

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146664.9 - phase: 0
(59 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
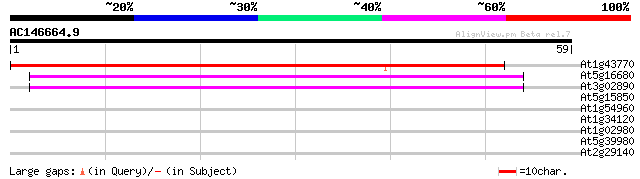
Score E
Sequences producing significant alignments: (bits) Value
At1g43770 hypothetical protein 53 3e-08
At5g16680 putative protein 44 2e-05
At3g02890 unknown protein 40 2e-04
At5g15850 CONSTANS-like 1 26 3.3
At1g54960 NPK1-related protein kinase 2 (ANP2) 25 5.6
At1g34120 putative inositol polyphosphate 5-phosphatase At5P1 25 5.6
At1g02980 25 5.6
At5g39980 putative protein 25 7.3
At2g29140 putative pumilio/Mpt5 family RNA-binding protein 25 9.5
>At1g43770 hypothetical protein
Length = 431
Score = 52.8 bits (125), Expect = 3e-08
Identities = 24/53 (45%), Positives = 35/53 (65%), Gaps = 1/53 (1%)
Query: 1 MGHLSTLACPKVHEEARYLPNMISANFLQKSTVWPESF-KNSGTNNFSIGIYF 52
+ H+S+LACPKVHE A L +SA L + VWP++F KN G + S+ ++F
Sbjct: 311 VAHVSSLACPKVHETASSLKGRLSAEILPRLEVWPKTFLKNGGPKDESVALFF 363
>At5g16680 putative protein
Length = 1280
Score = 43.5 bits (101), Expect = 2e-05
Identities = 19/52 (36%), Positives = 27/52 (51%)
Query: 3 HLSTLACPKVHEEARYLPNMISANFLQKSTVWPESFKNSGTNNFSIGIYFLS 54
HLSTLA P+V E P S N + + + WP F+ GT I ++F +
Sbjct: 829 HLSTLASPRVAEVVNKFPETFSLNEVPRKSTWPTQFEKLGTKEAHIALFFFA 880
>At3g02890 unknown protein
Length = 963
Score = 40.4 bits (93), Expect = 2e-04
Identities = 16/52 (30%), Positives = 30/52 (56%)
Query: 3 HLSTLACPKVHEEARYLPNMISANFLQKSTVWPESFKNSGTNNFSIGIYFLS 54
+LSTLA PKV E + P ++ N + + + WP F+++G + ++F +
Sbjct: 692 YLSTLASPKVVEVVKQFPEKVTLNEVPRLSSWPAQFQDTGAKEQHVALFFFA 743
>At5g15850 CONSTANS-like 1
Length = 355
Score = 26.2 bits (56), Expect = 3.3
Identities = 15/25 (60%), Positives = 17/25 (68%), Gaps = 1/25 (4%)
Query: 31 STVWPESFKNSGTNN-FSIGIYFLS 54
S + P S KNSG NN FSIG FL+
Sbjct: 141 SWLLPNSGKNSGNNNGFSIGDEFLN 165
>At1g54960 NPK1-related protein kinase 2 (ANP2)
Length = 651
Score = 25.4 bits (54), Expect = 5.6
Identities = 11/39 (28%), Positives = 21/39 (53%)
Query: 18 YLPNMISANFLQKSTVWPESFKNSGTNNFSIGIYFLSPH 56
++P ++ L+K +PES + TN +G+ +L H
Sbjct: 152 FVPGGSISSLLEKFGAFPESVVRTYTNQLLLGLEYLHNH 190
>At1g34120 putative inositol polyphosphate 5-phosphatase At5P1
Length = 586
Score = 25.4 bits (54), Expect = 5.6
Identities = 12/46 (26%), Positives = 23/46 (49%)
Query: 11 KVHEEARYLPNMISANFLQKSTVWPESFKNSGTNNFSIGIYFLSPH 56
++H ++LP+ ++AN L +S E G N+ I + + H
Sbjct: 416 EIHRRTQFLPHSLNANELPRSICNHERIIWLGDLNYRINLSYEKTH 461
>At1g02980
Length = 648
Score = 25.4 bits (54), Expect = 5.6
Identities = 11/37 (29%), Positives = 22/37 (58%)
Query: 3 HLSTLACPKVHEEARYLPNMISANFLQKSTVWPESFK 39
HLS + P + + + ++IS ++L++ T P +FK
Sbjct: 609 HLSKMFKPDIKMIKKRIEDLISRDYLERDTDNPNTFK 645
>At5g39980 putative protein
Length = 678
Score = 25.0 bits (53), Expect = 7.3
Identities = 9/27 (33%), Positives = 18/27 (66%)
Query: 12 VHEEARYLPNMISANFLQKSTVWPESF 38
VHEEA+Y P++ + N + ++ + + F
Sbjct: 145 VHEEAKYTPSVFAYNVVLRNVLRAKQF 171
>At2g29140 putative pumilio/Mpt5 family RNA-binding protein
Length = 964
Score = 24.6 bits (52), Expect = 9.5
Identities = 16/47 (34%), Positives = 23/47 (48%), Gaps = 1/47 (2%)
Query: 2 GHLSTLACPKVHEEARYLPNMISANFLQKSTVWPESFKNSGTNNFSI 48
GH S LA P H+ AR+ N S + L ++ ++ NS N I
Sbjct: 468 GHPSHLADPMYHQYARFSENADSFDLLNDPSM-DRNYGNSYMNMLEI 513
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.133 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,422,645
Number of Sequences: 26719
Number of extensions: 43638
Number of successful extensions: 99
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 92
Number of HSP's gapped (non-prelim): 9
length of query: 59
length of database: 11,318,596
effective HSP length: 35
effective length of query: 24
effective length of database: 10,383,431
effective search space: 249202344
effective search space used: 249202344
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146664.9


