINGVSEDGFKLRLFPFSLGDKAHIWEKNLP 60
Query: 97 VGSITSWDGMRRAFLARFSPHLKPLNLEIKSHGLIKKMGSLSMKRGRVSRKSLDFVPTMA 156
SIT+WD ++AFL++F + + L + G +K + + + P
Sbjct: 61 HDSITTWDDCKKAFLSKFFSNARTARLRNEISGFSQKTRESFCEAWERFKGYTNQCPHHG 120
Query: 157 W-KSGL*STPFNGLSYTTKMSVDAAAGGSLMNKNYTEAYALMK 198
+ K+ L ST + G+ +M +D ++ G+ NK+ E + L++
Sbjct: 121 FTKASLLSTLYRGVLLRIRMLLDTSSNGNFQNKDVEEGWELVE 163
>At2g10740 pseudogene
Length = 169
Score = 74.7 bits (182), Expect = 5e-14
Identities = 40/87 (45%), Positives = 53/87 (59%), Gaps = 3/87 (3%)
Query: 21 IVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN---QEAV*L 77
IV P V+ NNFEIK L++++Q N+F G P +DP H+ SF RL G N ++
Sbjct: 81 IVPPPVQNNNFEIKSGLISMIQGNKFHGLPMKDPLDHLDSFDRLCGLTNINGVTEDMFKQ 140
Query: 78 HLFPFSLRDRASAWFHSLEVGSITSWD 104
LFPFS+ D+A W +L SITSWD
Sbjct: 141 RLFPFSVGDKAHHWEKTLPSDSITSWD 167
>At3g42260 putative protein
Length = 355
Score = 72.8 bits (177), Expect = 2e-13
Identities = 49/140 (35%), Positives = 74/140 (52%), Gaps = 6/140 (4%)
Query: 12 PSMEEPQAIIVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSGTIKEN 71
P++++P+ I +G + K L++ +Q N+F G P EDP H+ SF RL G K N
Sbjct: 201 PNVDQPRNIGACDA-QGIITKDKGGLISTIQGNKFHGLPMEDPLDHLDSFDRLCGFTKIN 259
Query: 72 ---QEAV*LHLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPLNLEIKSH 128
++ L LFPFSL D+A W +L SI SW ++ FLA+F + + +
Sbjct: 260 GVTEDMFKLRLFPFSLGDKAHHWEKTLPPKSINSWGDCKKTFLAKFFSNARTTRFRNEIS 319
Query: 129 GLIKKMGSLSMK--RGRVSR 146
G +K S+K RG +SR
Sbjct: 320 GFTQKNYETSVKFARGDMSR 339
>At2g13250 putative Athila retroelement ORF1 protein
Length = 507
Score = 69.7 bits (169), Expect = 2e-12
Identities = 40/97 (41%), Positives = 56/97 (57%), Gaps = 3/97 (3%)
Query: 2 AERTLKEYATPSMEEPQAIIVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSF 61
A + + + P++ + I P VE NNFEIK +L+N+VQ ++F G EDP H+ F
Sbjct: 101 AHQPIGAFDEPNIRGNRNGIQAPPVENNNFEIKSSLINMVQTSKFHGLSMEDPLDHLDQF 160
Query: 62 LRLSGTIKEN---QEAV*LHLFPFSLRDRASAWFHSL 95
L T+K N ++A L LFPFSL DRA W +L
Sbjct: 161 DMLCSTVKINGISEDAFKLRLFPFSLGDRARIWEKNL 197
>At1g36130 hypothetical protein
Length = 914
Score = 68.9 bits (167), Expect = 3e-12
Identities = 55/199 (27%), Positives = 94/199 (46%), Gaps = 19/199 (9%)
Query: 4 RTLKEYATPSM-EEPQAIIVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFL 62
RTL ++ P + ++ IV P ++ N++E+KP LV Q+ F G E P HI F
Sbjct: 73 RTLGDFNRPDLFYANRSAIVPPPLQRNDYELKPGYFALVGQHPFHGLSHEQPMDHIERFE 132
Query: 63 RLSGTIKEN---QEAV*LHLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLK 119
L +IK N ++ + LFP+SL A +W L+ GS+ +W ++ AFL F
Sbjct: 133 DLVLSIKANGVSEDYLLCKLFPYSLAGEADSWLKQLKAGSLKTWRSIKIAFLNNFYD--- 189
Query: 120 PLNLEIKSHGLIKKMGSLSMKRGRVSRKSLDFVPTMAWKSGL*STPFNGLSYTTKMSVDA 179
+ KS L K+ + + + + + +K P +G S +M++DA
Sbjct: 190 ----DAKSEELRNKLSTFTQGPAEAFKAA-----WVRFKEYQRDCPHHGFS---EMALDA 237
Query: 180 AAGGSLMNKNYTEAYALMK 198
A+ G+ +A L++
Sbjct: 238 ASNGNFNTHYPADATTLIE 256
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.138 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,801,776
Number of Sequences: 26719
Number of extensions: 233213
Number of successful extensions: 646
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 42
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 570
Number of HSP's gapped (non-prelim): 56
length of query: 276
length of database: 11,318,596
effective HSP length: 98
effective length of query: 178
effective length of database: 8,700,134
effective search space: 1548623852
effective search space used: 1548623852
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146712.3


