

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146705.10 - phase: 0
(146 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
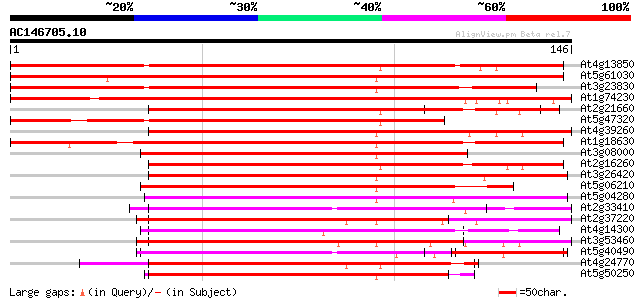
Score E
Sequences producing significant alignments: (bits) Value
At4g13850 glycine-rich RNA-binding protein AtGRP2 - like 187 2e-48
At5g61030 RNA-binding protein - like 180 2e-46
At3g23830 glycine-rich RNA binding protein, putative 169 6e-43
At1g74230 putative RNA-binding protein 163 3e-41
At2g21660 glycine-rich RNA binding protein 126 4e-30
At5g47320 40S ribosomal protein S19 120 2e-28
At4g39260 glycine-rich protein (clone AtGRP8) 120 3e-28
At1g18630 unknown protein 106 4e-24
At3g08000 RNA-binding protein like 101 2e-22
At2g16260 putative glycine-rich RNA-binding protein 100 2e-22
At3g26420 RNA-binding protein, putative 93 4e-20
At5g06210 unknown protein 90 4e-19
At5g04280 RNA-binding protein-like 87 4e-18
At2g33410 putative RNA-binding protein 87 4e-18
At2g37220 putative RNA-binding protein 86 9e-18
At4g14300 ribonucleoprotein like protein 85 1e-17
At3g53460 RNA-binding protein cp29 protein 85 2e-17
At5g40490 ribonucleoprotein -like 84 3e-17
At4g24770 31 kDa RNA binding protein (rbp31) 80 3e-16
At5g50250 RNA-binding protein-like 79 9e-16
>At4g13850 glycine-rich RNA-binding protein AtGRP2 - like
Length = 158
Score = 187 bits (474), Expect = 2e-48
Identities = 100/149 (67%), Positives = 117/149 (78%), Gaps = 7/149 (4%)
Query: 1 MAFCNKIGNLLRQGATQSTQAPVSSMLNYLRHMSSSKLFIGGLSYNVDDQSLRDAFTTYG 60
MAFCNK+G LLRQ + + PV+SML LR MS+ KLFIGGLS+ DD SLRDAF +G
Sbjct: 1 MAFCNKLGGLLRQNISSNGNVPVTSMLGSLRLMST-KLFIGGLSWGTDDASLRDAFAHFG 59
Query: 61 DVVEARVITDRETGRSRGFGFVNFTSEESATSALS-MDGQDLNGRNIRVSYANDRQAGGP 119
DVV+A+VI DRETGRSRGFGFVNF E +AT+A+S MDG++LNGR+IRV+ ANDR + P
Sbjct: 60 DVVDAKVIVDRETGRSRGFGFVNFNDEGAATAAISEMDGKELNGRHIRVNPANDRPS-AP 118
Query: 120 RP---GGGY-GGGGGYSGGGGGYSGGGGG 144
R GGGY GGGGGY GGGGGY GGGGG
Sbjct: 119 RAYGGGGGYSGGGGGYGGGGGGYGGGGGG 147
>At5g61030 RNA-binding protein - like
Length = 309
Score = 180 bits (457), Expect = 2e-46
Identities = 90/149 (60%), Positives = 115/149 (76%), Gaps = 5/149 (3%)
Query: 1 MAFCNKIGNLLRQGATQSTQAPVS----SMLNYLRHMSSSKLFIGGLSYNVDDQSLRDAF 56
MAF +K GN+L+Q + A VS S+ +R MSSSKLFIGG++Y++D+ SLR+AF
Sbjct: 1 MAFLSKFGNILKQTTNKQLNAQVSLSSPSLFQAIRCMSSSKLFIGGMAYSMDEDSLREAF 60
Query: 57 TTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-SMDGQDLNGRNIRVSYANDRQ 115
T YG+VV+ RVI DRETGRSRGFGFV FTS E+A+SA+ ++DG+DL+GR ++V+YANDR
Sbjct: 61 TKYGEVVDTRVILDRETGRSRGFGFVTFTSSEAASSAIQALDGRDLHGRVVKVNYANDRT 120
Query: 116 AGGPRPGGGYGGGGGYSGGGGGYSGGGGG 144
+GG GGGYGGGGG GG GGY GG GG
Sbjct: 121 SGGGFGGGGYGGGGGGYGGSGGYGGGAGG 149
Score = 28.1 bits (61), Expect = 1.7
Identities = 10/20 (50%), Positives = 11/20 (55%)
Query: 122 GGGYGGGGGYSGGGGGYSGG 141
GGGYG GG+ G Y G
Sbjct: 198 GGGYGSSGGFGSSGNTYGEG 217
>At3g23830 glycine-rich RNA binding protein, putative
Length = 136
Score = 169 bits (427), Expect = 6e-43
Identities = 84/138 (60%), Positives = 107/138 (76%), Gaps = 5/138 (3%)
Query: 1 MAFCNKIGNLLRQGATQSTQAPVSSMLNYLRHMSSSKLFIGGLSYNVDDQSLRDAFTTYG 60
MAFCNK+ +LRQG +QS+ PV+SML LR+MSS KLF+GGLS+ DD SL+ AFT++G
Sbjct: 1 MAFCNKLSGILRQGVSQSSNGPVTSMLGSLRYMSS-KLFVGGLSWGTDDSSLKQAFTSFG 59
Query: 61 DVVEARVITDRETGRSRGFGFVNFTSEESATSAL-SMDGQDLNGRNIRVSYANDRQAGGP 119
+V EA VI DRETGRSRGFGFV+F+ E+SA +A+ MDG++LNGR IRV+ A +R +
Sbjct: 60 EVTEATVIADRETGRSRGFGFVSFSCEDSANNAIKEMDGKELNGRQIRVNLATERSSA-- 117
Query: 120 RPGGGYGGGGGYSGGGGG 137
P +GGGGGY GGGGG
Sbjct: 118 -PRSSFGGGGGYGGGGGG 134
>At1g74230 putative RNA-binding protein
Length = 289
Score = 163 bits (412), Expect = 3e-41
Identities = 90/167 (53%), Positives = 116/167 (68%), Gaps = 23/167 (13%)
Query: 1 MAFCNKIGNLLRQGATQSTQAPVSSMLNYLRHMSSSKLFIGGLSYNVDDQSLRDAFTTYG 60
MAF +K+G L Q ++ T + SSML +R MSSSK+F+GG+SY+ D+ LR+AF+ YG
Sbjct: 1 MAFLSKVGRLFSQTSSHVTAS--SSMLQSIRCMSSSKIFVGGISYSTDEFGLREAFSKYG 58
Query: 61 DVVEARVITDRETGRSRGFGFVNFTSEESATSALSMDGQDLNGRNIRVSYANDRQAG-GP 119
+VV+A++I DRETGRSRGF FV FTS E A++A+ +DGQDL+GR IRV+YA +R +G G
Sbjct: 59 EVVDAKIIVDRETGRSRGFAFVTFTSTEEASNAMQLDGQDLHGRRIRVNYATERGSGFGG 118
Query: 120 R----PGGGYG---GG-----GGYSGGGGGYSG--------GGGGGW 146
R PGGGYG GG GGY GG GGY G GGGGG+
Sbjct: 119 RGFGGPGGGYGASDGGYGAPAGGYGGGAGGYGGNSSYSGNAGGGGGY 165
>At2g21660 glycine-rich RNA binding protein
Length = 176
Score = 126 bits (317), Expect = 4e-30
Identities = 64/111 (57%), Positives = 84/111 (75%), Gaps = 7/111 (6%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSALS- 95
+ F+GGL++ DD++L AF YGDV+++++I DRETGRSRGFGFV F E++ A+
Sbjct: 9 RCFVGGLAWATDDRALETAFAQYGDVIDSKIINDRETGRSRGFGFVTFKDEKAMKDAIEG 68
Query: 96 MDGQDLNGRNIRVSYANDRQAGGPRPGGGY--GGGGGY-SGGGGGYSGGGG 143
M+GQDL+GR+I V+ A R +GG GGG+ GGGGGY SGGGGGYSGGGG
Sbjct: 69 MNGQDLDGRSITVNEAQSRGSGG---GGGHRGGGGGGYRSGGGGGYSGGGG 116
Score = 39.3 bits (90), Expect = 7e-04
Identities = 18/30 (60%), Positives = 20/30 (66%)
Query: 109 SYANDRQAGGPRPGGGYGGGGGYSGGGGGY 138
SY R+ GG GGG GGG G SGGGGG+
Sbjct: 147 SYGGGRREGGGGYGGGEGGGYGGSGGGGGW 176
>At5g47320 40S ribosomal protein S19
Length = 212
Score = 120 bits (302), Expect = 2e-28
Identities = 66/114 (57%), Positives = 85/114 (73%), Gaps = 6/114 (5%)
Query: 1 MAFCNKIGNLLRQGATQSTQAPVSSMLNYLRHMSSSKLFIGGLSYNVDDQSLRDAFTTYG 60
MAFC K+G +QG PVSSML LR+MS+ KL+IGGLS D+ SL+DAF+++
Sbjct: 1 MAFCTKLGGHWKQGVN----VPVSSMLGSLRYMST-KLYIGGLSPGTDEHSLKDAFSSFN 55
Query: 61 DVVEARVITDRETGRSRGFGFVNFTSEESATSALS-MDGQDLNGRNIRVSYAND 113
V EARV+T++ TGRSRG+GFVNF SE+SA SA+S M+GQ+LNG NI V+ A D
Sbjct: 56 GVTEARVMTNKVTGRSRGYGFVNFISEDSANSAISAMNGQELNGFNISVNVAKD 109
>At4g39260 glycine-rich protein (clone AtGRP8)
Length = 169
Score = 120 bits (301), Expect = 3e-28
Identities = 61/119 (51%), Positives = 86/119 (72%), Gaps = 9/119 (7%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
+ F+GGL++ +D+ L+ F+ +GDV+++++I DRE+GRSRGFGFV F E++ A+
Sbjct: 7 RCFVGGLAWATNDEDLQRTFSQFGDVIDSKIINDRESGRSRGFGFVTFKDEKAMRDAIEE 66
Query: 96 MDGQDLNGRNIRVSYANDRQAGG-----PRPGGGY--GGGGGYS-GGGGGYSGGGGGGW 146
M+G++L+GR I V+ A R +GG GGGY GGGGGYS GGGGGYSGGGGGG+
Sbjct: 67 MNGKELDGRVITVNEAQSRGSGGGGGGRGGSGGGYRSGGGGGYSGGGGGGYSGGGGGGY 125
>At1g18630 unknown protein
Length = 155
Score = 106 bits (265), Expect = 4e-24
Identities = 62/146 (42%), Positives = 93/146 (63%), Gaps = 9/146 (6%)
Query: 1 MAFCNKIGNLLRQG-ATQSTQAPVSSMLNYLRHMSSSKLFIGGLSYNVDDQSLRDAFTTY 59
M ++ GN+ RQ A Q++ A + L+ ++ SK+F+GGLS + D + L++AF ++
Sbjct: 4 MGLFSRAGNIFRQPRALQASNAMLQGNLS----LTPSKIFVGGLSPSTDVELLKEAFGSF 59
Query: 60 GDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-SMDGQDLNGRNIRVSYANDRQAGG 118
G +V+A V+ DRE+G SRGFGFV + S E A +A+ +M ++L+GR I V A+ GG
Sbjct: 60 GKIVDAVVVLDRESGLSRGFGFVTYDSIEVANNAMQAMQNKELDGRIIGVHPADSGGGGG 119
Query: 119 PRPGGGYGGGGGYSGGGGGYSGGGGG 144
GGG+ GGY GG GGY+ GG G
Sbjct: 120 ---GGGFARRGGYGGGRGGYARGGFG 142
>At3g08000 RNA-binding protein like
Length = 143
Score = 101 bits (251), Expect = 2e-22
Identities = 50/86 (58%), Positives = 68/86 (78%), Gaps = 1/86 (1%)
Query: 35 SSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL 94
SSKLFIGGLS++VD+QSL+DAF+++G+V E R+ D+ +GRSRGFGFV+F E A SA
Sbjct: 40 SSKLFIGGLSWSVDEQSLKDAFSSFGEVAEVRIAYDKGSGRSRGFGFVDFAEEGDALSAK 99
Query: 95 -SMDGQDLNGRNIRVSYANDRQAGGP 119
+MDG+ L GR +R+S+A +R GGP
Sbjct: 100 DAMDGKGLLGRPLRISFALERVRGGP 125
>At2g16260 putative glycine-rich RNA-binding protein
Length = 185
Score = 100 bits (250), Expect = 2e-22
Identities = 54/118 (45%), Positives = 74/118 (61%), Gaps = 12/118 (10%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSALS- 95
+ F+GGL++ D+QS+ F +G+V ++++I DRETGRS+GF FV F E+S +A+
Sbjct: 45 RCFVGGLAWATDEQSIERCFNEFGEVFDSKIIIDRETGRSKGFRFVTFKDEDSMRTAIDR 104
Query: 96 MDGQDLNGRNIRVSYANDRQAGGPRPGGGYGGG--------GGYS-GGGGGYSGGGGG 144
M+GQ+L+GRNI GG GGYG G GGY+ GGGGGY GG GG
Sbjct: 105 MNGQELDGRNITAQARGSGTRGG--MVGGYGSGGYRGRRDQGGYNRGGGGGYGGGYGG 160
>At3g26420 RNA-binding protein, putative
Length = 245
Score = 93.2 bits (230), Expect = 4e-20
Identities = 50/111 (45%), Positives = 67/111 (60%), Gaps = 2/111 (1%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
+ FIGGL++ D+ LRDAF YG +VEA+V+ D+ +GRSRGFGF+ F +++ A+ +
Sbjct: 8 RCFIGGLAWTTSDRGLRDAFEKYGHLVEAKVVLDKFSGRSRGFGFITFDEKKAMDEAIAA 67
Query: 96 MDGQDLNGRNIRVSYANDRQAGGPRPG-GGYGGGGGYSGGGGGYSGGGGGG 145
M+G DL+GR I V A Q G R G G GY SGG GGG
Sbjct: 68 MNGMDLDGRTITVDKAQPHQGGAGRDNDGDRGRDRGYDRDRSRPSGGRGGG 118
>At5g06210 unknown protein
Length = 146
Score = 90.1 bits (222), Expect = 4e-19
Identities = 49/98 (50%), Positives = 63/98 (64%), Gaps = 9/98 (9%)
Query: 35 SSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL 94
+SKLFIGGLS+ +Q L +AF+ G VVEA+++ DR + RS+GFGFV F S + A AL
Sbjct: 33 ASKLFIGGLSFCTTEQGLSEAFSKCGQVVEAQIVMDRVSDRSKGFGFVTFASADEAQKAL 92
Query: 95 -SMDGQDLNGRNIRVSYANDRQAGGPRPGGGYGGGGGY 131
+GQ LNGR I V YA +Q+ GGGGGY
Sbjct: 93 MEFNGQQLNGRTIFVDYAKAKQS--------LGGGGGY 122
>At5g04280 RNA-binding protein-like
Length = 310
Score = 86.7 bits (213), Expect = 4e-18
Identities = 45/113 (39%), Positives = 67/113 (58%), Gaps = 3/113 (2%)
Query: 36 SKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL- 94
S++F+GGLS V D+ L AF+ +GD+++ +++ +R+TGRSRGFGF+ F + ++
Sbjct: 7 SRIFVGGLSPEVTDRDLERAFSRFGDILDCQIMLERDTGRSRGFGFITFADRRAMDESIR 66
Query: 95 SMDGQDLNGRNIRVSYANDR--QAGGPRPGGGYGGGGGYSGGGGGYSGGGGGG 145
M G+D R I V+ A + + G G G GYS G G GGGGGG
Sbjct: 67 EMHGRDFGDRVISVNRAEPKLGRDDGESHGSRGGRDSGYSIAGKGSFGGGGGG 119
Score = 31.2 bits (69), Expect = 0.20
Identities = 22/49 (44%), Positives = 25/49 (50%), Gaps = 9/49 (18%)
Query: 104 RNIRVSYANDR----QAGGPRPGGGYGGGGGYSGGGGGY-----SGGGG 143
R+I S +DR +AGGP GGG G G S G Y SGGGG
Sbjct: 251 RDIAPSAGSDRYGGGRAGGPIRGGGEEGRGFRSRAGAPYERPSRSGGGG 299
>At2g33410 putative RNA-binding protein
Length = 404
Score = 86.7 bits (213), Expect = 4e-18
Identities = 48/116 (41%), Positives = 65/116 (55%), Gaps = 3/116 (2%)
Query: 32 HMSSSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESAT 91
++ + K+F+GGL + R F TYG V +A ++ D+ T R RGFGFV+F SE+S
Sbjct: 106 NVRTKKIFVGGLPPALTSDEFRAYFETYGPVSDAVIMIDQTTQRPRGFGFVSFDSEDSVD 165
Query: 92 SALSMDGQDLNGRNIRVSYANDRQAG-GPRPGGGYGGGGGYSGGGGGYSGGGGGGW 146
L DLNG+ + V A + A G GGG G GG +GG GY G GG G+
Sbjct: 166 LVLHKTFHDLNGKQVEVKRALPKDANPGIASGGGRGSGG--AGGFPGYGGSGGSGY 219
Score = 65.5 bits (158), Expect = 1e-11
Identities = 32/88 (36%), Positives = 53/88 (59%), Gaps = 1/88 (1%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSALSM 96
KLFIGG+S++ D+ LR+ F+ +G+V++ V+ ++ TGR RGFGFV F S+ + +
Sbjct: 7 KLFIGGISWDTDENLLREYFSNFGEVLQVTVMREKATGRPRGFGFVAF-SDPAVIDRVLQ 65
Query: 97 DGQDLNGRNIRVSYANDRQAGGPRPGGG 124
D ++ R++ V A R+ P G
Sbjct: 66 DKHHIDNRDVDVKRAMSREEQSPAGRSG 93
Score = 27.7 bits (60), Expect = 2.3
Identities = 21/70 (30%), Positives = 28/70 (40%), Gaps = 3/70 (4%)
Query: 76 SRGFGFVNFTSEESATSALSMDGQDLNGRNIRVSYANDRQAGGPRPGGGYGGGGGYSGGG 135
S+G+G+ S SA G N+ ++ A GGYG G + G
Sbjct: 332 SQGYGYGGNDSSYGTPSAYGAVGG--RSGNMPNNHGGGGYADALDGSGGYGNHQG-NNGQ 388
Query: 136 GGYSGGGGGG 145
GY GG G G
Sbjct: 389 AGYGGGYGSG 398
>At2g37220 putative RNA-binding protein
Length = 289
Score = 85.5 bits (210), Expect = 9e-18
Identities = 42/82 (51%), Positives = 59/82 (71%), Gaps = 1/82 (1%)
Query: 34 SSSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSA 93
S +++++G LS+ VDD +L F+ G VVEARVI DR++GRS+GFGFV + S + +A
Sbjct: 202 SGNRVYVGNLSWGVDDMALESLFSEQGKVVEARVIYDRDSGRSKGFGFVTYDSSQEVQNA 261
Query: 94 L-SMDGQDLNGRNIRVSYANDR 114
+ S+DG DL+GR IRVS A R
Sbjct: 262 IKSLDGADLDGRQIRVSEAEAR 283
Score = 80.9 bits (198), Expect = 2e-16
Identities = 50/122 (40%), Positives = 63/122 (50%), Gaps = 12/122 (9%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTS-EESATSALS 95
KLF+G L +NVD L F + G+V VI D+ TGRSRGFGFV +S E +A
Sbjct: 92 KLFVGNLPFNVDSAQLAQLFESAGNVEMVEVIYDKITGRSRGFGFVTMSSVSEVEAAAQQ 151
Query: 96 MDGQDLNGRNIRVSYA------NDRQAGGPR-----PGGGYGGGGGYSGGGGGYSGGGGG 144
+G +L+GR +RV+ D + GPR G GYGGGGG G G G
Sbjct: 152 FNGYELDGRPLRVNAGPPPPKREDGFSRGPRSSFGSSGSGYGGGGGSGAGSGNRVYVGNL 211
Query: 145 GW 146
W
Sbjct: 212 SW 213
>At4g14300 ribonucleoprotein like protein
Length = 406
Score = 85.1 bits (209), Expect = 1e-17
Identities = 45/109 (41%), Positives = 61/109 (55%), Gaps = 5/109 (4%)
Query: 35 SSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL 94
+ K+F+GGL + D+ R F YG V + ++ D+ T R RGFGFV+F SE++ S L
Sbjct: 104 TKKIFVGGLPPTLTDEEFRQYFEVYGPVTDVAIMYDQATNRPRGFGFVSFDSEDAVDSVL 163
Query: 95 SMDGQDLNGRNIRVSYANDRQAGGPRPGGGYGGGGGYSGGGGGYSGGGG 143
DL+G+ + V A + A PGGG GG GG GGY G GG
Sbjct: 164 HKTFHDLSGKQVEVKRALPKDA---NPGGGGRSMGG--GGSGGYQGYGG 207
Score = 57.4 bits (137), Expect = 3e-09
Identities = 33/94 (35%), Positives = 51/94 (54%), Gaps = 14/94 (14%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFG------------FVNF 84
KLF+GG+S+ D+ LR+ FT YG+V +A V+ D+ TGR RGFG +V
Sbjct: 7 KLFVGGISWETDEDKLREHFTNYGEVSQAIVMRDKLTGRPRGFGIRKNRCIYFLLRYVLD 66
Query: 85 TSEESATSALSMDGQDLNGRNIRVSYANDRQAGG 118
+ A+S + Q ++GR ++ R +GG
Sbjct: 67 LGKVDVKRAMSREEQQVSGRTGNLN--TSRSSGG 98
Score = 35.4 bits (80), Expect = 0.011
Identities = 14/30 (46%), Positives = 16/30 (52%)
Query: 117 GGPRPGGGYGGGGGYSGGGGGYSGGGGGGW 146
G G GYG GGY+G GGY G G+
Sbjct: 244 GNGSNGAGYGAYGGYTGSAGGYGAGATAGY 273
Score = 31.6 bits (70), Expect = 0.16
Identities = 22/67 (32%), Positives = 28/67 (40%), Gaps = 1/67 (1%)
Query: 78 GFGFVNFTSEESATSALSMDGQDLNGRNIRVSYANDRQAGGPRPGGGYGGGGGYSGGGGG 137
G+G + S S G G + S ++ A G GG GGG + G GG
Sbjct: 316 GYGVPGAAPPTQSPSGYSNQGYGYGGYSGSDSGYGNQAAYGV-VGGRPSGGGSNNPGSGG 374
Query: 138 YSGGGGG 144
Y GGG G
Sbjct: 375 YMGGGYG 381
Score = 28.9 bits (63), Expect = 1.0
Identities = 22/72 (30%), Positives = 35/72 (48%), Gaps = 7/72 (9%)
Query: 76 SRGFGFVNFTSEESATSALSMDGQDLNGRNIRVSYANDRQAGGPRPGGGYGGGGGYS--- 132
++G+G+ ++ +S + G + G +N+ +GG GGGYG G S
Sbjct: 334 NQGYGYGGYSGSDSGYGNQAAYG--VVGGRPSGGGSNNPGSGGYM-GGGYGDGSWRSDPS 390
Query: 133 -GGGGGYSGGGG 143
G GGGY+ G G
Sbjct: 391 QGYGGGYNDGQG 402
>At3g53460 RNA-binding protein cp29 protein
Length = 342
Score = 84.7 bits (208), Expect = 2e-17
Identities = 43/86 (50%), Positives = 59/86 (68%), Gaps = 1/86 (1%)
Query: 34 SSSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSA 93
S ++L++G LS+ VDD +L + F G VVEARVI DR++GRS+GFGFV +S + A
Sbjct: 255 SGNRLYVGNLSWGVDDMALENLFNEQGKVVEARVIYDRDSGRSKGFGFVTLSSSQEVQKA 314
Query: 94 L-SMDGQDLNGRNIRVSYANDRQAGG 118
+ S++G DL+GR IRVS A R G
Sbjct: 315 INSLNGADLDGRQIRVSEAEARPPRG 340
Score = 84.3 bits (207), Expect = 2e-17
Identities = 54/127 (42%), Positives = 69/127 (53%), Gaps = 17/127 (13%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNF-TSEESATSALS 95
KLF+G LS+NVD L F + G+V VI D+ TGRSRGFGFV T+ E +A
Sbjct: 100 KLFVGNLSFNVDSAQLAQLFESAGNVEMVEVIYDKVTGRSRGFGFVTMSTAAEVEAAAQQ 159
Query: 96 MDGQDLNGRNIRV-----------SYANDRQAG--GPRPGGGYGG--GGGY-SGGGGGYS 139
+G + GR +RV S++ ++G G GGGYG GGGY S GGGY
Sbjct: 160 FNGYEFEGRPLRVNAGPPPPKREESFSRGPRSGGYGSERGGGYGSERGGGYGSERGGGYG 219
Query: 140 GGGGGGW 146
GGG+
Sbjct: 220 SERGGGY 226
>At5g40490 ribonucleoprotein -like
Length = 423
Score = 84.0 bits (206), Expect = 3e-17
Identities = 45/130 (34%), Positives = 68/130 (51%), Gaps = 19/130 (14%)
Query: 35 SSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL 94
+ K+F+GG+ +VDD ++ F +G++ E +++ D TGRSRGFGFV + SE+ L
Sbjct: 129 TKKIFVGGIPSSVDDDEFKEFFMQFGELKEHQIMRDHSTGRSRGFGFVTYESEDMVDHLL 188
Query: 95 SMDGQ-DLNGRNIRVS----------------YANDRQAGGPRPGGGYGG--GGGYSGGG 135
+ + +L+G + + + + R G G GYGG GGGY G G
Sbjct: 189 AKGNRIELSGTQVEIKKAEPKKPNSVTTPSKRFGDSRSNFGGGYGDGYGGGHGGGYGGPG 248
Query: 136 GGYSGGGGGG 145
G Y GGG G
Sbjct: 249 GPYKSGGGYG 258
Score = 51.2 bits (121), Expect = 2e-07
Identities = 22/75 (29%), Positives = 42/75 (55%), Gaps = 1/75 (1%)
Query: 34 SSSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSA 93
S+ K+F+GGL+ F YG++ ++ ++ DR+TG+ RGFGFV + ++ S
Sbjct: 40 SAGKIFVGGLARETTSAEFLKHFGKYGEITDSVIMKDRKTGQPRGFGFVTY-ADSSVVDK 98
Query: 94 LSMDGQDLNGRNIRV 108
+ D + G+ + +
Sbjct: 99 VIQDNHIIIGKQVEI 113
Score = 42.7 bits (99), Expect = 7e-05
Identities = 19/30 (63%), Positives = 20/30 (66%)
Query: 116 AGGPRPGGGYGGGGGYSGGGGGYSGGGGGG 145
AGG GGG GGG + GGGGG G GGGG
Sbjct: 385 AGGYGAGGGGNGGGSFYGGGGGRGGYGGGG 414
Score = 40.8 bits (94), Expect = 3e-04
Identities = 18/28 (64%), Positives = 18/28 (64%)
Query: 117 GGPRPGGGYGGGGGYSGGGGGYSGGGGG 144
GG R GGGY GG GG GGY GGGG
Sbjct: 367 GGYRGGGGYDMGGVGGGGAGGYGAGGGG 394
Score = 38.9 bits (89), Expect = 0.001
Identities = 20/36 (55%), Positives = 21/36 (57%), Gaps = 10/36 (27%)
Query: 121 PGGGYGG-----GGGYS-----GGGGGYSGGGGGGW 146
PGGGYGG GGGY G GGG G GGGG+
Sbjct: 334 PGGGYGGPSGSYGGGYGSSGIGGYGGGMGGAGGGGY 369
Score = 38.5 bits (88), Expect = 0.001
Identities = 19/33 (57%), Positives = 19/33 (57%), Gaps = 5/33 (15%)
Query: 118 GPRPGGGYGGGGGYS-----GGGGGYSGGGGGG 145
G GGGY GGGGY GGG G G GGGG
Sbjct: 362 GGAGGGGYRGGGGYDMGGVGGGGAGGYGAGGGG 394
Score = 38.1 bits (87), Expect = 0.002
Identities = 19/34 (55%), Positives = 20/34 (57%), Gaps = 4/34 (11%)
Query: 117 GGPRPGGGYGGGGGYSGGG----GGYSGGGGGGW 146
GG G G GGGGY GGG GG GGG GG+
Sbjct: 355 GGYGGGMGGAGGGGYRGGGGYDMGGVGGGGAGGY 388
Score = 37.7 bits (86), Expect = 0.002
Identities = 22/36 (61%), Positives = 22/36 (61%), Gaps = 9/36 (25%)
Query: 117 GGPRP--GGGYG--GGGGYSG-----GGGGYSGGGG 143
GGP GGGYG G GGY G GGGGY GGGG
Sbjct: 339 GGPSGSYGGGYGSSGIGGYGGGMGGAGGGGYRGGGG 374
Score = 37.0 bits (84), Expect = 0.004
Identities = 18/30 (60%), Positives = 18/30 (60%), Gaps = 3/30 (10%)
Query: 114 RQAGGPRPGGGYGGGGGYSGGGGGYSGGGG 143
R GG GGGY GG GGGGGY GG G
Sbjct: 298 RYGGG---GGGYNRGGYSMGGGGGYGGGPG 324
Score = 34.7 bits (78), Expect = 0.018
Identities = 18/27 (66%), Positives = 18/27 (66%), Gaps = 2/27 (7%)
Query: 116 AGGPRPGGG--YGGGGGYSGGGGGYSG 140
AGG GGG YGGGGG G GGG SG
Sbjct: 390 AGGGGNGGGSFYGGGGGRGGYGGGGSG 416
Score = 33.9 bits (76), Expect = 0.032
Identities = 19/35 (54%), Positives = 19/35 (54%), Gaps = 7/35 (20%)
Query: 117 GGPRPGGGY--GGGGGYSGG-----GGGYSGGGGG 144
GG GGY GGGGGY GG GG Y GGG
Sbjct: 303 GGGYNRGGYSMGGGGGYGGGPGDMYGGSYGEPGGG 337
Score = 33.5 bits (75), Expect = 0.041
Identities = 15/21 (71%), Positives = 17/21 (80%), Gaps = 2/21 (9%)
Query: 126 GGGGGYSGGGGGYSGGGGGGW 146
GGGGGY+ GGYS GGGGG+
Sbjct: 301 GGGGGYN--RGGYSMGGGGGY 319
Score = 32.7 bits (73), Expect = 0.070
Identities = 18/34 (52%), Positives = 19/34 (54%), Gaps = 9/34 (26%)
Query: 122 GGGYGGG------GGYSGGGGGY---SGGGGGGW 146
GGGYGGG G Y GGGY SG GGG+
Sbjct: 316 GGGYGGGPGDMYGGSYGEPGGGYGGPSGSYGGGY 349
Score = 32.0 bits (71), Expect = 0.12
Identities = 17/30 (56%), Positives = 17/30 (56%), Gaps = 3/30 (10%)
Query: 117 GGPRP--GGGYG-GGGGYSGGGGGYSGGGG 143
GGP GG YG GGGY G G Y GG G
Sbjct: 321 GGPGDMYGGSYGEPGGGYGGPSGSYGGGYG 350
Score = 29.3 bits (64), Expect = 0.78
Identities = 17/32 (53%), Positives = 17/32 (53%), Gaps = 3/32 (9%)
Query: 117 GGPRPGGGYGGGGG---YSGGGGGYSGGGGGG 145
GG GGG G GGG GG G GGG GG
Sbjct: 309 GGYSMGGGGGYGGGPGDMYGGSYGEPGGGYGG 340
>At4g24770 31 kDa RNA binding protein (rbp31)
Length = 329
Score = 80.5 bits (197), Expect = 3e-16
Identities = 42/87 (48%), Positives = 59/87 (67%), Gaps = 4/87 (4%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSALS- 95
++++G L ++VD+ L F+ +G VVEARV+ DRETGRSRGFGFV + + A+S
Sbjct: 245 RVYVGNLPWDVDNGRLEQLFSEHGKVVEARVVYDRETGRSRGFGFVTMSDVDELNEAISA 304
Query: 96 MDGQDLNGRNIRVSYANDRQAGGPRPG 122
+DGQ+L GR IRV+ A +R PR G
Sbjct: 305 LDGQNLEGRAIRVNVAEERP---PRRG 328
Score = 66.2 bits (160), Expect = 6e-12
Identities = 41/104 (39%), Positives = 59/104 (56%), Gaps = 4/104 (3%)
Query: 19 TQAPVSSMLNYLRHMSSSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRG 78
++ VS + +KLF+G L+Y+V+ Q+L F G V A VI +RET +SRG
Sbjct: 133 SEGAVSERAEFPEPSEEAKLFVGNLAYDVNSQALAMLFEQAGTVEIAEVIYNRETDQSRG 192
Query: 79 FGFVNFTS-EESATSALSMDGQDLNGRNIRVSYANDRQAGGPRP 121
FGFV +S +E+ T+ + DLNGR + V+ A R G RP
Sbjct: 193 FGFVTMSSVDEAETAVEKFNRYDLNGRLLTVNKAAPR---GSRP 233
>At5g50250 RNA-binding protein-like
Length = 289
Score = 79.0 bits (193), Expect = 9e-16
Identities = 37/79 (46%), Positives = 57/79 (71%), Gaps = 1/79 (1%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
++++G L ++VD L F+ +G VV+ARV++DRETGRSRGFGFV ++E A+ +
Sbjct: 208 RIYVGNLPWDVDSGRLERLFSEHGKVVDARVVSDRETGRSRGFGFVQMSNENEVNVAIAA 267
Query: 96 MDGQDLNGRNIRVSYANDR 114
+DGQ+L GR I+V+ A +R
Sbjct: 268 LDGQNLEGRAIKVNVAEER 286
Score = 60.8 bits (146), Expect = 2e-10
Identities = 36/87 (41%), Positives = 52/87 (59%), Gaps = 4/87 (4%)
Query: 36 SKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL- 94
+KLF+G L Y+VD Q+L F G V + VI +R+T +SRGFGFV ++ E A A+
Sbjct: 113 AKLFVGNLPYDVDSQALAMLFEQAGTVEISEVIYNRDTDQSRGFGFVTMSTVEEAEKAVE 172
Query: 95 SMDGQDLNGRNIRVSYANDRQAGGPRP 121
+ ++NGR + V+ A R G RP
Sbjct: 173 KFNSFEVNGRRLTVNRAAPR---GSRP 196
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,673,930
Number of Sequences: 26719
Number of extensions: 183310
Number of successful extensions: 11864
Number of sequences better than 10.0: 533
Number of HSP's better than 10.0 without gapping: 462
Number of HSP's successfully gapped in prelim test: 77
Number of HSP's that attempted gapping in prelim test: 1163
Number of HSP's gapped (non-prelim): 4161
length of query: 146
length of database: 11,318,596
effective HSP length: 90
effective length of query: 56
effective length of database: 8,913,886
effective search space: 499177616
effective search space used: 499177616
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146705.10


