

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146706.4 + phase: 0
(478 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
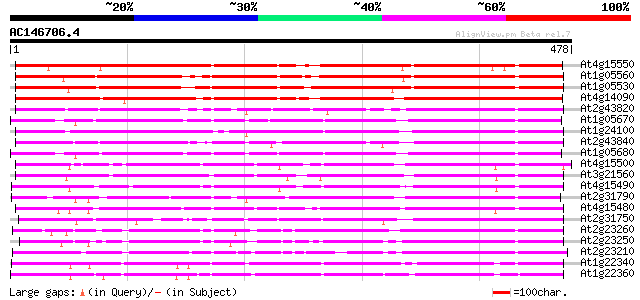
Score E
Sequences producing significant alignments: (bits) Value
At4g15550 UDP-glucose:indole-3-acetate beta-D-glucosyltransferas... 446 e-125
At1g05560 UDP-Glucose Transferase (UGT75B2) 411 e-115
At1g05530 hypothetical protein 405 e-113
At4g14090 glucosyltransferase like protein 402 e-112
At2g43820 putative glucosyltransferase 310 1e-84
At1g05670 UDP glycosyltransferase UGT74E1 292 3e-79
At1g24100 putative indole-3-acetate beta-glucosyltransferase 291 7e-79
At2g43840 putative glucosyltransferase 289 3e-78
At1g05680 indole-3-acetate beta-glucosyltransferase like protein 287 1e-77
At4g15500 indole-3-acetate beta-glucosyltransferase like protein 284 7e-77
At3g21560 UDP-glucose:indole-3-acetate beta-D-glucosyltransferas... 284 7e-77
At4g15490 indole-3-acetate beta-glucosyltransferase like protein 282 3e-76
At2g31790 putative glucosyltransferase 279 2e-75
At4g15480 indole-3-acetate beta-glucosyltransferase like protein 268 5e-72
At2g31750 putative glucosyltransferase 268 6e-72
At2g23260 putative glucosyltransferase 254 7e-68
At2g23250 putative glucosyltransferase 246 3e-65
At2g23210 putative glucosyltransferase 229 2e-60
At1g22340 hypothetical protein 212 3e-55
At1g22360 unknown protein 209 2e-54
>At4g15550 UDP-glucose:indole-3-acetate beta-D-glucosyltransferase
(iaglu)
Length = 474
Score = 446 bits (1146), Expect = e-125
Identities = 236/482 (48%), Positives = 329/482 (67%), Gaps = 38/482 (7%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISL--GAKVTFATTIHLYSR-LINKPTIPG-LSFATF 61
HFL +T+P QGHINP+L+ KRL GA+VTFA +I Y+R + + +P L FAT+
Sbjct: 13 HFLFVTFPAQGHINPSLELAKRLAGTISGARVTFAASISAYNRRMFSTENVPETLIFATY 72
Query: 62 SDGYDDGQKSFGDED------IVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLIL 115
SDG+DDG KS D ++MSE RRG E LT +I ++++N PFTC++YT++L
Sbjct: 73 SDGHDDGFKSSAYSDKSRQDATGNFMSEMRRRGKETLTELIEDNRKQNRPFTCVVYTILL 132
Query: 116 SWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLK 175
+W ++A E HLPS LLW+Q TVF IFY+YF+ + D I+ + + I LP L L
Sbjct: 133 TWVAELAREFHLPSALLWVQPVTVFSIFYHYFNGYEDAISEMANTPSSSIKLPSLPL-LT 191
Query: 176 SRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMI 235
RD+PSF+++SN Y F LP+ +EQI L EEINP++L+NT +E E +A++ V K++
Sbjct: 192 VRDIPSFIVSSNVYAFLLPAFREQIDSLKEEINPKILINTFQELEPEAMSSVP-DNFKIV 250
Query: 236 PIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKR 295
P+GPL+ TD S S+ +YI+WLD+K + SV+YVSFGTLAVLSK+
Sbjct: 251 PVGPLLTLR-------TDFS---------SRGEYIEWLDTKADSSVLYVSFGTLAVLSKK 294
Query: 296 QMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDEL--SCREELENNMNGKIVKWCSQ 353
Q+ E+ +AL+ S FLWVI DK + +++E+ +++ S REEL+ G +V WC Q
Sbjct: 295 QLVELCKALIQSRRPFLWVITDKSYRNKEDEQEKEEDCISSFREELDEI--GMVVSWCDQ 352
Query: 354 VEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRM--E 411
VL+HRS+GCF+THCGWNSTLESL SGVP+VAFPQW DQ NAKL+ED WKTG+R+ +
Sbjct: 353 FRVLNHRSIGCFVTHCGWNSTLESLVSGVPVVAFPQWNDQMMNAKLLEDCWKTGVRVMEK 412
Query: 412 HDEEGMVKV--EEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYL 469
+EEG+V V EEIR+C+E VM +K EE R NA +WKDLA AV+EGGSS +L++++
Sbjct: 413 KEEEGVVVVDSEEIRRCIEEVM--EDKAEEFRGNATRWKDLAAEAVREGGSSFNHLKAFV 470
Query: 470 ND 471
++
Sbjct: 471 DE 472
>At1g05560 UDP-Glucose Transferase (UGT75B2)
Length = 469
Score = 411 bits (1057), Expect = e-115
Identities = 228/474 (48%), Positives = 308/474 (64%), Gaps = 34/474 (7%)
Query: 6 HFLIITYPLQGHINPALQFTKRLIS-LGAKVTFATTIHLY--SRLINKPTIPGLSFATFS 62
HFL++T+P QGH+NP+L+F +RLI GA+VTF T + ++ S + N + LSF TFS
Sbjct: 5 HFLLVTFPAQGHVNPSLRFARRLIKRTGARVTFVTCVSVFHNSMIANHNKVENLSFLTFS 64
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVA 122
DG+DDG S ED G + L++ I ++K + P TCLIYT++L+WAPKVA
Sbjct: 65 DGFDDGGISTY-EDRQKRSVNLKVNGDKALSDFIEATKNGDSPVTCLIYTILLNWAPKVA 123
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
LPS LLWIQ A VF+I+Y +F + NKS + LP LS SL+ RDLPSF
Sbjct: 124 RRFQLPSALLWIQPALVFNIYYTHF------MGNKS-----VFELPNLS-SLEIRDLPSF 171
Query: 183 LLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIP 242
L SNT A + +E ++ L +E P++L+NT + E +AL I M+ +GPL+P
Sbjct: 172 LTPSNTNKGAYDAFQEMMEFLIKETKPKILINTFDSLEPEALTAFP--NIDMVAVGPLLP 229
Query: 243 SAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIAR 302
+ G T+ S D Y WLDSK E SV+YVSFGT+ LSK+Q+EE+AR
Sbjct: 230 TEIFSGS--TNKSVK------DQSSSYTLWLDSKTESSVIYVSFGTMVELSKKQIEELAR 281
Query: 303 ALLDSGFSFLWVIRDKKLQQQKEEEVDDDELS----CREELENNMNGKIVKWCSQVEVLS 358
AL++ FLWVI DK ++ K E ++ E+ R ELE G IV WCSQ+EVLS
Sbjct: 282 ALIEGKRPFLWVITDKSNRETKTEGEEETEIEKIAGFRHELEEV--GMIVSWCSQIEVLS 339
Query: 359 HRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMV 418
HR++GCF+THCGW+STLESL GVP+VAFP W+DQ TNAKL+E+ WKTG+R+ +++G+V
Sbjct: 340 HRAVGCFVTHCGWSSTLESLVLGVPVVAFPMWSDQPTNAKLLEESWKTGVRVRENKDGLV 399
Query: 419 KVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
+ EIR+CLE VM EK ELR NAKKWK LA A +EGGSS++N+ +++ DI
Sbjct: 400 ERGEIRRCLEAVM--EEKSVELRENAKKWKRLAMEAGREGGSSDKNMEAFVEDI 451
>At1g05530 hypothetical protein
Length = 455
Score = 405 bits (1042), Expect = e-113
Identities = 220/474 (46%), Positives = 305/474 (63%), Gaps = 31/474 (6%)
Query: 6 HFLIITYPLQGHINPALQFTKRLI-SLGAKVTFATTIHLYSRLI--NKPTIPGLSFATFS 62
HFL++T+P QGH+NP+L+F +RLI + GA+VTFAT + + R + N + LSF TFS
Sbjct: 5 HFLLVTFPAQGHVNPSLRFARRLIKTTGARVTFATCLSVIHRSMIPNHNNVENLSFLTFS 64
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVA 122
DG+DDG S D D+ + + F R G + L++ I +++ + P +CLIYT++ +W PKVA
Sbjct: 65 DGFDDGVISNTD-DVQNRLVHFERNGDKALSDFIEANQNGDSPVSCLIYTILPNWVPKVA 123
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
HLPS LWIQ A FDI+Y Y S + P L SL+ RDLPSF
Sbjct: 124 RRFHLPSVHLWIQPAFAFDIYYNY-----------STGNNSVFEFPNLP-SLEIRDLPSF 171
Query: 183 LLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIP 242
L SNT A +E + L EE NP++LVNT + E + L + I+M+ +GPL+P
Sbjct: 172 LSPSNTNKAAQAVYQELMDFLKEESNPKILVNTFDSLEPEFLTAIP--NIEMVAVGPLLP 229
Query: 243 SAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIAR 302
+ G + G D+ R Y WLDSK E SV+YVSFGT+ LSK+Q+EE+AR
Sbjct: 230 AEIFTGSES-----GKDLSRDHQSSSYTLWLDSKTESSVIYVSFGTMVELSKKQIEELAR 284
Query: 303 ALLDSGFSFLWVIRDKKLQQQK---EEEVDDDELS-CREELENNMNGKIVKWCSQVEVLS 358
AL++ G FLWVI DK ++ K EEE + ++++ R ELE G IV WCSQ+EVL
Sbjct: 285 ALIEGGRPFLWVITDKLNREAKIEGEEETEIEKIAGFRHELEEV--GMIVSWCSQIEVLR 342
Query: 359 HRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMV 418
HR++GCF+THCGW+S+LESL GVP+VAFP W+DQ NAKL+E++WKTG+R+ + EG+V
Sbjct: 343 HRAIGCFLTHCGWSSSLESLVLGVPVVAFPMWSDQPANAKLLEEIWKTGVRVRENSEGLV 402
Query: 419 KVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
+ EI +CLE VM K ELR NA+KWK LA A +EGGSS++N+ +++ +
Sbjct: 403 ERGEIMRCLEAVM--EAKSVELRENAEKWKRLATEAGREGGSSDKNVEAFVKSL 454
>At4g14090 glucosyltransferase like protein
Length = 456
Score = 402 bits (1033), Expect = e-112
Identities = 219/470 (46%), Positives = 301/470 (63%), Gaps = 32/470 (6%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFATFSDGY 65
H+L++T+P QGHINPALQ RLI GA VT++T + + R+ P+ GLSFA F+DG+
Sbjct: 13 HYLLVTFPAQGHINPALQLANRLIHHGATVTYSTAVSAHRRMGEPPSTKGLSFAWFTDGF 72
Query: 66 DDGQKSFGDEDIVSYMSEFTRRGSEFLTNII---LSSKQENHPFTCLIYTLILSWAPKVA 122
DDG KSF D+ I YMSE R GS L +II L + E P T +IY++++ W VA
Sbjct: 73 DDGLKSFEDQKI--YMSELKRCGSNALRDIIKANLDATTETEPITGVIYSVLVPWVSTVA 130
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
E HLP+TLLWI+ ATV DI+YYYF+ ++ + I LP L + + DLPSF
Sbjct: 131 REFHLPTTLLWIEPATVLDIYYYYFNTSYKHLFDVEP-----IKLPKLPL-ITTGDLPSF 184
Query: 183 LLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIP 242
L S AL +L+E I+ L E NP++LVNT E DAL V+ K+KMIPIGPL+
Sbjct: 185 LQPSKALPSALVTLREHIEALETESNPKILVNTFSALEHDALTSVE--KLKMIPIGPLVS 242
Query: 243 SAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAV-LSKRQMEEIA 301
S+ +GK S S +DY +WLDSK E+SV+Y+S GT A L ++ ME +
Sbjct: 243 SS--EGKTDLFKS---------SDEDYTKWLDSKLERSVIYISLGTHADDLPEKHMEALT 291
Query: 302 RALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRS 361
+L + FLW++R+K +++K+ E + + G +V WCSQ VL+H +
Sbjct: 292 HGVLATNRPFLWIVREKNPEEKKKNRF-------LELIRGSDRGLVVGWCSQTAVLAHCA 344
Query: 362 LGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVE 421
+GCF+THCGWNSTLESL SGVP+VAFPQ+ DQ T AKL+ED W+ G++++ EEG V E
Sbjct: 345 VGCFVTHCGWNSTLESLESGVPVVAFPQFADQCTTAKLVEDTWRIGVKVKVGEEGDVDGE 404
Query: 422 EIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLND 471
EIR+CLE VM GE+ EE+R NA+KWK +A A EGG S+ NL+ ++++
Sbjct: 405 EIRRCLEKVMSGGEEAEEMRENAEKWKAMAVDAAAEGGPSDLNLKGFVDE 454
>At2g43820 putative glucosyltransferase
Length = 449
Score = 310 bits (793), Expect = 1e-84
Identities = 190/473 (40%), Positives = 281/473 (59%), Gaps = 39/473 (8%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPG-LSFATFSDG 64
H L + YP QGHI P QF KRL G K T A T +++ + P + G +S AT SDG
Sbjct: 7 HVLAVPYPTQGHITPFRQFCKRLHFKGLKTTLALTTFVFNSI--NPDLSGPISIATISDG 64
Query: 65 YDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVAHE 124
YD G D I Y+ +F GS+ + +II + ++P TC++Y L WA VA E
Sbjct: 65 YDHGGFETADS-IDDYLKDFKTSGSKTIADIIQKHQTSDNPITCIVYDAFLPWALDVARE 123
Query: 125 LHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSFLL 184
L +T + Q V ++Y YI N S + + L F L+ +DLPSF
Sbjct: 124 FGLVATPFFTQPCAVNYVYYL------SYINNGSLQ----LPIEELPF-LELQDLPSFFS 172
Query: 185 ASNTYTFALPSLKEQI--QLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIP 242
S +Y P+ E + Q +N E VLVN+ +E EL N++ ++ IGP IP
Sbjct: 173 VSGSY----PAYFEMVLQQFINFEKADFVLVNSFQELELHE-NELWSKACPVLTIGPTIP 227
Query: 243 SAFLDGKDPTDNSFGGDVVRVDSKDDY--IQWLDSKDEKSVVYVSFGTLAVLSKRQMEEI 300
S +LD + +D + D+ +SKDD I WLD++ + SVVYV+FG++A L+ QMEE+
Sbjct: 228 SIYLDQRIKSDTGY--DLNLFESKDDSFCINWLDTRPQGSVVYVAFGSMAQLTNVQMEEL 285
Query: 301 ARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHR 360
A A+ S FSFLWV+R + EE++ L E N ++KW Q++VLS++
Sbjct: 286 ASAV--SNFSFLWVVRSSE-----EEKLPSGFL----ETVNKEKSLVLKWSPQLQVLSNK 334
Query: 361 SLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEE-GMVK 419
++GCF+THCGWNST+E+L GVPMVA PQWTDQ NAK I+DVWK G+R++ ++E G+ K
Sbjct: 335 AIGCFLTHCGWNSTMEALTFGVPMVAMPQWTDQPMNAKYIQDVWKAGVRVKTEKESGIAK 394
Query: 420 VEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
EEI ++ VM +GE+ +E+++N KKW+DLA ++ EGGS++ N+ ++++ +
Sbjct: 395 REEIEFSIKEVM-EGERSKEMKKNVKKWRDLAVKSLNEGGSTDTNIDTFVSRV 446
>At1g05670 UDP glycosyltransferase UGT74E1
Length = 453
Score = 292 bits (747), Expect = 3e-79
Identities = 175/477 (36%), Positives = 263/477 (54%), Gaps = 34/477 (7%)
Query: 1 MAQNHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIP------ 54
M + H +++ +P QGHI P QF KRL S K+T + +KP+ P
Sbjct: 1 MREGSHVIVLPFPAQGHITPMSQFCKRLASKSLKITLVL-------VSDKPSPPYKTEHD 53
Query: 55 GLSFATFSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLI 114
++ S+G+ +GQ+ ED+ YM L +I K +P L+Y
Sbjct: 54 TITVVPISNGFQEGQER--SEDLDEYMERVESSIKNRLPKLIEDMKLSGNPPRALVYDST 111
Query: 115 LSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSK-DETCLISLPGLSFS 173
+ W VAH L + + Q V I+Y+ F G + +K + L S P L
Sbjct: 112 MPWLLDVAHSYGLSGAVFFTQPWLVSAIYYHVFK--GSFSVPSTKYGHSTLASFPSLPI- 168
Query: 174 LKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIK 233
L + DLPSFL S++Y + L ++ +Q+ N + VL NT ++ E + L K
Sbjct: 169 LNANDLPSFLCESSSYPYILRTVIDQLS--NIDRVDIVLCNTFDKLE-EKLLKWIKSVWP 225
Query: 234 MIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLS 293
++ IGP +PS +LD + D ++G + + ++WL+SK SVVYVSFG+L VL
Sbjct: 226 VLNIGPTVPSMYLDKRLAEDKNYGFSLFGAKIAE-CMEWLNSKQPSSVVYVSFGSLVVLK 284
Query: 294 KRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQ 353
K Q+ E+A L SG FLWV+R+ + ++ E ++ E G V W Q
Sbjct: 285 KDQLIELAAGLKQSGHFFLWVVRETERRKLPENYIE----------EIGEKGLTVSWSPQ 334
Query: 354 VEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHD 413
+EVL+H+S+GCF+THCGWNSTLE L GVPM+ P W DQ TNAK +EDVWK G+R++ D
Sbjct: 335 LEVLTHKSIGCFVTHCGWNSTLEGLSLGVPMIGMPHWADQPTNAKFMEDVWKVGVRVKAD 394
Query: 414 EEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLN 470
+G V+ EE + +E VM + E+G+E+R+NA+KWK LA+ AV EGGSS++N+ +++
Sbjct: 395 SDGFVRREEFVRRVEEVM-EAEQGKEIRKNAEKWKVLAQEAVSEGGSSDKNINEFVS 450
>At1g24100 putative indole-3-acetate beta-glucosyltransferase
Length = 460
Score = 291 bits (744), Expect = 7e-79
Identities = 171/471 (36%), Positives = 258/471 (54%), Gaps = 27/471 (5%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFATFSDGY 65
H +I+ YP+QGH+NP +QF KRL+S KVT ATT + S + T P LS SDG+
Sbjct: 11 HVVILPYPVQGHLNPMVQFAKRLVSKNVKVTIATTTYTASSI----TTPSLSVEPISDGF 66
Query: 66 DDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVAHEL 125
D + +Y F GSE LT +I K + P CLIY L W +VA +
Sbjct: 67 DFIPIGIPGFSVDTYSESFKLNGSETLTLLIEKFKSTDSPIDCLIYDSFLPWGLEVARSM 126
Query: 126 HLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSFLLA 185
L + + TV + + + + + + LP LS+ +LPSF+
Sbjct: 127 ELSAASFFTNNLTVCSVLRKFSNGDFPLPADPNSAPFRIRGLPSLSYD----ELPSFV-- 180
Query: 186 SNTYTFALPSLKEQI--QLLNEEINPRVLVNTVEEFE-LDALNKVDVGKIKMIPIGPLIP 242
+ P + Q N E + VN E E + +K IGP+IP
Sbjct: 181 -GRHWLTHPEHGRVLLNQFPNHENADWLFVNGFEGLEETQDCENGESDAMKATLIGPMIP 239
Query: 243 SAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIAR 302
SA+LD + D +G +++ SK+ ++WL++K +SV +VSFG+ +L ++Q+ E+A
Sbjct: 240 SAYLDDRMEDDKDYGASLLKPISKE-CMEWLETKQAQSVAFVSFGSFGILFEKQLAEVAI 298
Query: 303 ALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRSL 362
AL +S +FLWVI++ + + E V+ + +V WC+Q+EVL+H S+
Sbjct: 299 ALQESDLNFLWVIKEAHIAKLPEGFVESTK----------DRALLVSWCNQLEVLAHESI 348
Query: 363 GCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDE-EGMVKVE 421
GCF+THCGWNSTLE L GVPMV PQW+DQ +AK +E+VWK G R + + E +VK E
Sbjct: 349 GCFLTHCGWNSTLEGLSLGVPMVGVPQWSDQMNDAKFVEEVWKVGYRAKEEAGEVIVKSE 408
Query: 422 EIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
E+ +CL+ VM +GE ++R ++KKWKDLA A+ EGGSS+R++ ++ +
Sbjct: 409 ELVRCLKGVM-EGESSVKIRESSKKWKDLAVKAMSEGGSSDRSINEFIESL 458
>At2g43840 putative glucosyltransferase
Length = 449
Score = 289 bits (739), Expect = 3e-78
Identities = 178/474 (37%), Positives = 271/474 (56%), Gaps = 41/474 (8%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFATFSDGY 65
H L + +P QGHI P QF KRL S G K T T +++ + P+ P +S AT SDGY
Sbjct: 7 HVLAVPFPSQGHITPIRQFCKRLHSKGFKTTHTLTTFIFNTIHLDPSSP-ISIATISDGY 65
Query: 66 DDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVAHEL 125
D G S + Y+ F GS+ + +II + ++P TC++Y + WA +A +
Sbjct: 66 DQGGFSSAGS-VPEYLQNFKTFGSKTVADIIRKHQSTDNPITCIVYDSFMPWALDLAMDF 124
Query: 126 HLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSFLLA 185
L + + Q+ V I Y + +G +T KD LP L+ +DLP+F+
Sbjct: 125 GLAAAPFFTQSCAVNYINYLSYINNGS-LTLPIKD------LP----LLELQDLPTFVTP 173
Query: 186 SNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFEL---DALNKVDVGKIKMIPIGPLIP 242
+ ++ + +Q N + VLVN+ + +L + L+KV ++ IGP +P
Sbjct: 174 TGSHLAYFEMVLQQFT--NFDKADFVLVNSFHDLDLHEEELLSKV----CPVLTIGPTVP 227
Query: 243 SAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIAR 302
S +LD + +DN + ++ + WLD + E SVVY++FG++A LS QMEEIA
Sbjct: 228 SMYLDQQIKSDNDYDLNLFDLKEAALCTDWLDKRPEGSVVYIAFGSMAKLSSEQMEEIAS 287
Query: 303 ALLDSGFSFLWVIR---DKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSH 359
A+ S FS+LWV+R + KL E VD D+ ++KW Q++VLS+
Sbjct: 288 AI--SNFSYLWVVRASEESKLPPGFLETVDKDK------------SLVLKWSPQLQVLSN 333
Query: 360 RSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEE-GMV 418
+++GCFMTHCGWNST+E L GVPMVA PQWTDQ NAK I+DVWK G+R++ ++E G+
Sbjct: 334 KAIGCFMTHCGWNSTMEGLSLGVPMVAMPQWTDQPMNAKYIQDVWKVGVRVKAEKESGIC 393
Query: 419 KVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
K EEI ++ VM +GEK +E++ NA KW+DLA ++ EGGS++ N+ +++ I
Sbjct: 394 KREEIEFSIKEVM-EGEKSKEMKENAGKWRDLAVKSLSEGGSTDININEFVSKI 446
>At1g05680 indole-3-acetate beta-glucosyltransferase like protein
Length = 453
Score = 287 bits (734), Expect = 1e-77
Identities = 171/477 (35%), Positives = 262/477 (54%), Gaps = 34/477 (7%)
Query: 1 MAQNHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIP------ 54
M + H +++ +P QGHI P QF KRL S G K+T + +KP+ P
Sbjct: 1 MREGSHLIVLPFPGQGHITPMSQFCKRLASKGLKLTLVL-------VSDKPSPPYKTEHD 53
Query: 55 GLSFATFSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLI 114
++ S+G+ +G++ D D YM L ++ K +P ++Y
Sbjct: 54 SITVFPISNGFQEGEEPLQDLD--DYMERVETSIKNTLPKLVEDMKLSGNPPRAIVYDST 111
Query: 115 LSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSK-DETCLISLPGLSFS 173
+ W VAH L + + Q V I+Y+ F G + +K + L S P
Sbjct: 112 MPWLLDVAHSYGLSGAVFFTQPWLVTAIYYHVFK--GSFSVPSTKYGHSTLASFPSFPM- 168
Query: 174 LKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIK 233
L + DLPSFL S++Y L + +Q+ N + VL NT ++ E L V
Sbjct: 169 LTANDLPSFLCESSSYPNILRIVVDQLS--NIDRVDIVLCNTFDKLEEKLLKWVQ-SLWP 225
Query: 234 MIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLS 293
++ IGP +PS +LD + D ++G + + ++WL+SK+ SVVY+SFG+L +L
Sbjct: 226 VLNIGPTVPSMYLDKRLSEDKNYGFSLFNAKVAE-CMEWLNSKEPNSVVYLSFGSLVILK 284
Query: 294 KRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQ 353
+ QM E+A L SG FLWV+R+ + + V+ E G IV W Q
Sbjct: 285 EDQMLELAAGLKQSGRFFLWVVRETETHKLPRNYVE----------EIGEKGLIVSWSPQ 334
Query: 354 VEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHD 413
++VL+H+S+GCF+THCGWNSTLE L GVPM+ P WTDQ TNAK ++DVWK G+R++ +
Sbjct: 335 LDVLAHKSIGCFLTHCGWNSTLEGLSLGVPMIGMPHWTDQPTNAKFMQDVWKVGVRVKAE 394
Query: 414 EEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLN 470
+G V+ EEI + +E VM +GEKG+E+R+NA+KWK LA+ AV EGGSS++++ +++
Sbjct: 395 GDGFVRREEIMRSVEEVM-EGEKGKEIRKNAEKWKVLAQEAVSEGGSSDKSINEFVS 450
>At4g15500 indole-3-acetate beta-glucosyltransferase like protein
Length = 475
Score = 284 bits (727), Expect = 7e-77
Identities = 179/491 (36%), Positives = 269/491 (54%), Gaps = 46/491 (9%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLIN----------KPTIPG 55
H +++++P QGHI+P L+ K + S G VTF TT + + KP G
Sbjct: 9 HVMLVSFPGQGHISPLLRLGKIIASKGLIVTFVTTEEPLGKKMRQANNIQDGVLKPV--G 66
Query: 56 LSFATFSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLIL 115
L F F + ++DG D D++ E + G + N++ K E P CLI +
Sbjct: 67 LGFLRF-EFFEDGFVYKEDFDLLQKSLEVS--GKREIKNLV--KKYEKQPVRCLINNAFV 121
Query: 116 SWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLK 175
W +A EL +PS +LW+Q+ +YYY H+ + T + T + +P +LK
Sbjct: 122 PWVCDIAEELQIPSAVLWVQSCACLAAYYYYHHQLVKFPTETEPEIT--VDVPFKPLTLK 179
Query: 176 SRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVD--VGKIK 233
++PSFL S+ + ++ EQI+ L++ + VL+ T +E E D ++ + ++
Sbjct: 180 HDEIPSFLHPSSPLSSIGGTILEQIKRLHKPFS--VLIETFQELEKDTIDHMSQLCPQVN 237
Query: 234 MIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLS 293
PIGPL A D GD+ + DS D I+WLDS++ SVVY+SFGTLA L
Sbjct: 238 FNPIGPLFTMAKTIRSD-----IKGDISKPDS--DCIEWLDSREPSSVVYISFGTLAFLK 290
Query: 294 KRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQ 353
+ Q++EIA +L+SG S LWV+R E V LE GKIV+WC Q
Sbjct: 291 QNQIDEIAHGILNSGLSCLWVLRPPLEGLAIEPHV--------LPLELEEKGKIVEWCQQ 342
Query: 354 VEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEH- 412
+VL+H ++ CF++HCGWNST+E+L SGVP++ FPQW DQ TNA + DV+KTGLR+
Sbjct: 343 EKVLAHPAVACFLSHCGWNSTMEALTSGVPVICFPQWGDQVTNAVYMIDVFKTGLRLSRG 402
Query: 413 --DEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLN 470
DE + + E + LE + GEK ELR NA++WK+ A +AV GG+S RN + +++
Sbjct: 403 ASDERIVPREEVAERLLEATV--GEKAVELRENARRWKEEAESAVAYGGTSERNFQEFVD 460
Query: 471 ---DIACITQI 478
D+ +T I
Sbjct: 461 KLVDVKTMTNI 471
>At3g21560 UDP-glucose:indole-3-acetate beta-D-glucosyltransferase,
putative
Length = 496
Score = 284 bits (727), Expect = 7e-77
Identities = 178/486 (36%), Positives = 270/486 (54%), Gaps = 44/486 (9%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRL---------INKPTIPG- 55
H +++++P QGH+NP L+ K L S G +TF TT ++ + KP G
Sbjct: 12 HVMLVSFPGQGHVNPLLRLGKLLASKGLLITFVTTESWGKKMRISNKIQDRVLKPVGKGY 71
Query: 56 LSFATFSDGY-DDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQ-ENHPFTCLIYTL 113
L + F DG +D + S + I+ E G + N++ K+ P TCLI
Sbjct: 72 LRYDFFDDGLPEDDEASRTNLTILRPHLELV--GKREIKNLVKRYKEVTKQPVTCLINNP 129
Query: 114 ILSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFS 173
+SW VA +L +P +LW+Q+ +YYY H D+ T + IS L
Sbjct: 130 FVSWVCDVAEDLQIPCAVLWVQSCACLAAYYYYHHNLVDFPTKTEPEIDVQISGMPL--- 186
Query: 174 LKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIK 233
LK ++PSF+ S+ ++ + +QI+ L++ + + ++T E D ++ + +
Sbjct: 187 LKHDEIPSFIHPSSPHSALREVIIDQIKRLHKTFS--IFIDTFNSLEKDIIDHMSTLSLP 244
Query: 234 MI--PIGPLIPSAFLDGKDPTDNSFGGDVVRV---DSKDDYIQWLDSKDEKSVVYVSFGT 288
+ P+GPL A + DVV+V + D ++WLDS+ SVVY+SFGT
Sbjct: 245 GVIRPLGPLYKMA---------KTVAYDVVKVNISEPTDPCMEWLDSQPVSSVVYISFGT 295
Query: 289 LAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIV 348
+A L + Q++EIA +L++ +FLWVIR ++L KE+ V +E+ + GKIV
Sbjct: 296 VAYLKQEQIDEIAYGVLNADVTFLWVIRQQELGFNKEKHVLPEEVKGK--------GKIV 347
Query: 349 KWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGL 408
+WCSQ +VLSH S+ CF+THCGWNST+E++ SGVP V FPQW DQ T+A + DVWKTG+
Sbjct: 348 EWCSQEKVLSHPSVACFVTHCGWNSTMEAVSSGVPTVCFPQWGDQVTDAVYMIDVWKTGV 407
Query: 409 RMEHD--EEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLR 466
R+ EE +V EE+ + L V KGEK EL++NA KWK+ A AAV GGSS+RNL
Sbjct: 408 RLSRGEAEERLVPREEVAERLREVT-KGEKAIELKKNALKWKEEAEAAVARGGSSDRNLE 466
Query: 467 SYLNDI 472
++ +
Sbjct: 467 KFVEKL 472
>At4g15490 indole-3-acetate beta-glucosyltransferase like protein
Length = 479
Score = 282 bits (722), Expect = 3e-76
Identities = 178/488 (36%), Positives = 267/488 (54%), Gaps = 42/488 (8%)
Query: 2 AQNHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLIN----------KP 51
+++ H +++++P QGH+NP L+ K + S G VTF TT + + + KP
Sbjct: 4 SRHTHVMLVSFPGQGHVNPLLRLGKLIASKGLLVTFVTTEKPWGKKMRQANKIQDGVLKP 63
Query: 52 TIPG-LSFATFSDGY-DDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCL 109
G + F FSDG+ DD +K F D ++ G + + N++ +E P TCL
Sbjct: 64 VGLGFIRFEFFSDGFADDDEKRF---DFDAFRPHLEAVGKQEIKNLVKRYNKE--PVTCL 118
Query: 110 IYTLILSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPG 169
I + W VA ELH+PS +LW+Q+ +YYY H + T D + + +P
Sbjct: 119 INNAFVPWVCDVAEELHIPSAVLWVQSCACLTAYYYYHHRLVKFPTKTEPDIS--VEIPC 176
Query: 170 LSFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVD- 228
L LK ++PSFL S+ YT + +Q++ + + ++T E E D ++ +
Sbjct: 177 LPL-LKHDEIPSFLHPSSPYTAFGDIILDQLKRFENHKSFYLFIDTFRELEKDIMDHMSQ 235
Query: 229 -VGKIKMIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFG 287
+ + P+GPL A + GD+ S D ++WLDS++ SVVY+SFG
Sbjct: 236 LCPQAIISPVGPLFKMA-----QTLSSDVKGDISEPAS--DCMEWLDSREPSSVVYISFG 288
Query: 288 TLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKI 347
T+A L + QMEEIA +L SG S LWV+R E V EL EE GKI
Sbjct: 289 TIANLKQEQMEEIAHGVLSSGLSVLWVVRPPMEGTFVEPHVLPREL---EE-----KGKI 340
Query: 348 VKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTG 407
V+WC Q VL+H ++ CF++HCGWNST+E+L +GVP+V FPQW DQ T+A + DV+KTG
Sbjct: 341 VEWCPQERVLAHPAIACFLSHCGWNSTMEALTAGVPVVCFPQWGDQVTDAVYLADVFKTG 400
Query: 408 LRMEHD--EEGMVKVEEI-RKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRN 464
+R+ EE +V E + K LE + GEK ELR NA++WK A AAV +GGSS+ N
Sbjct: 401 VRLGRGAAEEMIVSREVVAEKLLEATV--GEKAVELRENARRWKAEAEAAVADGGSSDMN 458
Query: 465 LRSYLNDI 472
+ +++ +
Sbjct: 459 FKEFVDKL 466
>At2g31790 putative glucosyltransferase
Length = 457
Score = 279 bits (714), Expect = 2e-75
Identities = 176/477 (36%), Positives = 259/477 (53%), Gaps = 36/477 (7%)
Query: 2 AQNHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIP---GLSF 58
A+ H L YPLQGHINP +Q KRL G +T+ + S+ +P ++
Sbjct: 4 AKKGHVLFFPYPLQGHINPMIQLAKRLSKKG----ITSTLIIASKDHREPYTSDDYSITV 59
Query: 59 ATFSDGY---DDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLIL 115
T DG+ + F D D F S LT+ I S+K ++P LIY +
Sbjct: 60 HTIHDGFFPHEHPHAKFVDLD------RFHNSTSRSLTDFISSAKLSDNPPKALIYDPFM 113
Query: 116 SWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLK 175
+A +A +L L + Q + + YY+ +E + + L S PG L
Sbjct: 114 PFALDIAKDLDLYVVAYFTQP-WLASLVYYHINEGTYDVPVDRHENPTLASFPGFPL-LS 171
Query: 176 SRDLPSFLLASNTYTFALPSLKEQI--QLLNEEINPRVLVNTVEEFELDALNKV-DVGKI 232
DLPSF +Y P L E + Q N +L NT ++ E + + D +
Sbjct: 172 QDDLPSFACEKGSY----PLLHEFVVRQFSNLLQADCILCNTFDQLEPKVVKWMNDQWPV 227
Query: 233 KMIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVL 292
K I GP++PS FLD + P D + + + + + ++WL ++ KSVVYV+FGTL L
Sbjct: 228 KNI--GPVVPSKFLDNRLPEDKDYELENSKTEPDESVLKWLGNRPAKSVVYVAFGTLVAL 285
Query: 293 SKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCS 352
S++QM+EIA A+ +G+ FLW +R+ + + + EE E +G + KW
Sbjct: 286 SEKQMKEIAMAISQTGYHFLWSVRESERSKLPSGFI--------EEAEEKDSGLVAKWVP 337
Query: 353 QVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEH 412
Q+EVL+H S+GCF++HCGWNSTLE+L GVPMV PQWTDQ TNAK IEDVWK G+R+
Sbjct: 338 QLEVLAHESIGCFVSHCGWNSTLEALCLGVPMVGVPQWTDQPTNAKFIEDVWKIGVRVRT 397
Query: 413 DEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYL 469
D EG+ EEI +C+ VM +GE+G+E+R+N +K K LAR A+ EGGSS++ + ++
Sbjct: 398 DGEGLSSKEEIARCIVEVM-EGERGKEIRKNVEKLKVLAREAISEGGSSDKKIDEFV 453
>At4g15480 indole-3-acetate beta-glucosyltransferase like protein
Length = 490
Score = 268 bits (685), Expect = 5e-72
Identities = 172/482 (35%), Positives = 266/482 (54%), Gaps = 37/482 (7%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATT------IHLYSRLIN---KPTIPG- 55
H +++++ QGH+NP L+ K + S G VTF TT + +++++ KP G
Sbjct: 19 HVMLVSFQGQGHVNPLLRLGKLIASKGLLVTFVTTELWGKKMRQANKIVDGELKPVGSGS 78
Query: 56 LSFATFSDGY--DDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTL 113
+ F F + + DD +++ D Y++ G ++ ++ ++ N P +CLI
Sbjct: 79 IRFEFFDEEWAEDDDRRA----DFSLYIAHLESVGIREVSKLVRRYEEANEPVSCLINNP 134
Query: 114 ILSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFS 173
+ W VA E ++P +LW+Q+ F +Y+Y + G + + LP +
Sbjct: 135 FIPWVCHVAEEFNIPCAVLWVQSCACFSAYYHY--QDGSVSFPTETEPELDVKLPCVPV- 191
Query: 174 LKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIK 233
LK+ ++PSFL S+ +T ++ Q + L++ VL+++ + E + ++ +
Sbjct: 192 LKNDEIPSFLHPSSRFTGFRQAILGQFKNLSKSFC--VLIDSFDSLEQEVIDYMS-SLCP 248
Query: 234 MIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLS 293
+ +GPL A D + GD+ + S D ++WLDS+ + SVVY+SFGT+A L
Sbjct: 249 VKTVGPLFKVARTVTSDVS-----GDICK--STDKCLEWLDSRPKSSVVYISFGTVAYLK 301
Query: 294 KRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQ 353
+ Q+EEIA +L SG SFLWVIR + E V EL +E G IV WC Q
Sbjct: 302 QEQIEEIAHGVLKSGLSFLWVIRPPPHDLKVETHVLPQEL---KESSAKGKGMIVDWCPQ 358
Query: 354 VEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEH- 412
+VLSH S+ CF+THCGWNST+ESL SGVP+V PQW DQ T+A + DV+KTG+R+
Sbjct: 359 EQVLSHPSVACFVTHCGWNSTMESLSSGVPVVCCPQWGDQVTDAVYLIDVFKTGVRLGRG 418
Query: 413 -DEEGMVKVEEI-RKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLN 470
EE +V EE+ K LE + GEK EELR+NA KWK A AAV GGSS++N R ++
Sbjct: 419 ATEERVVPREEVAEKLLEATV--GEKAEELRKNALKWKAEAEAAVAPGGSSDKNFREFVE 476
Query: 471 DI 472
+
Sbjct: 477 KL 478
>At2g31750 putative glucosyltransferase
Length = 456
Score = 268 bits (684), Expect = 6e-72
Identities = 169/478 (35%), Positives = 267/478 (55%), Gaps = 46/478 (9%)
Query: 8 LIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPG-----LSFATFS 62
L+ ++P+QGHINP LQF+KRL+S VTF TT ++ ++ + G LSF
Sbjct: 10 LVFSFPIQGHINPLLQFSKRLLSKNVNVTFLTTSSTHNSILRRAITGGATALPLSFVPID 69
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPF---TCLIYTL-ILSWA 118
DG+++ S + Y ++F S L+ +I S + + +CL Y L +
Sbjct: 70 DGFEEDHPS--TDTSPDYFAKFQENVSRSLSELISSMDPKPNAVVYDSCLPYVLDVCRKH 127
Query: 119 PKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRD 178
P VA + + Q++TV + ++ G++ + +++ L ++P LK D
Sbjct: 128 PGVA------AASFFTQSSTVNATYIHFLR--GEF--KEFQNDVVLPAMP----PLKGND 173
Query: 179 LPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIG 238
LP FL +N + Q +N + LVN+ +E E++ L + + + IG
Sbjct: 174 LPVFLYDNNLCRPLFELISSQF--VNVDDIDFFLVNSFDELEVEVLQWMK-NQWPVKNIG 230
Query: 239 PLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQME 298
P+IPS +LD + D +G ++ ++ + WLDSK SV+YVSFG+LAVL QM
Sbjct: 231 PMIPSMYLDKRLAGDKDYGINLFNAQV-NECLDWLDSKPPGSVIYVSFGSLAVLKDDQMI 289
Query: 299 EIARALLDSGFSFLWVIRD---KKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVE 355
E+A L +G +FLWV+R+ KKL E++ D G IV W Q++
Sbjct: 290 EVAAGLKQTGHNFLWVVRETETKKLPSNYIEDICD-------------KGLIVNWSPQLQ 336
Query: 356 VLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEE 415
VL+H+S+GCFMTHCGWNSTLE+L GV ++ P ++DQ TNAK IEDVWK G+R++ D+
Sbjct: 337 VLAHKSIGCFMTHCGWNSTLEALSLGVALIGMPAYSDQPTNAKFIEDVWKVGVRVKADQN 396
Query: 416 GMVKVEEIRKCL-EVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
G V EEI +C+ EV+ EKG+E+R+NA++ + AR A+ +GG+S++N+ ++ I
Sbjct: 397 GFVPKEEIVRCVGEVMEDMSEKGKEIRKNARRLMEFAREALSDGGNSDKNIDEFVAKI 454
>At2g23260 putative glucosyltransferase
Length = 456
Score = 254 bits (649), Expect = 7e-68
Identities = 165/478 (34%), Positives = 260/478 (53%), Gaps = 39/478 (8%)
Query: 3 QNHHFLIITYPLQGHINPALQFTKRLISLGAK---VTFATTIHLYSRL--INKPTIPGLS 57
Q H L++T P QGHINP L+ K L SL +K + AT L + KP P +
Sbjct: 7 QETHVLMVTLPFQGHINPMLKLAKHL-SLSSKNLHINLATIESARDLLSTVEKPRYP-VD 64
Query: 58 FATFSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSW 117
FSDG + + + + G+ L+ II E ++C+I + W
Sbjct: 65 LVFFSDGLPKEDPKAPE----TLLKSLNKVGAMNLSKII-----EEKRYSCIISSPFTPW 115
Query: 118 APKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSR 177
P VA ++ +LWIQA + ++Y Y+ + + + ++T + LP L L+ R
Sbjct: 116 VPAVAASHNISCAILWIQACGAYSVYYRYYMKTNSFPDLEDLNQT--VELPALPL-LEVR 172
Query: 178 DLPSFLLASNTYTF--ALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMI 235
DLPSF+L S F + + ++ + VLVN+ E E + + + K +I
Sbjct: 173 DLPSFMLPSGGAHFYNLMAEFADCLRYVKW-----VLVNSFYELESEIIESMADLK-PVI 226
Query: 236 PIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKR 295
PIGPL+ S FL G + G ++ S D ++WLD + SVVY+SFG++ +
Sbjct: 227 PIGPLV-SPFLLGDGEEETLDGKNLDFCKSDDCCMEWLDKQARSSVVYISFGSMLETLEN 285
Query: 296 QMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVE 355
Q+E IA+AL + G FLWVIR K+ Q ++ +E+ G +++W Q +
Sbjct: 286 QVETIAKALKNRGLPFLWVIRPKEKAQN---------VAVLQEMVKEGQGVVLEWSPQEK 336
Query: 356 VLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDE- 414
+LSH ++ CF+THCGWNST+E++ +GVP+VA+P WTDQ +A+L+ DV+ G+RM +D
Sbjct: 337 ILSHEAISCFVTHCGWNSTMETVVAGVPVVAYPSWTDQPIDARLLVDVFGIGVRMRNDSV 396
Query: 415 EGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
+G +KVEE+ +C+E V +G ++RR A + K +AR A+ GGSS RNL +++DI
Sbjct: 397 DGELKVEEVERCIEAVT-EGPAAVDIRRRAAELKRVARLALAPGGSSTRNLDLFISDI 453
>At2g23250 putative glucosyltransferase
Length = 438
Score = 246 bits (627), Expect = 3e-65
Identities = 169/475 (35%), Positives = 256/475 (53%), Gaps = 51/475 (10%)
Query: 9 IITYPLQGHINPALQFTKRLISLGAKVTFATTIH---LYSRLINKPTIPGLSFATFSDGY 65
++ QGH+NP L+F K L T ATT L S ++P P + A FSDG
Sbjct: 1 MVALAFQGHLNPMLKFAKHLARTNLHFTLATTEQARDLLSSTADEPHRP-VDLAFFSDGL 59
Query: 66 --DDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVAH 123
DD + D D ++ + G++ L+ II E F C+I W P VA
Sbjct: 60 PKDDPR----DPDTLA--KSLKKDGAKNLSKII-----EEKRFDCIISVPFTPWVPAVAA 108
Query: 124 ELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSFL 183
++P +LWIQA F ++Y Y+ + + + ++T + LP L L+ RDLPS +
Sbjct: 109 AHNIPCAILWIQACGAFSVYYRYYMKTNPFPDLEDLNQT--VELPALPL-LEVRDLPSLM 165
Query: 184 LAS-----NTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIG 238
L S NT + ++ VLVN+ E E + + + K +IPIG
Sbjct: 166 LPSQGANVNTLMAEFADCLKDVKW--------VLVNSFYELESEIIESMSDLK-PIIPIG 216
Query: 239 PLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQME 298
PL+ S FL G D D+ +VD D ++WLD + SVVY+SFG++ + Q+E
Sbjct: 217 PLV-SPFLLGNDEEKTL---DMWKVD--DYCMEWLDKQARSSVVYISFGSILKSLENQVE 270
Query: 299 EIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLS 358
IA AL + G FLWVIR K +K E V +E+ G + +W Q ++LS
Sbjct: 271 TIATALKNRGVPFLWVIRPK----EKGENVQ-----VLQEMVKEGKGVVTEWGQQEKILS 321
Query: 359 HRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDE-EGM 417
H ++ CF+THCGWNST+E++ +GVP+VA+P W DQ +A+L+ DV+ G+RM++D +G
Sbjct: 322 HMAISCFITHCGWNSTIETVVTGVPVVAYPTWIDQPLDARLLVDVFGIGVRMKNDAIDGE 381
Query: 418 VKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
+KV E+ +C+E V +G ++RR A + K AR+A+ GGSS +NL S+++DI
Sbjct: 382 LKVAEVERCIEAVT-EGPAAADMRRRATELKHAARSAMSPGGSSAQNLDSFISDI 435
>At2g23210 putative glucosyltransferase
Length = 453
Score = 229 bits (585), Expect = 2e-60
Identities = 160/475 (33%), Positives = 240/475 (49%), Gaps = 39/475 (8%)
Query: 3 QNHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFATFS 62
Q H L++ P QGH+NP L+F K L T AT L + L F
Sbjct: 7 QETHVLMVALPFQGHLNPMLKFAKHLARTNLHFTLATIESARDLLSSTDEPHSLVDLVF- 65
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVA 122
+ DG D + G+ + II E F C+I W P VA
Sbjct: 66 --FSDGLPKDDPRDHEPLTESLRKVGANNFSKII-----EGKRFDCIISVPFTPWVPAVA 118
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
++P +LWI+A F ++Y Y+ + + +D + LPGL F L+ RDLP+
Sbjct: 119 AAHNIPCAILWIEACAGFSVYYRYYMKTNSF--PDLEDPNQKVELPGLPF-LEVRDLPTL 175
Query: 183 LLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIP 242
+L S+ F + E ++ L + VL N+ E E + + K +IPIGPL+
Sbjct: 176 MLPSHGAIFNT-LMAEFVECLKDV--KWVLANSFYELESVIIESMFDLK-PIIPIGPLV- 230
Query: 243 SAFLDGKDPTDNSFGGDVVRVDSKDDY-IQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIA 301
S FL G D D G + + DDY ++WLD + ++ S+ Q+E IA
Sbjct: 231 SPFLLGADE-DKILDGKSLDMWKADDYCMEWLDKQV----------SILKSSENQVETIA 279
Query: 302 RALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRS 361
AL + G FLWVIR K +K E VD E++ G +++W Q ++L H +
Sbjct: 280 TALKNRGVPFLWVIRPK----EKAENVD-----VLEDMVEEGQGVVIEWGQQEKILCHMA 330
Query: 362 LGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHD-EEGMVKV 420
+ CF+THCGWNST+E++ SGVPMVA+P W DQ +A+L+ DV+ G+RM++D +G +KV
Sbjct: 331 ISCFVTHCGWNSTIETVVSGVPMVAYPTWFDQPLDARLLVDVFGIGVRMKNDVVDGELKV 390
Query: 421 EEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDIACI 475
E+ +C++ V KG ++RR A + K R+A+ GGS RNL ++NDI +
Sbjct: 391 AEVERCIDAVT-KGTDAADMRRRAAELKQATRSAMAPGGSLARNLDLFINDIKIV 444
>At1g22340 hypothetical protein
Length = 487
Score = 212 bits (540), Expect = 3e-55
Identities = 148/490 (30%), Positives = 249/490 (50%), Gaps = 36/490 (7%)
Query: 2 AQNHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINK------PTIPG 55
AQ H + + YP QGHINP L+ K L + G VTF T++ ++RL+ P
Sbjct: 9 AQKPHVVCVPYPAQGHINPMLKVAKLLYAKGFHVTFVNTLYNHNRLLRSRGPNALDGFPS 68
Query: 56 LSFATFSDGYD--DGQKSFGDEDI-VSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYT 112
F + DG DG ++ + +S E L I + K + P +C++
Sbjct: 69 FRFESIPDGLPETDGDRTQHTPTVCMSIEKNCLAPFKEILRRI--NDKDDVPPVSCIVSD 126
Query: 113 LILSWAPKVAHELHLPSTLLWIQAATVFD--IFYYYFHEHG------DYITNKSKDETCL 164
++S+ A EL +P + W +A F + +Y F E G + +K +T +
Sbjct: 127 GVMSFTLDAAEELGVPEVIFWTNSACGFMTILHFYLFIEKGLSPFKDESYMSKEHLDTVI 186
Query: 165 ISLPGLSFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDAL 224
+P + +L+ +D+PS++ +N L L +++ + +++NT +E E D +
Sbjct: 187 DWIPSMK-NLRLKDIPSYIRTTNPDNIMLNFLIREVE--RSKRASAIILNTFDELEHDVI 243
Query: 225 NKVDVGKIKMIPIGPLIPSAFLDGKDPTD-NSFGGDVVRVDSKDDYIQWLDSKDEKSVVY 283
+ + IGPL + + ++ G ++ R + + + WLD+K SV++
Sbjct: 244 QSMQSILPPVYSIGPLHLLVKEEINEASEIGQMGLNLWREEM--ECLDWLDTKTPNSVLF 301
Query: 284 VSFGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNM 343
V+FG + V+S +Q+EE A L S FLWVIR + + + + L+ E ++ M
Sbjct: 302 VNFGCITVMSAKQLEEFAWGLAASRKEFLWVIRPNLVVGEAMVVLPQEFLA--ETIDRRM 359
Query: 344 NGKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDV 403
+ WC Q +VLSH ++G F+THCGWNSTLESL GVPM+ +P +++Q TN K D
Sbjct: 360 ---LASWCPQEKVLSHPAIGGFLTHCGWNSTLESLAGGVPMICWPCFSEQPTNCKFCCDE 416
Query: 404 WKTGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVK-EGGSSN 462
W G+ + D VK EE+ + +M GEKG++LR A++W+ LA A + + GSS
Sbjct: 417 WGVGIEIGKD----VKREEVETVVRELM-DGEKGKKLREKAEEWRRLAEEATRYKHGSSV 471
Query: 463 RNLRSYLNDI 472
NL + ++ +
Sbjct: 472 MNLETLIHKV 481
>At1g22360 unknown protein
Length = 481
Score = 209 bits (533), Expect = 2e-54
Identities = 151/493 (30%), Positives = 244/493 (48%), Gaps = 41/493 (8%)
Query: 1 MAQNHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINK------PTIP 54
+AQ H + + YP QGHINP ++ K L + G +TF T++ ++RL+ +P
Sbjct: 5 VAQKQHVVCVPYPAQGHINPMMKVAKLLYAKGFHITFVNTVYNHNRLLRSRGPNAVDGLP 64
Query: 55 GLSFATFSDGYDDGQKSFGDEDIV----SYMSEFTRRGSEFLTNIILSSKQENHPFTCLI 110
F + DG + +DI S M E L I +++ + P +C++
Sbjct: 65 SFRFESIPDGLPETDVDV-TQDIPTLCESTMKHCLAPFKELLRQI--NARDDVPPVSCIV 121
Query: 111 YTLILSWAPKVAHELHLPSTLLWIQAATVF--DIFYYYFHEHG-------DYITNKSKDE 161
+S+ A EL +P L W +A F ++YY F E G Y+T + D
Sbjct: 122 SDGCMSFTLDAAEELGVPEVLFWTTSACGFLAYLYYYRFIEKGLSPIKDESYLTKEHLD- 180
Query: 162 TCLISLPGLSFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFEL 221
T + +P + +L+ +D+PSF+ +N L + + +++NT ++ E
Sbjct: 181 TKIDWIPSMK-NLRLKDIPSFIRTTNPDDIMLNFIIREADRAKRA--SAIILNTFDDLEH 237
Query: 222 DALNKVDVGKIKMIPIGPL-IPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKS 280
D + + + IGPL + G+ G ++ R ++ + + WL++K S
Sbjct: 238 DVIQSMKSIVPPVYSIGPLHLLEKQESGEYSEIGRTGSNLWREET--ECLDWLNTKARNS 295
Query: 281 VVYVSFGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELE 340
VVYV+FG++ VLS +Q+ E A L +G FLWVIR L E V + L+ +
Sbjct: 296 VVYVNFGSITVLSAKQLVEFAWGLAATGKEFLWVIRPD-LVAGDEAMVPPEFLTATADRR 354
Query: 341 NNMNGKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLI 400
+ WC Q +VLSH ++G F+THCGWNSTLESL GVPMV +P + +Q TN K
Sbjct: 355 -----MLASWCPQEKVLSHPAIGGFLTHCGWNSTLESLCGGVPMVCWPFFAEQQTNCKFS 409
Query: 401 EDVWKTGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVK-EGG 459
D W+ G+ + D VK EE+ + +M + EKG+ +R A++W+ LA A + + G
Sbjct: 410 RDEWEVGIEIGGD----VKREEVEAVVRELMDE-EKGKNMREKAEEWRRLANEATEHKHG 464
Query: 460 SSNRNLRSYLNDI 472
SS N +N +
Sbjct: 465 SSKLNFEMLVNKV 477
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,385,204
Number of Sequences: 26719
Number of extensions: 517959
Number of successful extensions: 2147
Number of sequences better than 10.0: 129
Number of HSP's better than 10.0 without gapping: 116
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 1656
Number of HSP's gapped (non-prelim): 173
length of query: 478
length of database: 11,318,596
effective HSP length: 103
effective length of query: 375
effective length of database: 8,566,539
effective search space: 3212452125
effective search space used: 3212452125
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146706.4


